Abstract
Development of functional nanoparticles can be encumbered by unanticipated material properties and biological events, which can affect nanoparticle effectiveness in complex, physiologically relevant systems1,2,3. Despite the advances in bottom-up nanoengineering and surface chemistry, reductionist functionalization approaches remain inadequate in replicating the complex interfaces present in nature and cannot avoid exposure of foreign materials. Here we report on the preparation of polymeric nanoparticles enclosed in the plasma membrane of human platelets, which are a unique population of cellular fragments that adhere to a variety of disease-relevant substrates4,5,6,7. The resulting nanoparticles possess a right-side-out unilamellar membrane coating functionalized with immunomodulatory and adhesion antigens associated with platelets. Compared to uncoated particles, the platelet membrane-cloaked nanoparticles have reduced cellular uptake by macrophage-like cells and lack particle-induced complement activation in autologous human plasma. The cloaked nanoparticles also display platelet-mimicking properties such as selective adhesion to damaged human and rodent vasculatures as well as enhanced binding to platelet-adhering pathogens. In an experimental rat model of coronary restenosis and a mouse model of systemic bacterial infection, docetaxel and vancomycin, respectively, show enhanced therapeutic efficacy when delivered by the platelet-mimetic nanoparticles. The multifaceted biointerfacing enabled by the platelet membrane cloaking method provides a new approach in developing functional nanoparticles for disease-targeted delivery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Pelaz, B. et al. Interfacing engineered nanoparticles with biological systems: anticipating adverse nano-bio interactions. Small 9, 1573–1584 (2013)
Salvati, A. et al. Transferrin-functionalized nanoparticles lose their targeting capabilities when a biomolecule corona adsorbs on the surface. Nature Nanotechnol. 8, 137–143 (2013)
Tenzer, S. et al. Rapid formation of plasma protein corona critically affects nanoparticle pathophysiology. Nature Nanotechnol. 8, 772–781 (2013)
Born, G. V. & Cross, M. J. The aggregation of blood platelets. J. Physiol. (Lond.) 168, 178–195 (1963)
Kieffer, N. & Phillips, D. R. Platelet membrane glycoproteins: functions in cellular interactions. Annu. Rev. Cell Biol. 6, 329–357 (1990)
Fitzgerald, J. R., Foster, T. J. & Cox, D. The interaction of bacterial pathogens with platelets. Nature Rev. Microbiol. 4, 445–457 (2006)
Yeaman, M. R. Platelets in defense against bacterial pathogens. Cell. Mol. Life Sci. 67, 525–544 (2010)
Peters, D. et al. Targeting atherosclerosis by using modular, multifunctional micelles. Proc. Natl Acad. Sci. USA 106, 9815–9819 (2009)
Chan, J. M. et al. Spatiotemporal controlled delivery of nanoparticles to injured vasculature. Proc. Natl Acad. Sci. USA 107, 2213–2218 (2010)
Bertram, J. P. et al. Intravenous hemostat: nanotechnology to halt bleeding. Sci. Transl. Med. 1, 11ra22 (2009)
Modery-Pawlowski, C. L. et al. Approaches to synthetic platelet analogs. Biomaterials 34, 526–541 (2013)
Simberg, D. et al. Biomimetic amplification of nanoparticle homing to tumors. Proc. Natl Acad. Sci. USA 104, 932–936 (2007)
Anselmo, A. C. et al. Platelet-like nanoparticles: mimicking shape, flexibility, and surface biology of platelets to target vascular injuries. ACS Nano 8, 11243–11253 (2014)
Olsson, M., Bruhns, P., Frazier, W. A., Ravetch, J. V. & Oldenborg, P. A. Platelet homeostasis is regulated by platelet expression of CD47 under normal conditions and in passive immune thrombocytopenia. Blood 105, 3577–3582 (2005)
