Abstract
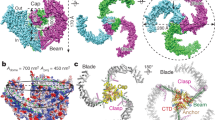
Piezo proteins are evolutionarily conserved and functionally diverse mechanosensitive cation channels. However, the overall structural architecture and gating mechanisms of Piezo channels have remained unknown. Here we determine the cryo-electron microscopy structure of the full-length (2,547 amino acids) mouse Piezo1 (Piezo1) at a resolution of 4.8 Å. Piezo1 forms a trimeric propeller-like structure (about 900 kilodalton), with the extracellular domains resembling three distal blades and a central cap. The transmembrane region has 14 apparently resolved segments per subunit. These segments form three peripheral wings and a central pore module that encloses a potential ion-conducting pore. The rather flexible extracellular blade domains are connected to the central intracellular domain by three long beam-like structures. This trimeric architecture suggests that Piezo1 may use its peripheral regions as force sensors to gate the central ion-conducting pore.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
The 3D cryo-electron microscopy density map has been deposited in the Electron Microscopy Data Bank (EMDB), with accession code EMD-6343. The coordinates of atomic models have been deposited in the Protein Data Bank (PDB) under the accession codes 4RAX for the CED and 3JAC for the full length.
Change history
04 November 2015
A grant number in the Acknowledgements was corrected.
References
Chalfie, M. Neurosensory mechanotransduction. Nature Rev. Mol. Cell Biol. 10, 44–52 (2009)
Coste, B. et al. Piezo1 and Piezo2 are essential components of distinct mechanically activated cation channels. Science 330, 55–60 (2010)
Coste, B. et al. Piezo proteins are pore-forming subunits of mechanically activated channels. Nature 483, 176–181 (2012)
Nilius, B. & Honoré, E. Sensing pressure with ion channels. Trends Neurosci. 35, 477–486 (2012)
Volkers, L., Mechioukhi, Y. & Coste, B. Piezo channels: from structure to function. Pflugers Arch. 467, 95–99 (2015)
Bae, C., Gottlieb, P. A. & Sachs, F. Human PIEZO1: removing inactivation. Biophys. J. 105, 880–886 (2013)
Gottlieb, P. A. & Sachs, F. Piezo1: properties of a cation selective mechanical channel. Channels 6, 214–219 (2012)
Kim, S. E., Coste, B., Chadha, A., Cook, B. & Patapoutian, A. The role of Drosophila Piezo in mechanical nociception. Nature 483, 209–212 (2012)
Faucherre, A., Nargeot, J., Mangoni, M. E. & Jopling, C. piezo2b regulates vertebrate light touch response. J. Neurosci. 33, 17089–17094 (2013)
Schneider, E. R. et al. Neuronal mechanism for acute mechanosensitivity in tactile-foraging waterfowl. Proc. Natl Acad. Sci. USA 111, 14941–14946 (2014)
Maksimovic, S. et al. Epidermal Merkel cells are mechanosensory cells that tune mammalian touch receptors. Nature 509, 617–621 (2014)
Woo, S. H. et al. Piezo2 is required for Merkel-cell mechanotransduction. Nature 509, 622–626 (2014)
Ranade, S. S. et al. Piezo2 is the major transducer of mechanical forces for touch sensation in mice. Nature 516, 121–125 (2014)
Ikeda, R. et al. Merkel cells transduce and encode tactile stimuli to drive Aβ-afferent impulses. Cell 157, 664–675 (2014)
Schrenk-Siemens, K. et al. PIEZO2 is required for mechanotransduction in human stem cell-derived touch receptors. Nature Neurosci. 18, 10–16 (2015)
Ranade, S. S. et al. Piezo1, a mechanically activated ion channel, is required for vascular development in mice. Proc. Natl Acad. Sci. USA 111, 10347–10352 (2014)
Li, J. et al. Piezo1 integration of vascular architecture with physiological force. Nature 515, 279–282 (2014)
Faucherre, A., Kissa, K., Nargeot, J., Mangoni, M. E. & Jopling, C. Piezo1 plays a role in erythrocyte volume homeostasis. Haematologica 99, 70–75 (2014)
Cahalan, S. M. et al. Piezo1 links mechanical forces to red blood cell volume. eLife 4, 07370 (2015)
