Abstract
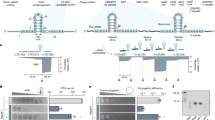
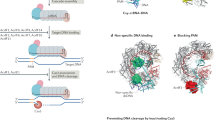
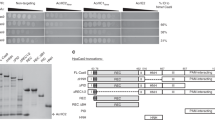
The battle for survival between bacteria and the viruses that infect them (phages) has led to the evolution of many bacterial defence systems and phage-encoded antagonists of these systems. Clustered regularly interspaced short palindromic repeats (CRISPR) and the CRISPR-associated (cas) genes comprise an adaptive immune system that is one of the most widespread means by which bacteria defend themselves against phages1,2,3. We identified the first examples of proteins produced by phages that inhibit a CRISPR–Cas system4. Here we performed biochemical and in vivo investigations of three of these anti-CRISPR proteins, and show that each inhibits CRISPR–Cas activity through a distinct mechanism. Two block the DNA-binding activity of the CRISPR–Cas complex, yet do this by interacting with different protein subunits, and using steric or non-steric modes of inhibition. The third anti-CRISPR protein operates by binding to the Cas3 helicase–nuclease and preventing its recruitment to the DNA-bound CRISPR–Cas complex. In vivo, this anti-CRISPR can convert the CRISPR–Cas system into a transcriptional repressor, providing the first example—to our knowledge—of modulation of CRISPR–Cas activity by a protein interactor. The diverse sequences and mechanisms of action of these anti-CRISPR proteins imply an independent evolution, and foreshadow the existence of other means by which proteins may alter CRISPR–Cas function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007)
Makarova, K. S. et al. Evolution and classification of the CRISPR-Cas systems. Nature Rev. Microbiol. 9, 467–477 (2011)
Jore, M. M., Brouns, S. J. J. & van der Oost, J. RNA in defense: CRISPRs protect prokaryotes against mobile genetic elements. Cold Spring Harb. Perspect. Biol. 4, a003657 (2012)
Bondy-Denomy, J., Pawluk, A., Maxwell, K. L. & Davidson, A. R. Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 493, 429–432 (2013)
van der Oost, J., Westra, E. R., Jackson, R. N. & Wiedenheft, B. Unravelling the structural and mechanistic basis of CRISPR–Cas systems. Nature Rev. Microbiol. 12, 479–492 (2014)
Haurwitz, R. E., Jinek, M., Wiedenheft, B., Zhou, K. & Doudna, J. A. Sequence- and structure-specific RNA processing by a CRISPR endonuclease. Science 329, 1355–1358 (2010)
Wiedenheft, B. et al. RNA-guided complex from a bacterial immune system enhances target recognition through seed sequence interactions. Proc. Natl Acad. Sci. USA 108, 10092–10097 (2011)
van Duijn, E. et al. Native tandem and ion mobility mass spectrometry highlight structural and modular similarities in clustered-regularly-interspaced shot-palindromic-repeats (CRISPR)-associated protein complexes from Escherichia coli and Pseudomonas aeruginosa. Mol. Cell. Proteomics 11, 1430–1441 (2012)
Westra, E. R. et al. CRISPR immunity relies on the consecutive binding and degradation of negatively supercoiled invader DNA by Cascade and Cas3. Mol. Cell 46, 595–605 (2012)
Huo, Y. et al. Structures of CRISPR Cas3 offer mechanistic insights into Cascade-activated DNA unwinding and degradation. Nature Struct. Mol. Biol. 21, 771–777 (2014)
Rollins, M. F., Schuman, J. T., Paulus, K., Bukhari, H. S. T. & Wiedenheft, B. Mechanism of foreign DNA recognition by a CRISPR RNA-guided surveillance complex from Pseudomonas aeruginosa. Nucleic Acids Res. 43, 2216–2222 (2015)
Semenova, E. et al. Interference by clustered regularly interspaced short palindromic repeat (CRISPR) RNA is governed by a seed sequence. Proc. Natl Acad. Sci. USA 108, 10098–10103 (2011)
Wurtzel, O. et al. The single-nucleotide resolution transcriptome of Pseudomonas aeruginosa grown in body temperature. PLoS Pathog. 8, e1002945 (2012)
Luo, M. L., Mullis, A. S., Leenay, R. T. & Beisel, C. L. Repurposing endogenous type I CRISPR-Cas systems for programmable gene repression. Nucleic Acids Res. 43, 674–681 (2015)
Rath, D., Amlinger, L., Hoekzema, M., Devulapally, P. R. & Lundgren, M. Efficient programmable gene silencing by Cascade. Nucleic Acids Res. 43, 237–246 (2015)
Cady, K. C. & O’Toole, G. A. Non-identity-mediated CRISPR-bacteriophage interaction mediated via the Csy and Cas3 proteins. J. Bacteriol. 193, 3433–3445 (2011)
Jackson, R. N. et al. Crystal structure of the CRISPR RNA-guided surveillance complex from Escherichia coli. Science 345, 1473–1479 (2014)