Sims, P. J., Rollins, S. A. & Wiedmer, T. Regulatory control of complement on blood platelets. Modulation of platelet procoagulant responses by a membrane inhibitor of the C5b-9 complex. J. Biol. Chem. 264, 19228–19235 (1989)
Nieswandt, B. & Watson, S. P. Platelet-collagen interaction: is GPVI the central receptor? Blood 102, 449–461 (2003)
Hu, C. M. et al. Erythrocyte membrane-camouflaged polymeric nanoparticles as a biomimetic delivery platform. Proc. Natl Acad. Sci. USA 108, 10980–10985 (2011)
Hu, C. M., Fang, R. H., Copp, J., Luk, B. T. & Zhang, L. A biomimetic nanosponge that absorbs pore-forming toxins. Nature Nanotechnol. 8, 336–340 (2013)
Hu, C. M., Fang, R. H., Luk, B. T. & Zhang, L. Nanoparticle-detained toxins for safe and effective vaccination. Nature Nanotechnol. 8, 933–938 (2013)
Gachet, C. et al. Alpha IIb beta 3 integrin dissociation induced by EDTA results in morphological changes of the platelet surface-connected canalicular system with differential location of the two separate subunits. J. Cell Biol. 120, 1021–1030 (1993)
Luk, B. T. et al. Interfacial interactions between natural RBC membranes and synthetic polymeric nanoparticles. Nanoscale 6, 2730–2737 (2014)
Hughes, C. E. et al. CLEC-2 activates Syk through dimerization. Blood 115, 2947–2955 (2010)
Kalluri, R. Basement membranes: structure, assembly and role in tumour angiogenesis. Nature Rev. Cancer 3, 422–433 (2003)
Rodriguez, P. L. et al. Minimal “Self” peptides that inhibit phagocytic clearance and enhance delivery of nanoparticles. Science 339, 971–975 (2013)
Law, S. K. A. & Dodds, A. W. The internal thioester and the covalent binding properties of the complement proteins C3 and C4. Protein Sci. 6, 263–274 (1997)
Terstappen, L. W. M. M., Nguyen, M., Lazarus, H. M. & Medof, M. E. Expression of the DAF (CD55) and CD59 antigens during normal hematopoietic cell differentiation. J. Leukoc. Biol. 52, 652–660 (1992)
Andersen, A. J., Hashemi, S. H., Andresen, T. L., Hunter, A. C. & Moghimi, S. M. Complement: alive and kicking nanomedicines. J. Biomed. Nanotechnol. 5, 364–372 (2009)
Siboo, I. R., Chambers, H. F. & Sullam, P. M. Role of SraP, a Serine-Rich Surface Protein of Staphylococcus aureus, in binding to human platelets. Infect. Immun. 73, 2273–2280 (2005)
Kamaly, N. et al. Development and in vivo efficacy of targeted polymeric inflammation-resolving nanoparticles. Proc. Natl Acad. Sci. USA 110, 6506–6511 (2013)
Hu, C. M. et al. ‘Marker-of-self’ functionalization of nanoscale particles through a top-down cellular membrane coating approach. Nanoscale 5, 2664–2668 (2013)
Acknowledgements
This work is supported by the National Institutes of Health under Award Numbers R01DK095168 (L.Z.), R01HL108735 (S.C.) and R01EY25090 (K.Z.), and partially by the Defense Threat Reduction Agency Joint Science and Technology Office for Chemical and Biological Defense under Grant Number HDTRA1-14-1-0064 (L.Z.). R.H.F. is supported by a National Institutes of Health R25CA153915 training grant from the National Cancer Institute.
Author information
Authors and Affiliations
Contributions
C.-M.J.H., R.H.F., K.-C.W., B.T.L., K.Z., S.C. and L.Z. conceived and designed the experiments; C.-M.J.H., R.H.F., K.-C.W., B.T.L., S.T., D.D., P.N., P.A., C.H.W., A.V.K., C.C., M.R., V.Q., S.H.P., J.Z., W.S., F.M.H., T.C.C. and W.G. performed all the experiments. The manuscript was written by C.-M.J.H., R.H.F., B.T.L., W.G. and L.Z. All authors discussed the results and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Schematic preparation of PNPs.
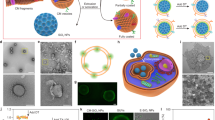
a, Poly(lactic-co-glycolic acid) (PLGA) nanoparticles are enclosed entirely in plasma membrane derived from human platelets. The resulting particles possess platelet-mimicking properties for immunocompatibility, subendothelium binding, and pathogen adhesion. b, Schematic depicting the process of preparing PNPs.
