Abstract
Following the discovery of BRD4 as a non-oncogene addiction target in acute myeloid leukaemia (AML)1,2, bromodomain and extra terminal protein (BET) inhibitors are being explored as a promising therapeutic avenue in numerous cancers3,4,5. While clinical trials have reported single-agent activity in advanced haematological malignancies6, mechanisms determining the response to BET inhibition remain poorly understood. To identify factors involved in primary and acquired BET resistance in leukaemia, here we perform a chromatin-focused RNAi screen in a sensitive MLL–AF9;NrasG12D-driven AML mouse model, and investigate dynamic transcriptional profiles in sensitive and resistant mouse and human leukaemias. Our screen shows that suppression of the PRC2 complex, contrary to effects in other contexts, promotes BET inhibitor resistance in AML. PRC2 suppression does not directly affect the regulation of Brd4-dependent transcripts, but facilitates the remodelling of regulatory pathways that restore the transcription of key targets such as Myc. Similarly, while BET inhibition triggers acute MYC repression in human leukaemias regardless of their sensitivity, resistant leukaemias are uniformly characterized by their ability to rapidly restore MYC transcription. This process involves the activation and recruitment of WNT signalling components, which compensate for the loss of BRD4 and drive resistance in various cancer models. Dynamic chromatin immunoprecipitation sequencing and self-transcribing active regulatory region sequencing of enhancer profiles reveal that BET-resistant states are characterized by remodelled regulatory landscapes, involving the activation of a focal MYC enhancer that recruits WNT machinery in response to BET inhibition. Together, our results identify and validate WNT signalling as a driver and candidate biomarker of primary and acquired BET resistance in leukaemia, and implicate the rewiring of transcriptional programs as an important mechanism promoting resistance to BET inhibitors and, potentially, other chromatin-targeted therapies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Zuber, J. et al. RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia. Nature 478, 524–528 (2011)
Dawson, M. A. et al. Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia. Nature 478, 529–533 (2011)
Delmore, J. E. et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 146, 904–917 (2011)
Asangani, I. A. et al. Therapeutic targeting of BET bromodomain proteins in castration-resistant prostate cancer. Nature 510, 278–282 (2014)
Puissant, A. et al. Targeting MYCN in neuroblastoma by BET bromodomain inhibition. Cancer Discov. 3, 308–323 (2013)
Herait, P. E. et al. BET-bromodomain inhibitor OTX015 shows clinically meaningful activity at nontoxic doses: interim results of an ongoing phase I trial in hematologic malignancies. Proc. 105th Annu. Meet. Am. Assoc. Cancer Res. Apr. 5–9 Abstract CT231 (2014)
Jang, M. K. et al. The bromodomain protein Brd4 is a positive regulatory component of P-TEFb and stimulates RNA polymerase II-dependent transcription. Mol. Cell 19, 523–534 (2005)
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013)
Roe, J.-S., Mercan, F., Rivera, K., Pappin, D. J. & Vakoc, C. R. BET bromodomain inhibition suppresses the function of hematopoietic transcription factors in acute myeloid leukemia. Mol. Cell 58, 1028–1039 (2015)
Filippakopoulos, P. et al. Selective inhibition of BET bromodomains. Nature 468, 1067–1073 (2010)
Fellmann, C. et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep. 5, 1704–1713 (2013)
De Raedt, T. et al. PRC2 loss amplifies Ras-driven transcription and confers sensitivity to BRD4-based therapies. Nature 514, 247–251 (2014)
Neff, T. et al. Polycomb repressive complex 2 is required for MLL-AF9 leukemia. Proc. Natl Acad. Sci. USA 109, 5028–5033 (2012)
Shi, J. et al. The Polycomb complex PRC2 supports aberrant self-renewal in a mouse model of MLL-AF9;NrasG12D acute myeloid leukemia. Oncogene 32, 930–938 (2013)
Zuber, J. et al. An integrated approach to dissecting oncogene addiction implicates a Myb-coordinated self-renewal program as essential for leukemia maintenance. Genes Dev. 25, 1628–1640 (2011)
Di Croce, L. & Helin, K. Transcriptional regulation by Polycomb group proteins. Nature Struct. Mol. Biol. 20, 1147–1155 (2013)
Rahman, S. et al. The Brd4 extraterminal domain confers transcription activation independent of pTEFb by recruiting multiple proteins, including NSD3. Mol. Cell. Biol. 31, 2641–2652 (2011)
Ballaré, C. et al. Phf19 links methylated Lys36 of histone H3 to regulation of Polycomb activity. Nature Struct. Mol. Biol. 19, 1257–1265 (2012)
