Abstract
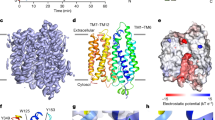
The altered activity of the fructose transporter GLUT5, an isoform of the facilitated-diffusion glucose transporter family, has been linked to disorders such as type 2 diabetes and obesity. GLUT5 is also overexpressed in certain tumour cells, and inhibitors are potential drugs for these conditions. Here we describe the crystal structures of GLUT5 from Rattus norvegicus and Bos taurus in open outward- and open inward-facing conformations, respectively. GLUT5 has a major facilitator superfamily fold like other homologous monosaccharide transporters. On the basis of a comparison of the inward-facing structures of GLUT5 and human GLUT1, a ubiquitous glucose transporter, we show that a single point mutation is enough to switch the substrate-binding preference of GLUT5 from fructose to glucose. A comparison of the substrate-free structures of GLUT5 with occluded substrate-bound structures of Escherichia coli XylE suggests that, in addition to global rocker-switch-like re-orientation of the bundles, local asymmetric rearrangements of carboxy-terminal transmembrane bundle helices TM7 and TM10 underlie a ‘gated-pore’ transport mechanism in such monosaccharide transporters.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mueckler, M. & Thorens, B. The SLC2 (GLUT) family of membrane transporters. Mol. Aspects Med. 34, 121–138 (2013)
Zhao, F. Q. & Keating, A. F. Functional properties and genomics of glucose transporters. Curr. Genomics 8, 113–128 (2007)
Simpson, I. A., Vannucci, S. J. & Maher, F. Glucose transporters in mammalian brain. Biochem. Soc. Trans. 22, 671–675 (1994)
James, D. E., Strube, M. & Mueckler, M. Molecular cloning and characterization of an insulin-regulatable glucose transporter. Nature 338, 83–87 (1989)
Burant, C. F., Takeda, J., Brot-Laroche, E., Bell, G. I. & Davidson, N. O. Fructose transporter in human spermatozoa and small intestine is GLUT5. J. Biol. Chem. 267, 14523–14526 (1992)
Kayano, T. et al. Human facilitative glucose transporters. Isolation, functional characterization, and gene localization of cDNAs encoding an isoform (GLUT5) expressed in small intestine, kidney, muscle, and adipose tissue and an unusual glucose transporter pseudogene-like sequence (GLUT6). J. Biol. Chem. 265, 13276–13282 (1990)
Blakemore, S. J. et al. The GLUT5 hexose transporter is also localized to the basolateral membrane of the human jejunum. Biochem. J. 309, 7–12 (1995)
Douard, V. & Ferraris, R. P. Regulation of the fructose transporter GLUT5 in health and disease. Am. J. Physiol. Endocrinol. Metab. 295, E227–E237 (2008)
Rand, E. B., Depaoli, A. M., Davidson, N. O., Bell, G. I. & Burant, C. F. Sequence, tissue distribution, and functional characterization of the rat fructose transporter GLUT5. Am. J. Physiol. 264, G1169–G1176 (1993)
Shepherd, P. R., Gibbs, E. M., Wesslau, C., Gould, G. W. & Kahn, B. B. Human small intestine facilitative fructose/glucose transporter (GLUT5) is also present in insulin-responsive tissues and brain. Investigation of biochemical characteristics and translocation. Diabetes 41, 1360–1365 (1992)
Zisman, A. et al. Targeted disruption of the glucose transporter 4 selectively in muscle causes insulin resistance and glucose intolerance. Nature Med. 6, 924–928 (2000)
Barone, S. et al. Slc2a5 (Glut5) is essential for the absorption of fructose in the intestine and generation of fructose-induced hypertension. J. Biol. Chem. 284, 5056–5066 (2009)
Douard, V. & Ferraris, R. P. The role of fructose transporters in diseases linked to excessive fructose intake. J. Physiol. 591, 401–414 (2013)
Zamora-Leon, S. P. et al. Expression of the fructose transporter GLUT5 in human breast cancer. Proc. Natl Acad. Sci. USA 93, 1847–1852 (1996)
