Abstract
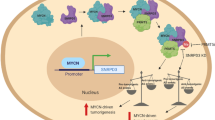
MYC (also known as c-MYC) overexpression or hyperactivation is one of the most common drivers of human cancer. Despite intensive study, the MYC oncogene remains recalcitrant to therapeutic inhibition. MYC is a transcription factor, and many of its pro-tumorigenic functions have been attributed to its ability to regulate gene expression programs1,2,3. Notably, oncogenic MYC activation has also been shown to increase total RNA and protein production in many tissue and disease contexts4,5,6,7. While such increases in RNA and protein production may endow cancer cells with pro-tumour hallmarks, this increase in synthesis may also generate new or heightened burden on MYC-driven cancer cells to process these macromolecules properly8. Here we discover that the spliceosome is a new target of oncogenic stress in MYC-driven cancers. We identify BUD31 as a MYC-synthetic lethal gene in human mammary epithelial cells, and demonstrate that BUD31 is a component of the core spliceosome required for its assembly and catalytic activity. Core spliceosomal factors (such as SF3B1 and U2AF1) associated with BUD31 are also required to tolerate oncogenic MYC. Notably, MYC hyperactivation induces an increase in total precursor messenger RNA synthesis, suggesting an increased burden on the core spliceosome to process pre-mRNA. In contrast to normal cells, partial inhibition of the spliceosome in MYC-hyperactivated cells leads to global intron retention, widespread defects in pre-mRNA maturation, and deregulation of many essential cell processes. Notably, genetic or pharmacological inhibition of the spliceosome in vivo impairs survival, tumorigenicity and metastatic proclivity of MYC-dependent breast cancers. Collectively, these data suggest that oncogenic MYC confers a collateral stress on splicing, and that components of the spliceosome may be therapeutic entry points for aggressive MYC-driven cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Eilers, M. & Eisenman, R. N. Myc’s broad reach. Genes Dev. 22, 2755–2766 (2008)
Sabo, A. & Amati, B. Genome recognition by MYC. Cold Spring Harb. Perspect. Med. 4, a014191 (2014)
Dang, C. V. MYC, metabolism, cell growth, and tumorigenesis. Cold Spring Harb. Perspect. Med. 3, a014217 (2013)
Lin, C. Y. et al. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 151, 56–67 (2012)
Nie, Z. et al. c-Myc is a universal amplifier of expressed genes in lymphocytes and embryonic stem cells. Cell 151, 68–79 (2012)
Ruggero, D. The role of Myc-induced protein synthesis in cancer. Cancer Res. 69, 8839–8843 (2009)
Barna, M. et al. Suppression of Myc oncogenic activity by ribosomal protein haploinsufficiency. Nature 456, 971–975 (2008)
Luo, J., Solimini, N. L. & Elledge, S. J. Principles of cancer therapy: oncogene and non-oncogene addiction. Cell 136, 823–837 (2009)
Kessler, J. D. et al. A SUMOylation-dependent transcriptional subprogram is required for Myc-driven tumorigenesis. Science 335, 348–353 (2012)
Masciadri, B. et al. Characterization of the BUD31 gene of Saccharomyces cerevisiae. Biochem. Biophys. Res. Commun. 320, 1342–1350 (2004)
Wahl, M. C., Will, C. L. & Luhrmann, R. The spliceosome: design principles of a dynamic RNP machine. Cell 136, 701–718 (2009)