Shi, J. et al. Role of SWI/SNF in acute leukemia maintenance and enhancer-mediated Myc regulation. Genes Dev. 27, 2648–2662 (2013)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
He, T. C. et al. Identification of c-MYC as a target of the APC pathway. Science 281, 1509–1512 (1998)
Reya, T. et al. A role for Wnt signalling in self-renewal of haematopoietic stem cells. Nature 423, 409–414 (2003)
Wang, Y. et al. The Wnt/β-catenin pathway is required for the development of leukemia stem cells in AML. Science 327, 1650–1653 (2010)
Arnold, C. D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013)
Noubissi, F. K. et al. CRD-BP mediates stabilization of βTrCP1 and c-myc mRNA in response to β-catenin signalling. Nature 441, 898–901 (2006)
Kolligs, F. T. et al. ITF-2, a downstream target of the Wnt/TCF pathway, is activated in human cancers with beta-catenin defects and promotes neoplastic transformation. Cancer Cell 1, 145–155 (2002)
Klijn, C. et al. A comprehensive transcriptional portrait of human cancer cell lines. Nature Biotechnol. 33, 306–312 (2014)
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012)
Thorne, C. A. et al. Small-molecule inhibition of Wnt signaling through activation of casein kinase 1α. Nature Chem. Biol. 6, 829–836 (2010)
Knoechel, B. et al. An epigenetic mechanism of resistance to targeted therapy in T cell acute lymphoblastic leukemia. Nature Genet. 46, 364–370 (2014)
Fong, C. Y. et al. BET inhibitor resistance emerges from leukaemia stem cells. Nature http://dx.doi.org/10.1038/nature14888 (2015)
Fellmann, C. et al. Functional identification of optimized RNAi triggers using a massively parallel sensor assay. Mol. Cell 41, 733–746 (2011)
Dow, L. E. et al. A pipeline for the generation of shRNA transgenic mice. Nature Protocols 7, 374–393 (2012)
Goecks, J., Nekrutenko, A. & Taylor, J. Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 11, R86 (2010)
Zuber, J. et al. Mouse models of human AML accurately predict chemotherapy response. Genes Dev. 23, 877–889 (2009)
Lito, P. et al. Disruption of CRAF-mediated MEK activation is required for effective MEK inhibition in KRAS mutant tumors. Cancer Cell 25, 697–710 (2014)
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001)
Jonkers, J. et al. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nature Genet. 29, 418–425 (2001)
Chou, T. C. Theoretical basis, experimental design, and computerized simulation of synergism and antagonism in drug combination studies. Pharmacol. Rev. 58, 621–681 (2006)
Zuber, J. et al. Toolkit for evaluating genes required for proliferation and survival using tetracycline-regulated RNAi. Nature Biotechnol. 29, 79–83 (2011)
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet.journal 17, 10 (2011)
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359 (2012)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-seq. Bioinformatics 25, 1105–1111 (2009)
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015)
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-seq. Nature Methods 5, 621–628 (2008)
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014)
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Bennett, J. M. et al. Proposed revised criteria for the classification of acute myeloid leukemia. A report of the French–American–British cooperative group. Ann. Intern. Med. 103, 620–625 (1985)
Bennett, J. M. et al. Proposals for the classification of the acute leukaemias. French–American–British (FAB) co-operative group. Br. J. Haematol. 33, 451–458 (1976)
Vardiman, J. W. et al. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: rationale and important changes. Blood 114, 937–951 (2009)
Acknowledgements
We thank M. Weißenböck, B. Hopfgartner and M. Fellner for technical support, S.-M. Kula and the IMP/IMBA Molecular Biology Service for help with library construction, G. Schmauß, T. Lendl, M. Weninger and G. Stengl and the Biooptics Service Facility for FACS; A. Sommer and his team at Campus Science Support Facilities (http://www.csf.ac.at) for Illumina sequencing, S. W. Lowe for reagents and discussions, and all members of the Zuber laboratory for reagents, protocols and discussions. This research was funded by a Starting Grant of the European Research Council (ERC no. 336860; to J.Z.), SFB grants F4704 and F4710 of the Austrian Science Fund (FWF), a Fellowship of the People Programme (Marie Curie Actions) of the European Union (to P.R.). F.M. is an EMBO long-term fellow (EMBO ALTF 491-2014) and research in the Stark laboratory is supported by an ERC Starting grant (no. 242922; to A.S.). The Zuber laboratory and research at the IMP is generously supported by Boehringer Ingelheim.