Warburg, O. On respiratory impairment in cancer cells. Science 124, 269–270 (1956)
Yan, N. Structural advances for the major facilitator superfamily (MFS) transporters. Trends Biochem. Sci. 38, 151–159 (2013)
Madej, M. G., Sun, L., Yan, N. & Kaback, H. R. Functional architecture of MFS D-glucose transporters. Proc. Natl Acad. Sci. USA 111, E719–E727 (2014)
Maiden, M. C., Davis, E. O., Baldwin, S. A., Moore, D. C. & Henderson, P. J. Mammalian and bacterial sugar transport proteins are homologous. Nature 325, 641–643 (1987)
Pao, S. S., Paulsen, I. T. & Saier, M. H., Jr Major facilitator superfamily. Microbiol. Mol. Biol. Rev. 62, 1–34 (1998)
Farwick, A., Bruder, S., Schadeweg, V., Oreb, M. & Boles, E. Engineering of yeast hexose transporters to transport D-xylose without inhibition by D-glucose. Proc. Natl Acad. Sci. USA 111, 5159–5164 (2014)
Deng, D. et al. Crystal structure of the human glucose transporter GLUT1. Nature 510, 121–125 (2014)
Sun, L. et al. Crystal structure of a bacterial homologue of glucose transporters GLUT1–4. Nature 490, 361–366 (2012)
Quistgaard, E. M., Low, C., Moberg, P., Tresaugues, L. & Nordlund, P. Structural basis for substrate transport in the GLUT-homology family of monosaccharide transporters. Nature Struct. Mol. Biol. 20, 766–768 (2013)
Wisedchaisri, G., Park, M. S., Iadanza, M. G., Zheng, H. & Gonen, T. Proton-coupled sugar transport in the prototypical major facilitator superfamily protein XylE. Nat. Commun. 5, 4521 (2014)
Iancu, C. V., Zamoon, J., Woo, S. B., Aleshin, A. & Choe, J. Y. Crystal structure of a glucose/H+ symporter and its mechanism of action. Proc. Natl Acad. Sci. USA 110, 17862–17867 (2013)
Garcia, J. C., Strube, M., Leingang, K., Keller, K. & Mueckler, M. M. Amino acid substitutions at tryptophan 388 and tryptophan 412 of the HepG2 (Glut1) glucose transporter inhibit transport activity and targeting to the plasma membrane in Xenopus oocytes. J. Biol. Chem. 267, 7770–7776 (1992)
Li, Q. et al. Cloning and functional characterization of the human GLUT7 isoform SLC2A7 from the small intestine. Am. J. Physiol. Gastrointest. Liver Physiol. 287, G236–G242 (2004)
Mueckler, M. & Makepeace, C. Analysis of transmembrane segment 10 of the Glut1 glucose transporter by cysteine-scanning mutagenesis and substituted cysteine accessibility. J. Biol. Chem. 277, 3498–3503 (2002)
Andersson, M. et al. Proton-coupled dynamics in lactose permease. Structure 20, 1893–1904 (2012)
Law, C. J. et al. Salt-bridge dynamics control substrate-induced conformational change in the membrane transporter GlpT. J. Mol. Biol. 378, 828–839 (2008)
Schürmann, A. et al. Role of conserved arginine and glutamate residues on the cytosolic surface of glucose transporters for transporter function. Biochemistry 36, 12897–12902 (1997)
Seatter, M. J., De la Rue, S. A., Porter, L. M. & Gould, G. W. QLS motif in transmembrane helix VII of the glucose transporter family interacts with the C-1 position of D-glucose and is involved in substrate selection at the exofacial binding site. Biochemistry 37, 1322–1326 (1998)
Hruz, P. W. & Mueckler, M. M. Cysteine-scanning mutagenesis of transmembrane segment 7 of the GLUT1 glucose transporter. J. Biol. Chem. 274, 36176–36180 (1999)
Manolescu, A., Salas-Burgos, A. M., Fischbarg, J. & Cheeseman, C. I. Identification of a hydrophobic residue as a key determinant of fructose transport by the facilitative hexose transporter SLC2A7 (GLUT7). J. Biol. Chem. 280, 42978–42983 (2005)
Kasahara, T., Maeda, M., Boles, E. & Kasahara, M. Identification of a key residue determining substrate affinity in the human glucose transporter GLUT1. Biochim. Biophys. Acta 1788, 1051–1055 (2009)
Karpowich, N. K. & Wang, D. N. Structural biology. Symmetric transporters for asymmetric transport. Science 321, 781–782 (2008)