Bonnal, S., Vigevani, L. & Valcarcel, J. The spliceosome as a target of novel antitumour drugs. Nature Rev. Drug Discov. 11, 847–859 (2012)
Lagisetti, C. et al. Optimization of antitumor modulators of pre-mRNA splicing. J. Med. Chem. 56, 10033–10044 (2013)
Kanazawa, S., Soucek, L., Evan, G., Okamoto, T. & Peterlin, B. M. c-Myc recruits P-TEFb for transcription, cellular proliferation and apoptosis. Oncogene 22, 5707–5711 (2003)
Rahl, P. B. et al. c-Myc regulates transcriptional pause release. Cell 141, 432–445 (2010)
Koh, C. M. et al. MYC regulates the core pre-mRNA splicing machinery as an essential step in lymphomagenesis. Nature 523, 96–100 (2015)
Garneau, N. L., Wilusz, J. & Wilusz, C. J. The highways and byways of mRNA decay. Nature Rev. Mol. Cell Biol. 8, 113–126 (2007)
Mezquita, P., Parghi, S. S., Brandvold, K. A. & Ruddell, A. Myc regulates VEGF production in B cells by stimulating initiation of VEGF mRNA translation. Oncogene 24, 889–901 (2005)
Minn, A. J. et al. Distinct organ-specific metastatic potential of individual breast cancer cells and primary tumors. J. Clin. Invest. 115, 44–55 (2005)
Di Cosimo, S. & Baselga, J. Management of breast cancer with targeted agents: importance of heterogeneity. Nature Rev. Clin. Oncol. 7, 139–147 (2010)
David, C. J., Chen, M., Assanah, M., Canoll, P. & Manley, J. L. HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature 463, 364–368 (2010)
Sabò, A. et al. Selective transcriptional regulation by Myc in cellular growth control and lymphomagenesis. Nature 511, 488–492 (2014)
Walz, S. et al. Activation and repression by oncogenic MYC shape tumour-specific gene expression profiles. Nature 511, 483–487 (2014)
Lin, C. J. et al. Targeting synthetic lethal interactions between Myc and the eIF4F complex impedes tumorigenesis. Cell Rep. 1, 325–333 (2012)
Graubert, T. A. et al. Recurrent mutations in the U2AF1 splicing factor in myelodysplastic syndromes. Nature Genet. 44, 53–57 (2011)
Papaemmanuil, E. et al. Somatic SF3B1 mutation in myelodysplasia with ring sideroblasts. N. Engl. J. Med. 365, 1384–1395 (2011)
Hubert, C. G. et al. Genome-wide RNAi screens in human brain tumor isolates reveal a novel viability requirement for PHF5A. Genes Dev. 27, 1032–1045 (2013)
Adler, A. S. et al. An integrative analysis of colon cancer identifies an essential function for PRPF6 in tumor growth. Genes Dev. 28, 1068–1084 (2014)
Cunningham, J. T., Moreno, M. V., Lodi, A., Ronen, S. M. & Ruggero, D. Protein and nucleotide biosynthesis are coupled by a single rate-limiting enzyme, PRPS2, to drive cancer. Cell 157, 1088–1103 (2014)
Liu, Y. C. et al. Global regulation of nucleotide biosynthetic genes by c-Myc. PLoS ONE 3, e2722 (2008)
Meerbrey, K. L. et al. The pINDUCER lentiviral toolkit for inducible RNA interference in vitro and in vivo. Proc. Natl Acad. Sci. USA 108, 3665–3670 (2011)
Schuhmacher, M. et al. Control of cell growth by c-Myc in the absence of cell division. Curr. Biol. 9, 1255–1258 (1999)
Malovannaya, A. et al. Streamlined analysis schema for high-throughput identification of endogenous protein complexes. Proc. Natl Acad. Sci. USA 107, 2431–2436 (2010)
Zapp, M. L. & Berget, S. M. Evidence for nuclear factors involved in recognition of 5′ splice sites. Nucleic Acids Res. 17, 2655–2674 (1989)