Author information
Authors and Affiliations
Contributions
P.R. designed the shRNA library and performed the RNAi screen, P.R. and M.R. planned and performed most experiments and analysed data. T.N. and E.A. performed bioinformatics analysis of RNA- and ChIP-seq data. F.M., Ł.M.B. and A.S. performed and analysed STARR-seq experiments. M.M. generated validated BRD2/3/4 shRNAs and performed experiments. J.R.-S. and C.V. contributed ChIP-seq data and provided advice. B.P., S.C.-R. and P.V. performed analyses in primary patient samples. S.D. performed experiments in PDAC. T.H. improved protocols for shRNA deep-sequencing. J.J. contributed critical reagents and performed CFU assays. L.E.D. provided critical reagents. N.S., U.T.-G. and D.G. profiled JQ1 sensitivity and performed RNA-seq in human cell lines. M.P. and N.K. provided critical advice and support. J.Z. designed experiments, analysed data, and supervised this research. P.R., M.R. and J.Z. co-wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Multiplexed shRNAmir screening for chromatin-associated dependencies in AML maintenance and BET resistance.
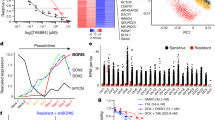
a, Scatter plot illustrating the correlation of normalized reads per shRNA at all three time points (T0, T1 and T2; top to bottom) compared to the plasmid pool in all three independent replicates (left to right). The schematic to the right illustrates the different sampling time points subjected to deep sequencing. Top hits enriched under JQ1 treatment and positive controls from the initial negative-selection screen (T1) are marked with coloured dots according to the legend on the right. b, Pooled negative-selection screening depicting changes in representation of 2,917 shRNAs during 7 days of culture. shRNA fold depletion values were calculated by dividing the number of reads after 7 days of culture (T1) by the number of reads obtained from the plasmid pool, and are plotted as the mean of three replicates in ascending order. Completely depleted shRNAs (0 reads at T1) obtained a fold depletion value of 1 × 10−3. Positive control shRNAs targeting essential genes are marked in red; negative control shRNAs are depicted in green. c, Scatter plot depicting all genes ranked by the sum of their average depletion score of all shRNAs across all three replicates. Top scoring hits were defined as genes for which at least two shRNAs showed an average depletion of eightfold after 7 days of shRNA expression and are marked in red (45 genes). d, IC50 determination for JQ1 in murine MLL–AF9;NrasG12D AML cells (RN2). Obtained numbers of viable cells per ml were normalized to DMSO (n = 3, mean ± s.e.m.). e, Table showing the top ten enriched shRNAs at T2. shRNAs targeting Suz12, Dnmt3a and Psip1 are strongly enriched in all three independent replicates. f, Relative mRNA abundance (RPKM) of shRNA target genes in JQ1-resistant leukaemia cells expressing the indicated shRNAs, plotted relative to leukaemia cells expressing Ren.713.