McHugh, B. J. et al. Integrin activation by Fam38A uses a novel mechanism of R-Ras targeting to the endoplasmic reticulum. J. Cell Sci. 123, 51–61 (2010)
Pathak, M. M. et al. Stretch-activated ion channel Piezo1 directs lineage choice in human neural stem cells. Proc. Natl Acad. Sci. USA 111, 16148–16153 (2014)
Albuisson, J. et al. Dehydrated hereditary stomatocytosis linked to gain-of-function mutations in mechanically activated PIEZO1 ion channels. Nature Commun. 4, 1884 (2013)
Andolfo, I. et al. Multiple clinical forms of dehydrated hereditary stomatocytosis arise from mutations in PIEZO1 . Blood 121, 3925–3935 (2013)
Bae, C., Gnanasambandam, R., Nicolai, C., Sachs, F. & Gottlieb, P. A. Xerocytosis is caused by mutations that alter the kinetics of the mechanosensitive channel PIEZO1. Proc. Natl Acad. Sci. USA 110, E1162–E1168 (2013)
Beneteau, C. et al. Recurrent mutation in the PIEZO1 gene in two families of hereditary xerocytosis with fetal hydrops. Clin. Genet. 85, 293–295 (2014)
Shmukler, B. E. et al. Dehydrated stomatocytic anemia due to the heterozygous mutation R2456H in the mechanosensitive cation channel PIEZO1: a case report. Blood Cells Mol. Dis. 52, 53–54 (2014)
Zarychanski, R. et al. Mutations in the mechanotransduction protein PIEZO1 are associated with hereditary xerocytosis. Blood 120, 1908–1915 (2012)
Coste, B. et al. Gain-of-function mutations in the mechanically activated ion channel PIEZO2 cause a subtype of distal arthrogryposis. Proc. Natl Acad. Sci. USA 110, 4667–4672 (2013)
McMillin, M. J. et al. Mutations in PIEZO2 cause Gordon syndrome, Marden–Walker syndrome, and distal arthrogryposis type 5. Am. J. Hum. Genet. 94, 734–744 (2014)
Zhang, X. et al. Crystal structure of an orthologue of the NaChBac voltage-gated sodium channel. Nature 486, 130–134 (2012)
Payandeh, J., Scheuer, T., Zheng, N. & Catterall, W. A. The crystal structure of a voltage-gated sodium channel. Nature 475, 353–358 (2011)
Kawate, T., Michel, J. C., Birdsong, W. T. & Gouaux, E. Crystal structure of the ATP-gated P2X(4) ion channel in the closed state. Nature 460, 592–598 (2009)
Paulsen, C. E., Armache, J. P., Gao, Y., Cheng, Y. & Julius, D. Structure of the TRPA1 ion channel suggests regulatory mechanisms. Nature 520, 511–517 (2015)
Liao, M., Cao, E., Julius, D. & Cheng, Y. Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504, 107–112 (2013)
Liu, Z., Gandhi, C. S. & Rees, D. C. Structure of a tetrameric MscL in an expanded intermediate state. Nature 461, 120–124 (2009)
Bass, R. B., Strop, P., Barclay, M. & Rees, D. C. Crystal structure of Escherichia coli MscS, a voltage-modulated and mechanosensitive channel. Science 298, 1582–1587 (2002)
Chang, G., Spencer, R. H., Lee, A. T., Barclay, M. T. & Rees, D. C. Structure of the MscL homolog from Mycobacterium tuberculosis: a gated mechanosensitive ion channel. Science 282, 2220–2226 (1998)
Kung, C., Martinac, B. & Sukharev, S. Mechanosensitive channels in microbes. Annu. Rev. Microbiol. 64, 313–329 (2010)
Brohawn, S. G., del Mármol, J. & MacKinnon, R. Crystal structure of the human K2P TRAAK, a lipid- and mechano-sensitive K+ ion channel. Science 335, 436–441 (2012)
Coste, B. et al. Piezo1 ion channel pore properties are dictated by C-terminal region. Nature Commun. 6, 7223 (2015)
Hou, X., Pedi, L., Diver, M. M. & Long, S. B. Crystal structure of the calcium release-activated calcium channel Orai. Science 338, 1308–1313 (2012)
Penna, A. et al. The CRAC channel consists of a tetramer formed by Stim-induced dimerization of Orai dimers. Nature 456, 116–120 (2008)
Kamajaya, A., Kaiser, J. T., Lee, J., Reid, M. & Rees, D. C. The structure of a conserved Piezo channel domain reveals a topologically distinct β sandwich fold. Structure 22, 1520–1527 (2014)
Prole, D. L. & Taylor, C. W. Identification and analysis of putative homologues of mechanosensitive channels in pathogenic protozoa. PLoS One 8, e66068 (2013)
Gonzales, E. B., Kawate, T. & Gouaux, E. Pore architecture and ion sites in acid-sensing ion channels and P2X receptors. Nature 460, 599–604 (2009)