Dignam, J. D., Lebovitz, R. M. & Roeder, R. G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11, 1475–1489 (1983)
Echeverria, G. V. & Cooper, T. A. Muscleblind-like 1 activates insulin receptor exon 11 inclusion by enhancing U2AF65 binding and splicing of the upstream intron. Nucleic Acids Res. 42, 1893–1903 (2014)
Dölken, L. et al. High-resolution gene expression profiling for simultaneous kinetic parameter analysis of RNA synthesis and decay. RNA 14, 1959–1972 (2008)
Marcotte, R. et al. Essential gene profiles in breast, pancreatic, and ovarian cancer cells. Cancer Discov. 2, 172–189 (2012)
Shao, D. D. et al. ATARiS: computational quantification of gene suppression phenotypes from multisample RNAi screens. Genome Res. 23, 665–678 (2013)
Acknowledgements
We would like to thank J. Rosen, S. Butler, K. Neugebauer, M. Moore, S. Elledge, T. Davoli, members of T.F.W., C.A.S. and T.A.C. laboratories for comments, and P. Yu for bioinformatics support. The authors also acknowledge the joint participation by Adrienne Helis Melvin Medical Research Foundation through its direct engagement in the continuous active conduct of medical research in conjunction with Baylor College of Medicine for cancer research. The Dan L. Duncan Cancer Center Shared Resources was supported by the NCI P30CA125123 Center Grant and provided technical assistance including Cell-Based Assay Screening Service (D. Liu), Genomic and RNA Profiling Resource (L. White), Biostatistics & Informatics Shared Resource (S. Hilsenbeck), Cytometry and Cell Sorting (J. Sederstrom; P30 AI036211 and S10 RR024574), and the Proteomics and Metabolomics Core Facility (Cancer Prevention and Research Institute of Texas, RP12009). T.Y.-T.H. was supported by NIH pre-doctoral fellowship (NCI 1F30CA180447) and CPRIT training grant (RP101499). M.O. and R.J.B. were supported by The Gillson Longenbaugh Foundation. R.J.B. was supported by Alex’s Lemonade Stand Foundation. T.F.W. was supported by CPRIT (RP120583), the Susan G. Komen for the Cure (KG090355), the NIH (1R01CA178039-01 and U54- CA149196) and the DOD Breast Cancer Research Program (BC120604).
Author information
Authors and Affiliations
Contributions
T.Y.-T.H., N.J.N., R.M., C.S.B., G.V.E., T.S., S.J.K., S.T., K.L.K., J.D.H., K.S., R.J.B., S.O.S., A.J., C.L. and M.O. performed the experiments. L.M.S., A.S., R.D.-V., A.R. and C.A.S. performed statistical analyses. I.G., S.Y.J., J.R.N., X.H.-F.Z., T.A.C., T.R.W., B.G.N., C.A.S. and T.F.W. devised or supervised experiments. T.Y.-T.H. and T.F.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Validation of BUD31 as a MYC-synthetic lethal gene in HMECs.
a, qRT–PCR analysis of BUD31 mRNA level (mean ± s.d., n = 3 biological replicates). b, Clonogenicity of MYC-ER HMECs with or without MYC hyperactivation or BUD31 depletion (mean ± s.e.m., n = 4 biological replicates, **P < 0.01, two-tailed Student’s t-test). c, Caspase-3/7 activation by caspase luminescence assay (mean ± s.e.m., n = 3, ***P < 0.001, one-way ANOVA). d, Flag-tagged protein levels in MYC-ER HMECs in which vinculin was used as a loading control.
Extended Data Figure 2 BUD31 interacts with core spliceosomal factors and is required for spliceosomal assembly and pre-mRNA splicing.