Extended Data Figure 2 PRC2 suppression confers resistance to JQ1.
a, Competitive proliferation assays of MLL–AF9;NrasG12D leukaemia cells transduced with pLMN constructs expressing the indicated shRNAs. Shown is the fraction of GFP+/shRNA+ cells (relative to the initial ratio) under treatment with JQ1 (50 nM) or DMSO over time (continuation from Fig. 1d). JQ1 treatment was initiated at the indicated time points (red arrow). b, Competitive proliferation assays evaluating one validated shRNA targeting Eed as an additional core component of the PRC2 complex (as in a). c, Competitive proliferation assays of Tet-on competent MLL–AF9;NrasG12D leukaemia cells expressing the indicated shRNAs from a Tet‐inducible vector (pRT3GEN-miR30). Transduced cells were selected with G418 and subsequently mixed with wild-type (wt) cells in a ratio of 95% to 5%. shRNA-expression was induced using dox treatment (1 µg ml−1; from day 0), and the fraction of GFP+ cells was measured over time and plotted relative to day 2. JQ1 treatment (50 nM) was initiated in one of the duplicate samples at the indicated time point (red arrow). Once the percentage of viable cells was below 10%, measurements were discontinued (indicated in the graph by the discontinuation of the respective sample). The negative effect induced by Suz12 suppression is reverted upon treatment with JQ1, which is not the case when Myc is suppressed. d, Bar chart showing colony-forming cell frequencies of MLL–AF9;NrasG12D leukaemia cells expressing Ren.713 or Suz12.1676 shRNAs in the presence of DMSO or 200 nM JQ1; type 1, myeloblasts; type 2, maturing myeloblasts; type 3, terminally differentiated myeloid cells (n = 3, mean ± s.e.m.). e, Pie charts depicting the fraction of JQ1-response genes which are re-expressed to the indicated extent in mouse resistant AML cells expressing shRNAs targeting Dnmt3a and Psip1 (continuation of Fig. 2b). JQ1-response genes (Fig. 2a) were grouped into four categories based on the divergence of their expression in resistant AML compared to AML expressing Ren.713 (not changed compared to expression after 24 h JQ1, less than 1.5-fold; restoration relative to DMSO control: full restoration, less than 1.5-fold; enhanced, above 1.5-fold; partial restoration, restored but less than 1.5 fold) f, Heat map showing the distribution of H3K27me3, H3K36me3 and H3K4me3 ChIP-seq peaks (fold enrichment >5 over input, FDR <1%) relative to Brd4 enhancer and promoter binding sites in MLL–AF9;NrasG12D AML cells with and without JQ1 treatment. g, ChIP-seq occupancy profiles of Brd4 and H3K27me3 and H3K36me3 chromatin marks at enhancer regions downstream of Myc and upstream of Tifab following 3 days of treatment with vehicle or JQ1 (25 nM). y axis reflects the number of normalized cumulative tag counts in each region.
Extended Data Figure 3 Changes in the regulatory landscape of BET-inhibitor-resistant mouse AML cells.
a, Global H3K27 acetylation density of Suz12.1842-expressing resistant MLL–AF9;NrasG12D leukaemia cells under long term (LT) treatment with 50 nM JQ1 (red bar), after 4 days of drug withdrawal (orange bar) and in Ren.713 controls (blue bar; statistical significance determined using Student’s t-test). b, Left panel, sorted fold change (FC) ratios of H3K27ac peaks in long-term JQ1-treated MLL–AF9;NrasG12D leukaemia cells expressing Suz12.1842 compared to cells expressing Ren.713 control shRNA. Included were all peaks showing > 10 reads per million in at least one condition. Right panel, top 15 gained peaks and their associated genes (defined using the closest transcription start site, TSS). The Myc proximal enhancer in the first intron of Pvt1 is highlighted in red as one of the most differentially enriched peaks (FC = 4.18). c, qRT–PCR validation of presented H3K27ac ChIP-seq at the indicated regions downstream of the Myc locus (n = 3, mean ± s.e.m., statistical significance determined using Student’s t-test). d, Gene set enrichment analysis plots of three publicly available gene sets associated with signalling pathways comparing expression changes in resistant MLL–AF9;NrasG12D AML cells induced by suppression of PRC2 complex members, compared to control cells expressing Ren.713 shRNA (n = 2) or empty vector (continuation of Fig. 2e, f). e, Core signature genes of KEGG-curated Wnt and TGF-β gene sets with increased expression in resistant murine MLL–AF9;NrasG12D cells, compared to sensitive cells. Red coloured bars indicate association with H3K27 methylation in JQ1-sensitive MLL–AF9;NrasG12D AML cells. f, H3K27 methylation density at three exemplified genes with high expression changes in JQ1-resistant murine AML.