Radestock, S. & Forrest, L. R. The alternating-access mechanism of MFS transporters arises from inverted-topology repeats. J. Mol. Biol. 407, 698–715 (2011)
Solcan, N. et al. Alternating access mechanism in the POT family of oligopeptide transporters. EMBO J. 31, 3411–3421 (2012)
Fukuda, M. et al. Structural basis for dynamic mechanism of nitrate/nitrite antiport by NarK. Nature Commun. 6, 7097 (2015)
Kota, J., Gilstring, C. F. & Ljungdahl, P. O. Membrane chaperone Shr3 assists in folding amino acid permeases preventing precocious ERAD. J. Cell Biol. 176, 617–628 (2007)
Newstead, S., Kim, H., von Heijne, G., Iwata, S. & Drew, D. High-throughput fluorescent-based optimization of eukaryotic membrane protein overexpression and purification in Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 104, 13936–13941 (2007)
Drew, D. et al. GFP-based optimization scheme for the overexpression and purification of eukaryotic membrane proteins in Saccharomyces cerevisiae. Nature Protocols 3, 784–798 (2008)
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006)
Sonoda, Y. et al. Benchmarking membrane protein detergent stability for improving throughput of high-resolution X-ray structures. Structure 19, 17–25 (2011)
Suharni, et al. Proteoliposome-based selection of a recombinant antibody fragment against the human M2 muscarinic acetylcholine receptor. Monoclon. Antib. Immunodiagn. Immunother. 33, 378–385 (2014)
Sonoda, Y. et al. Tricks of the trade used to accelerate high-resolution structure determination of membrane proteins. FEBS Lett. 584, 2539–2547 (2010)
Sarkar, H. K., Thorens, B., Lodish, H. F. & Kaback, H. R. Expression of the human erythrocyte glucose transporter in Escherichia coli. Proc. Natl Acad. Sci. USA 85, 5463–5467 (1988)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D. 50, 760–763 (1994)
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D 66, 22–25 (2010)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Blanc, E. et al. Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr. D 60, 2210–2221 (2004)
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr. Protocols Protein Sci. Ch. 2, Unit 2.9. (2007)
Notredame, C., Higgins, D. G. & Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302, 205–217 (2000)
Acknowledgements
We are grateful to D. Slotboom, A. Cameron and S. Newstead for discussions and comments, and J. Mansfield for assistance with large-scale yeast fermentations and H. Unno with rGLUT5 crystallization. Data were collected at the European Synchrotron Radiation Facility, Diamond Light Source, and SPring-8 (proposal numbers 2011A1393, 2011B1229, 2012A1184, 2012B1253, 2013A1241, 2013B1237, 2014A1348 and 2014B1407), with assistance from beamline scientists. This work was funded by the Knut and Alice Wallenberg Foundation (D.D), The Royal Society through the University Research Fellow scheme (D.D), the BBSRC (BB/G02325/1 to S.I.), the ERATO Human Receptor Crystallography Project of the Japan Science and Technology Agency (JST) (S.I.), by the Research Acceleration Program of the JST (S.I.), by the Targeted Proteins Research Program of the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan (S.I.), and by Grants-in-Aids for Scientific Research from the MEXT (No. 22570114 to N.N.), and by the Platform for Drug Discovery, Informatics, and Structural Life Science from the MEXT (T.K.). The authors are grateful for the use of the Membrane Protein Laboratory funded by the Wellcome Trust (grant 062164/Z/00/Z) at the Diamond Light Source Limited, and The Centre for Biomembrane Research (CBR), supported by the Swedish Foundation for Strategic Research. H.J.K. was a recipient of a Human Frontiers Postdoctoral fellowship and D.D. acknowledges support from EMBO through the Young Investigator Program (YIP).