Extended Data Figure 2 PNP preparation and storage.
a, Isolation of platelet rich plasma (PRP) was achieved by centrifugation at 100g. PRP was collected from the top layer (yellow) separated from the red blood cells (red, bottom layer). b, Collected human platelets under light microscopy, which possess a distinctive morphology from c, red blood cells. Scale bars, 10 μm. d, Transmission electron micrographs of platelet membrane vesicles and e, PNPs, both of which were negatively stained with 1% uranyl acetate. Scale bars, 200 nm. f, Dynamic light scattering measurements of PNPs in 10% sucrose show that the particles retain their size and stability after a freeze-thaw cycle and re-suspension upon lyophilization (n = 3). Bars represent means ± s.d. g, Transmission electron micrograph shows retentions of the core-shell structure of PNPs after a freeze-thaw cycle in 10% sucrose. Scale bar, 100 nm. h, Transmission electron micrograph shows retentions of the core-shell structure of PNPs upon resuspension after lyophilization in 10% sucrose. Scale bar, 100 nm.
Extended Data Figure 3 Overall protein content on PNPs resolved by western blotting.
Primary platelet membrane protein/protein subunits including CD47, CD55, CD59, αIIb, α2, α5, α6, β1, β3, GPIbα, GPIV, GPV, GPVI, GPIX, and CLEC-2 were monitored in platelet rich plasma, platelets, platelet vesicles, and PNPs. Platelets derived from four different protocols, including commercial blood anti-coagulated in EDTA, freshly drawn blood anti-coagulated in EDTA, freshly drawn blood anti-coagulated in heparin, and transfusion-grade platelet rich plasma anti-coagulated in acid-citrate-dextrose (ACD), were examined to compare the membrane protein expression. Each sample was normalized to equivalent overall protein content before western blotting. It was observed that the PNP preparation resulted in membrane protein retention and enrichment very similar across the different platelet sources.
Extended Data Figure 4 Platelet membrane sidedness on PNPs.
a, Transmission electron micrograph of PNPs primary-stained with anti-CD47 (intracellular), secondary-stained with immunogold, and negatively stained with 2% vanadium. The immunogold staining revealed presence of intracellular CD47 domains on collapsed platelet membrane vesicles, but not on PNPs. b, Transmission electron micrograph of PNPs primary-stained with anti-CD47 (extracellular), secondary-stained with immunogold, and negatively stained with 2% vanadium. PNPs were shown to display extracellular CD47 domains. All scale bars, 100 nm. c, 2 μm polystyrene beads were functionalized with anti-CD47 against the protein’s extracellular domain, anti-CD47 against the protein’s intracellular domain, or a sham antibody. Flow cytometric analysis of the different beads after DiD-loaded PNP incubation showed the highest particle retention to beads functionalized with anti-CD47 against the protein’s extracellular domain. d, Normalized fluorescence intensity of PNP retention to the different antibody-functionalized beads. Bars represent means ± s.e.m.
Extended Data Figure 5 PNP binding to collagen and extracellular matrix.
a–f, Collagen-coated tissue culture slides seeded with human umbilical vein endothelial cells (HUVECs) were incubated with PNP solution for 30 s. Fluorescence microscopy samples demonstrate differential PNP adherence to exposed collagen versus covered endothelial surfaces. a–c, Representative fluorescence images visualizing DiD-loaded PNPs (red), cellular cytosol (green), and cellular nuclei (blue). d–f, Images showing only the red and blue channels to highlight the differential localization of PNPs. Scale bar, 10 μm. g, Fluorescence quantification of PNP per unit area on collagen and endothelial surfaces. Bars represent means ± s.d. (n = 10). h, i, PNP adherence to arterial extracellular matrix (ECM) as visualized by SEM. h, SEM images of the ECM of a decellularized human umbilical cord artery. Left, scale bar, 1 μm; right, scale bar, 500 nm. i, SEM images of the ECM of a decellularized human umbilical cord artery after PNP incubation. Left, scale bar, 1 μm; right, scale bar, 500 nm.