Extended Data Figure 4 Resistant MLL–AF9;NrasG12D AML cells generated through Suz12 suppression are not enriched for LSC-associated surface markers or expression signatures.
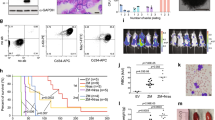
a, Immunophenotyping of JQ1-resistant mouse AML cells expressing two independent Suz12 shRNAs stably cultured in 50 nM JQ1 for more than 4 weeks (LT) compared to cells expressing Ren.713 control shRNA. Data are representative of two independent biological replicates. b, Percentage of c-Kit+ cells in CD45.2+ bone marrow cells isolated from terminally diseased CD45.1+ mice following transplantation with MLL–AF9;NrasG12D cells expressing Ren.713 or Suz12.1676 and in vivo treatment with DMSO carrier or JQ1 (50 mg kg−1 per day) (n = 5, mean ± s.e.m.). c, Gene set enrichment analysis evaluating changes in macrophage and LSC gene signatures in resistant MLL–AF9;NrasG12D AML expressing three PRC2 shRNAs compared to cells expressing Ren.713 shRNA (n = 2) or empty vector (see also Fig. 2e, f).
Extended Data Figure 5 Comparison of JQ1-response genes in sensitive and resistant cancer cell lines.
a, JQ1 sensitivity profiling in 246 human cancer cell lines of different tissue contexts. Shown are GI50 values determined using Alamar blue staining after 72 h. b, Principal component analysis of steady-state transcriptomes (based on RPKM) and c, transcription changes (based on fold change, FC) following 2 h of JQ1 treatment (200 nM) in indicated sensitive and resistant cancer cell lines of different tissue context. Steady-state profiles cluster based on tissue context, while neither baseline nor dynamic expression analysis can accurately distinguish sensitive and resistant contexts. d, MYC mRNA levels (RPKM) at different time points after JQ1 treatment (200 nM) in indicated cell lines, relative to levels in DMSO-treated cells. Individual cell line pairs are grouped for their tissue context and coloured according to their sensitivities (green, sensitive; red, resistant). e, Number of genes twofold up- or downregulated upon JQ1 treatment (200 nM) after 2 h and 24 h (minimum expression >3 RPKM). f, Pairwise overlap of commonly up- or downregulated genes after 2 h of JQ1 treatment (200 nM) relative to DMSO control. Each cell colour corresponds to the relative number of commonly up- or downregulated genes in the cell lines listed in the respective row and column. The total number of genes regulated per cell line is indicated in black next to each cell line. Only little overlap is observed between cell lines of the same tissue context as well as between JQ1-sensitive or -resistant cell lines.
Extended Data Figure 6 shRNA-based analysis of BRD2/3/4-dependent target genes and detailed analysis of effects on HEXIM1 and MYC transcription in different cell lines.