Author information
Authors and Affiliations
Contributions
N.N., S.I. and D.D. designed the project. Cloning, expression screening and initial crystallization of rat and bovine GLUT5 was carried out by H.J.K., Y.So. and D.D. Crystal optimization of bovine GLUT5 was carried out by H.J.K. and G.V. Data collection, structure determination and refinement of bovine GLUT5 was carried out by G.V. Generation of rat GLUT5 scFv fragment was carried out by N.N., Y.N., T.M., Y.N.-N., O.K.-A., H.I., T.A., T.K. and T.Ha. Expression and purification of the Fv fragment was carried out by N.N., Y.N., Y.Sa., H.A. and T.Hi. Co-crystallization of rat GLUT5–Fv complex and data collection was performed by N.N. and Y.N. with assistance from T.Hi. and S.I. Structure determination and refinement of rat GLUT5–Fv was carried out by T.S. Experiments for functional analysis were designed by M.K. and D.D. and carried out by M.K., D.D., S.A.H. and A.A.Q. Modelling of GLUT5 was carried out by M.C. The manuscript was prepared by N.N., H.J.K., G.V., S.I. and D.D. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Anisotropy descriptors of bGLUT5 data reported by the UCLA-MBI Diffraction Anisotropy Server and 2Fo − Fc electron density maps for the bovine and rat GLUT5 structures.
a, Degree of anisotropy of bGLUT5 data, resolution limits for the 3 principal axes (left), and panel illustrating steps along correction of bGLUT5 data for anisotropy (right). b, Representative portions of the electron density map (1.5σ) for bGLUT5 overall model (left) and a close-up of the substrate binding site (right); residues highlighted are numbered based on rGLUT5 for the sake of clarity. c, Electron density (1.0σ) for rGLUT5 showing one of the inter-bundle salt-bridge clusters that form in the open outward-facing conformation.
Extended Data Figure 2 Superimposition of open inward-facing bGLUT5 and hGLUT1 structures, and comparison of the substrate-binding site in bGLUT5 and inward-facing XylE
a, Ribbon representation of inward-facing bGLUT5 (coloured as in Fig. 1a) and inward-facing hGLUT1 (light grey) structures, as viewed in the plane of the membrane. The d-glucopyranoside moiety of the detergent molecule bound to GLUT1 (n-nonyl-β-d-glucopyranoside (β-NG)) is shown in stick representation. Density for ICH5 at the C terminus is missing in both hGLUT1 and bGLUT5 inward-facing structures and highlighted with the dotted ellipse. The beginning of TM1 kinks further outwards in the bGLUT5 structure compared to hGLUT1 and residues 1–18 could not be built. The r.m.s.d. (root mean square deviation) after superposition of the two structures is 1.12 Å for 364 pairs of Cα atoms (see Methods). b, The substrate-binding in the inward-facing bGLUT5 structure (coloured as in Fig. 1) is very similar to that seen in inward-facing XylE (4JA4) structure (shown in light grey). Only non-conserved residues and the equivalent glutamine to Q166 are labelled for XylE.
Extended Data Figure 3 Structure of the rat GLUT5–Fv complex.
a, Cartoon representation of the complex between rGLUT5 (grey) and 4D111Fv (heavy-chain variable region (VH) is in blue; light-chain variable region (VL) is in red). 4D111Fv binds to the cytoplasmic domain of GLUT5, including ICH2 (residues 226, 230, 234), the loop between ICH2 and ICH3 (residues 238, 240, 241), and ICH3 (residue 243), with ∼ 848 Å2 of buried surface area at the interface. b, Packing of the rat GLUT5–Fv complex molecules in the crystal. The unit cell is represented as green lines.
Extended Data Figure 4 Sequence alignment of rat GLUT5 (rGLUT5), bovine GLUT5 (bGLUT5), human GLUT5 and GLUT7 (hGLUT5 and hGLUT7), human GLUT1–4 (hGLUT1–4), Saccharomyces cerevisiae HXT7, Plasmodium falciparum PfHT), Arabidopsis thaliana GlcT and Escherichia coli XylE.