Vilceanu, D. & Stucky, C. L. TRPA1 mediates mechanical currents in the plasma membrane of mouse sensory neurons. PLoS One 5, e12177 (2010)
Kwan, K. Y., Glazer, J. M., Corey, D. P., Rice, F. L. & Stucky, C. L. TRPA1 modulates mechanotransduction in cutaneous sensory neurons. J. Neurosci. 29, 4808–4819 (2009)
Smart, O. S., Goodfellow, J. M. & Wallace, B. A. The pore dimensions of gramicidin A. Biophys. J. 65, 2455–2460 (1993)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Cowtan, K. D. & Main, P. Improvement of macromolecular electron-density maps by the simultaneous application of real and reciprocal space constraints. Acta Crystallogr. D 49, 148–157 (1993)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Radermacher, M., Wagenknecht, T., Verschoor, A. & Frank, J. Three-dimensional reconstruction from a single-exposure, random conical tilt series applied to the 50S ribosomal subunit of Escherichia coli . J. Microsc. 146, 113–136 (1987)
Shaikh, T. R. et al. SPIDER image processing for single-particle reconstruction of biological macromolecules from electron micrographs. Nature Protocols 3, 1941–1974 (2008)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nature Methods 10, 584–590 (2013)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nature Methods 9, 853–854 (2012)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Voorhees, R. M., Fernández, I. S., Scheres, S. H. & Hegde, R. S. Structure of the mammalian ribosome-Sec61 complex to 3.4 Å resolution. Cell 157, 1632–1643 (2014)
Greber, B. J. et al. The complete structure of the large subunit of the mammalian mitochondrial ribosome. Nature 515, 283–286 10.1038/nature13895 (2014)
Brown, A. et al. Structure of the large ribosomal subunit from human mitochondria. Science 346, 718–722 (2014)
Fernández, I. S., Bai, X. C., Murshudov, G., Scheres, S. H. & Ramakrishnan, V. Initiation of translation by cricket paralysis virus IRES requires its translocation in the ribosome. Cell 157, 823–831 (2014)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nature Methods 11, 63–65 (2014)
Pettersen, E. F. et al. UCSF Chimera – a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007)
Yachdav, G. et al. PredictProtein – an open resource for online prediction of protein structural and functional features. Nucleic Acids Res. 42, W337–W343 (2014)
Bernsel, A., Viklund, H., Hennerdal, A. & Elofsson, A. TOPCONS: consensus prediction of membrane protein topology. Nucleic Acids Res. 37, W465–W468 (2009)
Sonnhammer, E. L., von Heijne, G. & Krogh, A. A hidden Markov model for predicting transmembrane helices in protein sequences. Proc. Int. Conf. Intell. Syst. Mol. Biol. 6, 175–182 (1998)
Tusnády, G. E. & Simon, I. The HMMTOP transmembrane topology prediction server. Bioinformatics 17, 849–850 (2001)
Käll, L., Krogh, A. & Sonnhammer, E. L. Advantages of combined transmembrane topology and signal peptide prediction – the Phobius web server. Nucleic Acids Res. 35, W429–W432 (2007)
Acknowledgements
We thank H. Yu and J. Chai for discussion and proofreading of the manuscript. We thank the staff at beamline BL17U of the Shanghai Synchrotron Radiation Facility (SSRF) and beamline 3W1A of the Beijing Synchrotron Radiation Facility (BSRF) for their assistance in data collection. K. Wu and H. Wang are acknowledged for technique help. We also thank the National Center for Protein Sciences (Beijing, China) for technical support with cryo-electron microscopy data collection and for computation resources. This work was supported by grants from the Ministry of Science and Technology (2012CB911101 and 2011CB910502 to M.Y., 2015CB910102 to B.X. and 2013CB910404 to N.G.), the National Natural Science Foundation of China (21532004, 31570733, 31030020 and 31170679 to M.Y., 31422016 to N.G. and 31422027 to B.X.) and the Ministry of Education (the Young Thousand Talent program to B.X.).