Extended Data Figure 6 Pharmacokinetics, biodistribution and safety of PNPs.
a, DiD-loaded PNPs were injected intravenously through the tail vein of Sprague–Dawley rats. At various time points, blood was withdrawn via tail vein blood sampling for fluorescence quantification to evaluate the systemic circulation lifetime of the nanoparticles (n = 6). b, Biodistribution of the PNP nanoparticles in balloon-denuded Sprague–Dawley rats at 2 h and 48 h after intravenous nanoparticle administration through the tail vein (n = 6). c, Comprehensive metabolic panel of rats after injections with human-derived PNPs and PBS (n = 6). The rats received intravenous injections of PNPs and PBS on day 0 and day 5, and the blood test conducted on day 10 did not reveal significant changes between the two groups, indicating normal liver and kidney functions after the PNP administration. All bars and markers represent means ± s.d.
Extended Data Figure 7 PNP targeting of damaged vasculatures upon intravenous injection to rats with angioplasty-induced arterial denudation.
a, Fluorescence microscopy of the aortic branch revealed selective PNP binding to the denuded artery (right) as opposed to the undamaged artery (left) (PNP fluorescence in red). b, Fluorescence images acquired from the control artery, which did not reveal PNP fluorescence upon focusing on either the endothelium (top) or the smooth muscle layer (bottom) (nuclei in blue). c, Fluorescence images acquired from the denuded artery, which revealed significant PNP retention as fluorescent punctates (PNP fluorescence in red) above the smooth muscle layer. d, Fluorescence image of arterial cross-section acquired from the control artery, which showed nuclei of endothelial cells above the collagen layer (autofluorescence in green) and an absence of PNP fluorescence. e, Fluorescence image of arterial cross-section acquired from the denuded artery, which showed PNP retention as fluorescent punctates on the collagen layer (PNP fluorescence in red; collagen autofluorescence in green) and an absence of endothelial cell nuclei. All scale bars, 100 μm.
Extended Data Figure 8 Characterizations of drug-loaded cell membrane cloaked nanoparticles.
a, Physicochemical properties of drug-loaded cell membrane cloaked nanoparticles. b, TEM visualization of docetaxel-loaded PNPs (PNP-Dtxl). Scale bar, 200 nm. c, Drug release profile of PNP-Dtxl compared to polyethylene glycol (PEG)-PLGA diblock nanoparticles of equivalent size and docetaxel loading (n = 3). d, TEM visualization of vancomycin-loaded PNPs (PNP-Vanc). Scale bar, 200 nm. e, Drug release profiles of PNP-Vanc and RBCNP-Vanc (n = 3). Bars represent means ± s.d.
Extended Data Figure 9 Treatment of an experimental rat model of coronary restenosis.
a–e, H&E-stained arterial cross-sections reveal the vascular structure of non-damaged arteries (serving as baseline, a) and denuded arteries after treatment with PNP-Dtxl (b), PBS (c), PNP with no docetaxel content (d), or free docetaxel (e). Scale bar, 200 μm.
Extended Data Figure 10 PNP adherence to MRSA252 bacteria.
a, Flow cytometric analysis of MRSA252 bacteria after incubation with different DiD-loaded nanoformulations. b, Pellets of MRSA252 after incubation with DiD-loaded RBCNPs (left) and DiD-loaded PNPs (right) show differential retention of nanoformulation with MRSA252 upon pelleting of the bacteria. c, A pseudocoloured SEM image of PNPs binding to MRSA252 under high magnification (MRSA coloured in gold, PNP coloured in orange). Scale bar, 400 nm.
Rights and permissions
About this article
Cite this article
Hu, CM., Fang, R., Wang, KC. et al. Nanoparticle biointerfacing by platelet membrane cloaking. Nature 526, 118–121 (2015). https://doi.org/10.1038/nature15373
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15373
This article is cited by
-
Physiological principles underlying the kidney targeting of renal nanomedicines
Nature Reviews Nephrology (2024)
-
Molecular Mechanisms of Intracellular Delivery of Nanoparticles Monitored by an Enzyme-Induced Proximity Labeling
Nano-Micro Letters (2024)
-
Acetylcholine Analog-Modified Albumin Nanoparticles for the Enhanced and Synchronous Brain Delivery of Saponin Components of Panax Notoginseng
Pharmaceutical Research (2024)
-
Systematic evaluation of membrane-camouflaged nanoparticles in neutralizing Clostridium perfringens ε-toxin
Journal of Nanobiotechnology (2023)
-
Expanding applications of allogeneic platelets, platelet lysates, and platelet extracellular vesicles in cell therapy, regenerative medicine, and targeted drug delivery
Journal of Biomedical Science (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.