a, Determination of knockdown levels of a set of BRD2, BRD3 and BRD4 shRNAs determined using an established fluorescence-based shRNAmir reporter assay11. Knockdown levels were quantified relative to Ren.713 control shRNA. b, Competitive proliferation assays in MOLM-13 cells functionally evaluating the potency of BRD2, BRD3 and BRD4 shRNAs over time using a Tet‐regulated vector (pRT3GEN) in presence of doxycycline. Only shRNAs targeting BRD4 induce a proliferative disadvantage and lead to rapid depletion of GFP+ cells over time. Red labels indicate most potent shRNAs based on results obtained from reporter and competitive proliferation assay. c, Determination of BRD2, BRD3 and BRD4 mRNA levels in K-562 and MOLM-13 cells expressing the indicated shRNA. d, Number of genes commonly up- or downregulated with a fold change (FC) >3 in MOLM-13 or K-562 cells following expression of indicated validated shRNAs or treatment with 200 nM JQ1 for 24 h. JQ1-induced expression changes show the largest overlap with cells expressing a validated BRD4 shRNA, suggesting that suppression of BRD4 is the key effector mechanism of JQ1 in leukaemia. e, Venn diagrams showing the overlap of genes commonly up- or downregulated following 2 h and 24 h of JQ1 either in all contexts, or specifically in sensitive or resistant leukaemias. f, HEXIM1/Hexim1 expression (RPKM) in all 18 analysed human cell lines and murine MLL–AF9;NrasG12D AML cells after 2 h and 24 h treatment with 200 nM JQ1, compared to DMSO control (statistical significance was determined using a paired Student’s t-test). g, Relative MYC mRNA levels determined by qRT–PCR quantification in the indicated cell lines after incubation with 200 nM JQ1 measured at different treatment time points. Cell lines are grouped according to their sensitivity and the respective IC50 is presented below. Resistant cell lines rapidly restore MYC transcription (n = 3, mean ± s.e.m.).
Extended Data Figure 7 Dynamic STARR-seq analysis of enhancer activity in K-562 cells.
a, Schematic representation of the STARR-seq cloning and screening strategy in K-562 cells. BACs available for the extended MYC locus (covering approximately 91% of a 3.1 Mb region at the MYC locus) and 25 genic control BACs were fragmented and cloned into a modified STARR-seq vector containing a minimal MYC promoter. This library was then screened for enhancer activity using STARR-seq in K-562 cells with or without 250 nM JQ1. The schematic shows the underlying principle of STARR-seq: a bona fide enhancer can activate its own transcription from a minimal MYC promoter. Messenger RNA corresponding to active enhancer elements will therefore become more abundant among the cellular RNA compared to inactive fragments. b, PVT1 mRNA levels (RPKM) at different time points after JQ1 treatment (200 nM) in indicated cell lines, relative to levels in DMSO-treated cells. PVT1 expression is generally reduced upon JQ1 treatment indicating no association with enhancer activation.
Extended Data Figure 8 Analysis and functional validation of Wnt signalling as a key driver of BET resistance.
a, Protein levels of TCF4 and IGF2BP1 in K-562 cells transduced with pRT3GEN expressing indicated shRNAs after 7 days of doxycycline treatment, compared to Ren.713 and wild-type (wt) control samples. b, Competitive proliferation assay of JQ1-sensitive MOLM-13 cells expressing GFP-linked TCF4 cDNA or empty vector. Plotted is the relative fraction of GFP+ cells 72 h after JQ1 treatment using the indicated doses (n = 3, means ± s.e.m., ***P < 0.001; **P < 0.01; *P < 0.05 as determined by Student’s t-test). Cells overexpressing TCF4 exhibit a dose-dependent competitive advantage under JQ1 treatment. c, Protein levels of TCF4 after overexpression of TCF4 cDNA subcloned into pMSCV-PGK-NeoR-IRES-GFP in MOLM-13 after 4 weeks of G418 (0.5 mg ml−1) selection, compared to MOLM-13 transduced with empty control vector and to TCF4 protein levels in K-562 cells. d, ChIP qRT–PCR analysis of TCF7L2 binding to AXIN2, SP5, MYC promoter and the PVT1 enhancer element in K-562 cells at indicated time points after treatment with 200 nM JQ1 (n = 2 biological replicates, mean ± s.e.m.). TCF7L2 binding increases gradually over time at promoters of Wnt target genes and the PVT1 enhancer at the