Author information
Authors and Affiliations
Contributions
M.Y. directed the study. J.G., M.C. and R.L. performed protein purification, detergent screening and crystallization. W.L. performed electron microscopy sample preparation, data collection and structural determination with N.L; Q.Z. was responsible for molecular cloning (with P.Z.), protein purification, detergent screening and biochemical and confocal imaging studies. N.G. directed electron microscopy studies and wrote part of the manuscript. B.X. initiated the project and directed molecular cloning, protein expression and purification and wrote most of the manuscript. All authors contributed to discussion of the data and editing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Biochemical characterization of the recombinant protein of Piezo1–pp–GST.
a, A representative trace of gel filtration chromatography of the Piezo1–pp–GST protein. b, Protein samples of the indicated fractions were subjected to SDS–PAGE and Coomassie blue staining. Fractions of 8.0 ml and 8.5 ml (elution volume) were used for the negative-staining electron microscopy and native gel analyses, respectively.
Extended Data Figure 2 Negative-staining electron microscopy examination of Piezo1 in different detergents.
a, A representative micrograph of negatively stained Piezo1 purified with C12E10. b, 2D class averages of Piezo1 particles (C12E10). c, A representative micrograph of negatively stained Piezo1 purified with C12E8. d, 2D class averages of Piezo1 particles (C12E8). e, A representative micrograph of negatively stained Piezo1, with amphipol A8-35 as detergent. f, 2D class averages of Piezo1 particles (amphipol A8-35).
Extended Data Figure 3 Initial model of Piezo1 generated from the random conical tilt method and validation of the model using cryo-electron microscopy data from a TF20 microscope.
a, b, Representative micrographs of negatively stained Piezo1 in C12E10 collected in random conical tilt (RCT) pairs (a, untilted and b, 50° tilted). c, Top view of an RCT reconstruction, showing an overall threefold symmetry for the Piezo1 complex, is shown on the left. The right-hand side shows the top view of the refined model, obtained by a structural refinement of all particles from untilted micrographs. d, e, Model validation was performed by refinement of cryo-electron microscopy particles collected with TF20, with a Gaussian ball (d) or the RCT model (e) as initial reference. The 3D volumes are shown in top, side and bottom views. During the refinement, both the symmetry-free (C1) and symmetry-imposed (C3) reconstructions were tested. Note that some of these reconstructions have incorrect handedness.
Extended Data Figure 4 Representative raw particles of Piezo1 collected with the Titan Krios electron microscope fitted with a K2 electron detector.
A collection of raw particles of Piezo1 (eluted with C12E10), collected with Titan Krios (300 kV).
Extended Data Figure 5 Workflow of 3D classification of Piezo1 particles.
a, The schematic diagram of a series of 3D classification procedures with RELION is shown (also see Methods). After several rounds of 2D classification, the remaining 120,000 particles were subjected to three rounds of 3D classification without imposing any symmetry. A final set of particles (class 4 after the second round of 3D classification), with its reconstruction best matching threefold symmetry, was subjected to 3D refinement (C3 imposed). Notably, further 3D classification of this class resulted in generally similar structures (vertically arranged panels) without detectable improvement of conformational homogeneity. A top view of the soft mask used in structural refinement is also shown (yellow). b, Distribution of particle orientations in the last iteration of the refinement. c, Gold-standard Fourier shell correlation (FSC) curves of the final density map. The FSC curves were calculated with (red) or without (blue) the application of a soft mask to the two half-set maps. The final FSC curve (red) was corrected for the soft-mask-induced effect. Reported resolutions were based on FSC = 0.143 criteria.