Structure elements of rat GLUT5 are indicated above the alignment, and coloured as in Fig. 1a. Strictly conserved residues are highlighted in black-filled boxes, and highly conserved residues are shaded in grey. Green boxes highlight central cavity residues that are specific to GLUT5 and red boxes highlight those that are conserved among GLUTs. Purple boxes highlight residues forming the salt bridges between cytosolic TM segments. A blue box (TM5) highlights Gln166, whose mutation to glutamic acid, as present in GLUT7, weakens d-fructose binding but supports strong d-glucose binding in rGLUT5. The brown box (TM8) highlights Glu336 that is conserved across all the GLUTs and replaced with glutamic acid in XylE. Red bars underneath the alignment indicate the sugar porter (SP) family motifs18,19. Note that because bGLUT5 and hGLUT5 have an additional amino acid at position 8, their numbering differs from rGLUT5 by 1 amino acid. For clarity, bGLUT5 residues are labelled using rGLUT5 numbering.
Extended Data Figure 5 d-fructose binding monitored by tryptophan fluorescence quenching.
a, Cartoon representation of the outward-facing rGLUT5 structure, as viewed from the plane of the membrane with the colouring as shown in Fig. 1a. Atoms in all tryptophan residues are shown as spheres and tryptophan W419, whose fluorescence is quenched by substrate, is labelled. b, Emission fluorescence spectra for purified deglycosylated rGLUT5 wild-type-like mutant N50Y (referred to as WT), shown in the range of 320–360 nm with an excitation wavelength of 295 nm after the addition of 40 mM d-fructose (top), and 40 mM l-fructose (bottom). Emission fluorescence spectra for purified wild-type protein that had been previously incubated with the inhibitor HgCl2 is also shown for d-fructose (middle). c, Tryptophan fluorescence quenching (excitation 295 nm; emission 338 nm) after incubation of purified rGLUT5 N50Y with either 40 mM d-fructose (filled bar) or l-fructose, d-glucose, d-mannose, d-xylose or d-galactose as labelled (open bars). Tryptophan fluorescence quenching for purified wild-type protein that had been previously incubated with the inhibitor HgCl2 is also shown for d-fructose (open bar). d, As in c, but for rGLUT5 with a single tryptophan residue (W419), which contains the following mutations: N50Y, W70F, W191F, W239F, W265F, W275F, W338F and W370F. No tryptophan quenching was observed for d-fructose (5 mM HgCl2), l-fructose, d-glucose or d-galactose. In all experiments errors bars indicate s.e.m.; n = 3.
Extended Data Figure 6 Substrate specificity in GLUT5.
a, Time-dependent uptake of d-[14C]-fructose by rGLUT5 wild type (open squares and triangles) and the deglycosylated mutant N50Y (filled squares and triangles) in proteoliposomes incubated with or without the inhibitor HgCl2 as labelled. Non-specific uptake was estimated with 0.1 mM l-[14C]-glucose for wild type (filled circles) and the N50Y mutant (open circles). In all experiments errors bars represent a spread of duplicates. Inset shows SDS–PAGE analysis of the purified rat GLUT5 wild type and the deglycosylated N50Y mutant. b, Tryptophan fluorescence quenching (excitation 295 nm; emission 338 nm), after incubation of purified rat GLUT5 mutant (N50Y, W70F, W191F, W239F, W265F, W275F, W338F, W370F) that contains one single tryptophan residue, W419, with increasing concentrations of d-fructose (filled squares) and to the protein previously incubated with the inhibitor mercury chloride (open circles). c, Slab through the surface of the outward-facing rGLUT5 structure as viewed in the plane of membrane. The structure of substrate-bound XylE structure was further superimposed onto rGLUT5 and is shown here as a grey ribbon. In XylE, Trp392 (Trp388 in hGLUT1) is located at the bottom of the cavity (spheres; magenta) and coordinates d-xylose (stick form; yellow). In GLUT5, the equivalent residue is an alanine, making the cavity deeper. d, d-fructose binding as measured by tryptophan fluorescence quenching (excitation 295 nm; emission 338 nm) after incubation with 40 mM d-fructose for wild type (open bar), and TM7 mutations of Ile295 (interacts with TM10 residues) and Tyr296 and Tyr297 residues. Equivalently located tyrosine residues in XylE occlude the sugar-binding site from the outside22. Fluorescence quenching for the mutants are displayed as a percentage of total wild-type binding. In all experiments errors bars indicate s.e.m.; n = 3.