Mulepati, S., Héroux, A. & Bailey, S. Crystal structure of a CRISPR RNA-guided surveillance complex bound to a ssDNA target. Science 345, 1479–1484 (2014)
Pawluk, A., Bondy-Denomy, J., Cheung, V. H. W., Maxwell, K. L. & Davidson, A. R. A new group of phage anti-CRISPR genes inhibits the type I-E CRISPR-Cas system of Pseudomonas aeruginosa. MBio 5, e00896 (2014)
Westra, E. R., Buckling, A. & Fineran, P. C. CRISPR–Cas systems: beyond adaptive immunity. Nature Rev. Microbiol. 12, 317–326 (2014)
Cady, K. C., Bondy-Denomy, J., Heussler, G. E., Davidson, A. R. & O’Toole, G. A. The CRISPR/Cas adaptive immune system of Pseudomonas aeruginosa mediates resistance to naturally occurring and engineered phages. J. Bacteriol. 194, 5728–5738 (2012)
Acknowledgements
We thank W. Navarre and E. Westra for reading the manuscript. This work was supported by an Operating Grant to A.R.D. (MOP-130482) and to K.L.M. (MOP-136845), both of which were from the Canadian Institutes of Health Research (CIHR). J.B.-D. was supported by a CIHR Canada Graduate Scholarship Doctoral Award and an Ontario Graduate Scholarship award. Research in the Wiedenheft laboratory is supported by the National Institutes of Health (P20GM103500 and R01GM108888), the National Science Foundation EPSCoR (EPS-110134), the M.J. Murdock Charitable Trust, and the Montana State University Agricultural Experimental Station.
Author information
Authors and Affiliations
Contributions
J.B.-D. designed, performed and supervised experiments and wrote the manuscript. B.G., S.S., M.D., and Y.H.-R. performed experiments. M.F.R. and B.W. provided essential reagents and experimental assistance. K.L.M. supervised experiments. A.R.D. designed and supervised experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 AcrF2 interacts with the Csy complex.
a, b, Purified Csy complex was fractionated by SEC alone (a) or in the presence of AcrF2 (b). Fractions were analysed on a silver nitrate stained SDS–PAGE gel. The input (IN) and fractions are shown.
Extended Data Figure 2 AcrF3, not AcrF1, interacts with Cas3.
a, Cas3 was fractionated by SEC alone or in the presence of AcrF3 or AcrF1. Overlays of plots of elution volume versus optical density at 280 nm of the column eluates are shown. The numbers represent the fractions that were selected for analysis. b–e, Silver nitrate stained SDS–PAGE gels are shown from SEC experiments with Cas3 (b), AcrF3 (c), Cas3 with AcrF3 (d) or Cas3 with AcrF1 (e). The sample that was loaded onto the SEC column is shown as input (In) and fractions from the same elution positions are indicated numerically. AcrF3 is seen eluting in fractions 4–8 only in the presence of Cas3. There is also a visible shift in the Cas3 elution profile in the presence of AcrF3 but not AcrF1 (fractions 3–5).
Extended Data Figure 3 AcrF1 and AcrF2 prevent target recognition by the Csy complex.
Isothermal titration calorimetry (ITC) assays showing the Csy complex binding to an 8-nucleotide ssDNA target that comprises the seed region. No binding is observed in the presence of AcrF1, AcrF2 or with a non-target (the reverse complement sequence of the target) ssDNA substrate. A representative run is shown for each condition with the dissociation constant (Kd) value and error of fit from that particular run. Over multiple runs (n = 6) with the Csy complex binding to the ssDNA ligand, the average Kd value was 90 nM ± 37.
Extended Data Figure 4 Expression of phzM is repressed by the Csy complex.
The Csy complex was targeted to the promoter of the gene phzM, and repression efficiency was assayed by RT–qPCR (see Methods). The per cent repression of phzM in the indicated strains expressing a phzM-targeting crRNA relative to wild-type (WT) PA14 with an empty plasmid is shown. All values were normalized to rpsL, a gene encoding a ribosomal protein. Means ± s.d. are shown.
Extended Data Figure 5 AcrF4 interacts with the Csy complex.
Untagged AcrF4 was expressed in E. coli BL21 cells and a crude lysate of these cells was mixed with the Csy complex bound to Ni-NTA beads via a 6×His tag on Csy3. a, The flow through (FT), wash 1 (W1), and two elution fractions (E1, E2) from the Ni-NTA column are shown, as well as a comparison to pure Csy complex. b, The Ni-NTA elution fractions were fractionated by SEC, demonstrating a stable interaction between the Csy complex and AcrF4. The input (In) lane shows the sample that was loaded on the SEC column and numbered fractions are analysed on SDS–PAGE gels.