Extended Data Figure 6 The trimeric CEDs form the cap domain of Piezo1.
a, A representative micrograph of negatively stained Piezo1(ΔCED) in C12E10. b, 2D class averages of negatively stained Piezo1(ΔCED) particles. It is evident that the central cap domain is absent from these average images. c, Sequence alignment of the CED region of Piezo1 from Mus musculus and Caenorhabditis elegans. Identical residues are highlighted in blue. Secondary structures are indicated by cartoons above the primary sequence. Sequence alignment was performed using Clustal W2 (ref. 71). d, Structure alignment of the trimeric CED of Piezo1 from M. musculus and C. elegans. The three CEDs are coloured in purple, cyan and green, respectively. The CED of C. elegans is coloured in orange. e, A representative trace of gel filtration of the CED of Piezo1. The molecular weights are labelled. Protein samples of the indicated fractions were subjected to SDS–PAGE and Coomassie blue staining (bottom). f, Transparent surface representation of the segmented density map of the cap, superimposed with the trimeric CED crystal structure. The trimeric CEDs are coloured as in d.
Extended Data Figure 7 Local resolution map of the final density map.
a, The final 3D density map of Piezo1 is coloured according to the local resolutions estimated by the software of ResMap. The density map is shown in three different views (top, bottom and side, respectively). b, The final 3D density map (transparent) is superimposed with a poly-alanine model and the crystal structure of the trimeric CED. Three protomers are coloured cyan, purple and green, respectively.
Extended Data Figure 8 Density connections between the transmembrane helices and between the helices in the compact CTD.
a, Alanine models of five representative pairs of transmembrane helices are displayed with their densities (mesh) superimposed. The transmembrane region is highlighted by a light purple shade with the intracellular and extracellular sides indicated. b, An alanine model of the anchor motif with its density superimposed (mesh). Four helices (α1anchor–α4anchor) connecting PH1 and OH are labelled. The transmembrane region is highlighted by a light purple shade with the intracellular and extracellular sides indicated. c, An alanine model of the last four helices (α1CTD–α4CTD) of the trimeric CTD, superimposed with the density of the CTD (mesh).
Extended Data Figure 9 Secondary structure analyses of the C-terminal segments of Piezo1 proteins from different species.
Sequence alignment of the C-terminal regions of Piezo1 from different species. The alignment was performed using Clustal W2 (ref. 71). The anchor motif and the CTD are highlighted in green and pink, respectively. For clarification, the sequences of CEDs were omitted and are indicated by red dashed lines. Secondary structures (α-helices) predicted with PredictProtein72 are shown as black lines. Transmembrane segments were predicted using multiple web servers including Topcons73 (green lines), TMHMM2 (ref. 74) (blue lines), HMMTOP75 (orange lines) and Phobius76 (pink lines).
Rights and permissions
About this article
Cite this article
Ge, J., Li, W., Zhao, Q. et al. Architecture of the mammalian mechanosensitive Piezo1 channel. Nature 527, 64–69 (2015). https://doi.org/10.1038/nature15247
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15247
This article is cited by
-
Piezo1 expression in chondrocytes controls endochondral ossification and osteoarthritis development
Bone Research (2024)
-
Piezo1 regulates meningeal lymphatic vessel drainage and alleviates excessive CSF accumulation
Nature Neuroscience (2024)
-
Advances in using ultrasound to regulate the nervous system
Neurological Sciences (2024)
-
The Piezo channel is a mechano-sensitive complex component in the mammalian inner ear hair cell
Nature Communications (2024)
-
Intraluminal chloride regulates lung branching morphogenesis: involvement of PIEZO1/PIEZO2
Respiratory Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.