Extended Data Figure 7 The intracellular helical domain (ICH).
a, Cytoplasmic view of the ICH domain after superposition of the open, outward-facing rGLUT5–sFv (grey) and outward-facing occluded E. coli XylE (teal) (4GBY) structures. b, In the outward-facing GLUT5 structure ICH1–ICH3 are linked together by several salt bridges (side chains are labelled and shown as sticks in yellow). In contrast, no polar interactions are formed between ICH5 and either ICH1–ICH3 or cytoplasmic ends of N-terminal TM bundle helices. A salt bridge forms (dotted line in magenta), however, between Glu225 in ICH3 and Arg407 in TM11, which also forms part of the inter-bundle salt-bridge network (side chains are labelled and shown as sticks in cyan). c, In the inward-facing GLUT5 structure, this inter-bundle salt-bridge network is not formed, because the cytoplasmic ends of the N- and C-terminal bundle have moved apart; consistently, the ICH domain functional role is proposed to act as a scaffold domain that further helps to stabilize the outward-facing conformation21.
Extended Data Figure 8 Access to the central cavity and substrate-binding site is gated by TM7 on the outside and TM10 on the inside.
a, Superposition of outward-facing open GLUT5 and outward-facing occluded E. coli XylE (4GBY) structures. The TM numbering for outward-facing occluded XylE has an additional asterisk. The inward-facing GLUT5 structure is coloured as in Fig. 1a and that of XylE in grey. The bound d-xylose is shown in stick representation in green. The r.m.s.d. is 1.38 Å for 290 pairs of Cα atoms (see Methods). b, Superposition of inward-open GLUT5 and inward-occluded E. coli XylE structure (4JA3) with colouring and annotation as described in a. The r.m.s.d. is 1.80 Å for 274 pairs of Cα atoms (see Methods). The bound d-xylose in 4GBY is represented in stick form in green. The ICH domain is not shown for clarity. c, Superposition of inward-facing open GLUT5 and inward-facing open XylE (4JA4) structures as viewed from the cytoplasmic side with colouring and annotation as described in a. The ICH domain is not shown for clarity. The r.m.s.d. is 1.70 Å for 273 pairs of Cα atoms (see Methods).
Supplementary information
Video 1: Alternating access model of GLUT5 transport
This video shows morphing between the open outward-facing, outward occluded, inward-occluded and open inward-facing GLUT5; conformations as viewed parallel to the membrane. The open outward- and inward-facing conformations are based on the GLUT5 structures reported in this study, and occluded states were modeled based on homologous XylE crystal structures (Methods). Morphing between conformations was generated using PyMol. Helix colouring is as in Fig. 1a. (MOV 20436 kb)
Video 2: Alternating access model of GLUT5 transport
This video shows the same as Video 1, but viewed from the extracellular side of the membrane. (MOV 17095 kb)
Video 3: Alternating access model of GLUT5 transport
This video shows the same as Video 1, but viewed from the intracellular side of the membrane. (MOV 16468 kb)
Rights and permissions
About this article
Cite this article
Nomura, N., Verdon, G., Kang, H. et al. Structure and mechanism of the mammalian fructose transporter GLUT5. Nature 526, 397–401 (2015). https://doi.org/10.1038/nature14909
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14909
This article is cited by
-
Multiple roles for the cytoplasmic C-terminal domains of the yeast cell surface receptors Rgt2 and Snf3 in glucose sensing and signaling
Scientific Reports (2024)
-
Rare sugar l-sorbose exerts antitumor activity by impairing glucose metabolism
Communications Biology (2023)
-
Structure and mechanism of oxalate transporter OxlT in an oxalate-degrading bacterium in the gut microbiota
Nature Communications (2023)
-
Establishing mammalian GLUT kinetics and lipid composition influences in a reconstituted-liposome system
Nature Communications (2023)
-
GLUT3 inhibitor discovery through in silico ligand screening and in vivo validation in eukaryotic expression systems
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.