Extended Data Figure 6 AcrF1 and AcrF2 bind the Csy complex at distinct locations.
a, Purified Csy1–Csy2 heterodimer with an MBP and 6×His tag fused to Csy1 was fractionated by SEC in the presence or absence of AcrF1 (boxes indicate the Csy1–Csy2 peak). b, Purified MBP/6×His-tagged Csy3 was fractionated in the presence or absence of AcrF2. These are complementary experiments to those seen in Fig. 3b and c, respectively. Input (In) and selected fractions are shown on SDS–PAGE gels. c, AcrF1 and AcrF2 were incubated with the Csy complex singly or in combination. Asterisks designate which anti-CRISPR was added first to the reactions containing both anti-CRISPR proteins. The addition order did not affect the result since there is no competition for binding sites between these two anti-CRISPR proteins. After incubation, each mixture was fractionated by SEC and the peak Csy complex fraction is shown on an SDS–PAGE gel. In each experiment the anti-CRISPR proteins are in excess relative to the Csy complex.
Extended Data Figure 7 AcrF1 and AcrF2 interact with an RNase-A-treated Csy complex.
a, The Csy complex was treated with a low concentration (600 nM, +) of RNase A or a high concentration of RNase A (70 µM, ++). This mixture was fractionated by SEC, revealing Csy4 dissociation at the higher RNase A concentration. Pre-treatment of the Csy complex with RNase A, with the subsequent addition of AcrF1 or AcrF2 followed by SEC fractionation was then conducted. Peak Csy complex fractions are shown on an SDS–PAGE gel. b, A TBE-urea denaturing gel is shown, stained with SYBR gold, showing the native crRNA in the Csy complex and the protected fragments remaining after 70 µM RNase A treatment. c, Quantification of Coomassie blue stained gels from three independent preparations of the respective proteins is shown. Anti-CRISPR proteins bound with unaltered stoichiometry to RNase-A-pre-treated Csy complexes. Error bars represent s.d.
Extended Data Figure 8 Twofold dilutions used to quantify anti-CRISPR binding stoichiometry.
Csy complexes with crRNA molecules possessing spacers of differing lengths (16, 32, or 48 nucleotides) were purified and fractionated by SEC in the presence of AcrF1. A representative Coomassie blue stained SDS–PAGE gel is shown, with twofold dilutions of the peak fraction containing the Csy complex and co-eluting AcrF1. Arrows on the bottom of the gel indicate comparable dilutions based on the levels of Csy1. Note the increasing abundance of Csy3 and AcrF1. b, Lanes with arrows from the gel in a are shown next to each other for comparison.
Extended Data Figure 9 dsDNA binds to the Csy complex after SEC fractionation.
a, The same samples from Fig. 4a were run on a denaturing TBE-urea gel, stained with SYBR gold, to reveal the crRNA (two species are apparent), and the Csy-complex-bound 50 bp dsDNA. In these experiments, DNA was prebound to the Csy complex, and AcrF1 or AcrF2 were subsequently added to the DNA-saturated Csy complex. This mixture was then fractionated by SEC and the Csy-complex-containing peak fractions were analysed. b, A schematic showing the crRNA sequence with repeat-derived regions shown in black and the variable 32-nucleotide spacer region in red. The seed-interacting region that is critical for target recognition (nucleotides 1–5, 7, 8) is in bold. DNA oligonucleotides used in this study are shown, with labels ‘A’, ‘B’ and ‘C’ corresponding to the targets shown in Fig. 4c. The 8-nucleotide ssDNA substrate was used in ITC experiments (Extended Data Fig. 3), and the 50 bp dsDNA in EMSAs (Figs 1d and 4b).
Rights and permissions
About this article
Cite this article
Bondy-Denomy, J., Garcia, B., Strum, S. et al. Multiple mechanisms for CRISPR–Cas inhibition by anti-CRISPR proteins. Nature 526, 136–139 (2015). https://doi.org/10.1038/nature15254
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15254
This article is cited by
-
CRISPR-Cas9 mediated phage therapy as an alternative to antibiotics
Animal Diseases (2023)
-
Diversity and dynamics of the CRISPR-Cas systems associated with Bacteroides fragilis in human population
BMC Genomics (2022)
-
Alternative functions of CRISPR–Cas systems in the evolutionary arms race
Nature Reviews Microbiology (2022)
-
Disarming of type I-F CRISPR-Cas surveillance complex by anti-CRISPR proteins AcrIF6 and AcrIF9
Scientific Reports (2022)
-
Digging into the lesser-known aspects of CRISPR biology
International Microbiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.