a, 134 core spliceosomal proteins are listed. Proteins in red are shown to interact with BUD31, as discovered by Flag–BUD31 immunoprecipitation mass spectrometry and BUD31 BiFC. b, Heat map of BUD31-interacting spliceosomal proteins, organized by spliceosome sub-complexes. A black-green colour scale depicts normalized BiFC interaction values between spliceosomal proteins and negative control protein (technical replicates in two left lanes) and BUD31 (technical replicates in two right lanes). c, Spliceosomal snRNPs (coloured circles) interact in a stepwise manner to excise intronic sequences from pre-mRNA. snRNPs with proteins identified from the BUD31 immunoprecipitation and mass spectrometry are noted (blue outline) to be BUD31-associated. d, Co-immunoprecipitation of Flag–BUD31 for non-spliceosomal proteins. Input and immunoprecipitation blots probed by EIF2S1 and EIF3I were taken at different exposures to minimize background signal. e, Interaction between N-YFP-tagged BUD31 and C-YFP-tagged spliceosomal (DDX46) or cytoplasmic proteins (TRIM9, SOCS2 and EPHA8) was assessed by cellular fluorescence (mean ± s.e.m., n = 3 technical replicates). f, Nuclear extracts with or without BUD31 knockdown were incubated with pre-mRNA substrate, and RT–PCR of unspliced RNA (top) and spliced RNA (bottom) was performed, using primers at the indicated arrows (left). BUD31 protein levels in the nuclear extracts were normalized to vinculin expression (middle) and quantified (right). g, Radioactively labelled pre-mRNA (MINX) was incubated with nuclear extracts with or without BUD31 depletion. RNA purified from the splicing reaction was run on a denaturing gel and imaged by autoradiography. The identities of prominent bands are based on size. Asterisk denotes putative intron-lariat band. h, After in vitro splicing was performed as described previously, products were electrophoresed on native gel, and spliceosome complexes were visualized by autoradiography. Complex A and nonspecific H complexes are labelled. i, Phosphorimager quantification of the ratio of RNA in complex A compared to that in complex H. j, Interaction between N-YFP-tagged wild-type (WT) or mutant BUD31 and C-YFP-tagged splicing factors was assessed by cellular fluorescence (mean ± s.e.m., n = 2 technical replicates, ***P < 0.001, two-tailed Student’s t-test).
Extended Data Figure 3 HMECs with oncogenic activation of HER2 and EGFR do not require BUD31.
a, Cell number changes in HMECs with inducible shBUD31 and constitutive HER2 or EGFR expression (mean ± s.e.m.; n = 4 technical replicates; *P < 0.05, two-tailed Student’s t-test). HER2 and EGFR protein is normalized to vinculin (right). b, MYC protein levels in HMECs with constitutive HER2 or EGFR expression. c, MYC induction by tamoxifen in MYC-ER HMECs does not increase cell proliferation over time (mean ± s.e.m., n = 8 technical replicates).
Extended Data Figure 4 Partial knockdown of core splicing factors is MYC-synthetic lethal in HMECs.
a–d, mRNA levels for core splicing factors SF3B1 (a), U2AF1 (b), EFTUD2 (c) and SNRPF (d) were evaluated by qRT–PCR (mean ± s.d., n = 3 technical replicates). e–i, Caspase-3/7 luminescence in MYC-ER HMECs with partial suppression of core spliceosomal proteins (e–h) or spliceosome inhibitor SD6 (i) (mean ± s.e.m., n = 3 technical replicates, ***P < 0.001, one-way ANOVA).
Extended Data Figure 5 BUD31 loss in MYC-hyperactivated cells destabilizes mRNA.
a, b, MYC-ER HMECs with inducible shBUD31 treated with actinomycin D for 5 h were labelled with oligo(dT)25 LNA probes via fluorescence in situ hybridization. Cellular FITC intensity was assessed within cellular (a) and nuclear (DAPI+) (b) regions. Data are represented as the difference in cellular FITC intensity between 0 and 5 h of actinomycin D treatment in each cell state (mean ± s.e.m., n = 150, ***P < 0.001, two-tailed Student’s t-test).
Extended Data Figure 6 BUD31 depletion in MYC-hyperactivated cells enhances intron retention and decreases expression of cell-essential genes.
In MYC-hyperactive cells, 17 representative genes display increased IR and decreased steady-state RNA levels after BUD31 knockdown. Depletion of these genes by shRNA decreased cell viability (mean barcode abundance ± s.e.m.). Twofold decrease in barcode abundance is noted by the dashed red line. All values are reflective of three biological replicates, and genes are colour-coded based on their Gene Ontology term annotation.
Extended Data Figure 7 MYC-dependent breast cancer cells require BUD31 for in vitro and in vivo growth.
a, Relative cell number of SUM159 cells with doxycycline-inducible shBUD31 in vitro (mean ± s.e.m., n = 8 technical replicates, ***P < 0.001, two-tailed Student’s t-test). b, Caspase-3/7 luminescence in BUD31-depleted SUM159 cells (mean ± s.e.m., n = 3 technical replicates, ***P < 0.001, two-tailed Student’s t-test). c, d, SUM159 cells engineered with dox-inducible shBUD31 were subcutaneously transplanted into mice and randomized onto dox treatment (−dox n = 10, +dox n = 9). Loss of BUD31 in SUM159 xenografts inhibits tumour growth (mean ± s.e.m., ***P < 0.001 at day 21, two-tailed Student’s t-test) (c) and prolongs progression-free survival (d) in nude mice (P-value, log-rank test).
Extended Data Figure 8 BUD31 depletion does not affect levels of MYC protein.
a, MYC protein levels in MYC-ER HMECs with inducible shBUD31 expression normalized to vinculin expression. To confirm specificity of MYC antibody, HMECs without the MYC-ER construct were engineered to express inducible MYC shRNA. b, MYC protein levels in SUM159 and LM2 cells with inducible shBUD31 normalized to vinculin expression. To confirm specificity of MYC antibody, SUM159 cells were engineered to express inducible MYC shRNA.
Extended Data Figure 9 Schematic for in vivo barcode-based competition assay.
LM2 cells transduced with inducible shRNAs targeting negative control genes or candidate genes were mixed at an equal ratio. This mixed population was transplanted into mice, and tumours were allowed to form in the presence or absence of dox. At the experimental endpoint, genomic DNA was isolated for comparisons of relative barcode (shRNA) abundance in tumour genomic DNA.
Extended Data Figure 10 Spliceosome inhibitor SD6 inhibits MYC-dependent cancer cells in vitro and in vivo.
a, MYC-dependent breast cancer cells (SUM159 and LM2) and MYC-normal immortalized epithelial cells (F7 and HME1) were cultured with SD6 at low density and analysed for clonogenic growth. b, MYC-repressible human B-cell line P493-6 was treated with or without 100 nM SD6 in the absence or presence of MYC hyperactivation for four days, and cells were counted for relative cell number changes (mean ± s.e.m., n = 3 biological replicates, ***P < 0.001, one-way ANOVA). c, Kaplan–Meier survival analysis of nude mice with pulmonary seeding of LM2 cells treated with or without SD6 for 10 days (vehicle n = 7, SD6 n = 6, P-value by log-rank test).
Rights and permissions
About this article
Cite this article
Hsu, TT., Simon, L., Neill, N. et al. The spliceosome is a therapeutic vulnerability in MYC-driven cancer. Nature 525, 384–388 (2015). https://doi.org/10.1038/nature14985
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14985
This article is cited by
-
MYCN and SNRPD3 cooperate to maintain a balance of alternative splicing events that drives neuroblastoma progression
Oncogene (2024)
-
Cuproptosis engages in c-Myc-mediated breast cancer stemness
Journal of Translational Medicine (2023)
-
MYC up-regulation confers vulnerability to dual inhibition of CDK12 and CDK13 in high-risk Group 3 medulloblastoma
Journal of Experimental & Clinical Cancer Research (2023)
-
Tumor educated platelet: the novel BioSource for cancer detection
Cancer Cell International (2023)
-
Bud31-mediated alternative splicing is required for spermatogonial stem cell self-renewal and differentiation
Cell Death & Differentiation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.