MYC locus. e, Protein levels of Apc in 3T3 murine fibroblast cells 7 days after infection with the indicated shRNAs cloned into pLMP compared to Ren.713 and wild-type control samples. f, Relative Axin2 mRNA expression levels, determined by qRT–PCR normalized to B2m, after expression of the indicated shRNAs targeting Apc. g, Competitive proliferation assays of MLL–AF9;NrasG12D leukaemia cells transduced with pLMP constructs expressing shRNAs targeting Apc, in combination with JQ1 (50 nM) or DMSO over time (continuation from Fig. 4d). h, Top, bioluminescent imaging of mice transplanted with 1 × 105 MLL–AF9;NrasG12D leukaemia cells expressing constitutively active Ctnnb1 (Ctnnb1.4x). Treatment with JQ1 (50 mg kg−1 per day) or DMSO carrier started at day 1 after injection. Bottom, Kaplan–Meier survival curves of control and JQ1-treated mice demonstrate decreased survival rates in mice treated with JQ1 (n = 5). Statistical significance was calculated using the log-rank test. i, Competitive proliferation assays of BCR/ABLp210;p53−/− and MLL/ENL;NrasG12D leukaemia cells transduced with constitutively active Ctnnb1 (Ctnnb1.4x) or empty vector control in the absence or presence of JQ1. Measurements started 4 days after transduction together with JQ1 treatment (50 nM). Shown is the fraction of GFP+ cells (relative to initial) over time.
Extended Data Figure 9 Wnt signatures are generally associated with BET resistance and Wnt activation drives resistance of pancreatic cancer models.
The BETi_Resistance_Rathert signature was defined by combining the core enriched genes from the two significant Wnt gene sets (KEGG_WNT_SIGNALING and ST_WNT_SIGNALING) in murine resistant AML, filtered for significant upregulation (DESeq padj <0.1). These were combined with the Wnt-associated genes found differentially expressed in resistant human leukaemia cell lines and primary patient samples, resulting in a total of 26 genes. a, Left, microarray expression data of all 26 signature genes was curated from the Cancer Cell Line Encyclopedia (CCLE)28 and normalized to the geometric mean of each individual gene throughout all samples (relative expression). The sum of the relative expression of all genes (resistance index) was plotted for all CCLE cell lines showing a GI50 > 450 nM (resistant, 54 total) or a GI50 < 150 nM (sensitive, 55 total) based on our sensitivity profiling (statistical significance was determined using Student’s t-test). Right, gene set enrichment analysis plot comparing the expression of 26 signature genes associated with JQ1 resistance across 54 resistant (GI50 > 450 nM) and 55 sensitive cell lines (GI50 < 150 nM) available from CCLE. b, Left, as in a the resistance index was calculated as the sum of relative expression values of all 26 signature genes, which were based on RPKM extracted from an independent RNA-seq data set27. Plotted are all 49 resistant (GI50 > 450 nM) and all 50 sensitive (GI50 < 150 nM) cell lines analysed in both sensitivity profiling and RNA-seq27 (statistical significance determined using Student’s t-test). Right, gene set enrichment analysis plot comparing the expression of 26 signature genes associated with JQ1 resistance across 49 resistant (GI50 > 450 nM) and 50 sensitive cell lines (GI50 < 150 nM) available from ref. 27. c, Competitive proliferation assays of murine pancreatic adenocarcinoma (KRPC2) cells transduced with pLEPC constructs expressing potent validated shRNAs targeting Apc, cultured in the presence of JQ1 (800 nM) or DMSO. Shown is the relative number of mCherry+ cells over time, relative to initial. d, Cell viability of murine KRPC2 was determined following 5 days of treatment with pyrvinium and/or JQ1 as indicated (n = 3, mean ± s.e.m., statistical significance determined using Student’s t-test). The combination index (CI) for drug combinations was calculated using the CompuSyn software and percentage inhibition (fraction affected, Fa) resulting from combined action of the two drugs versus effects of either drug alone. CI values <1.0 indicate synergism of the two agents. e, As in d, cell viability was determined for human ASPC-1 pancreatic cancer cells following 5 days of treatment with pyrvinium and/or JQ1 as indicated (n = 3, mean ± s.e.m., statistical significance determined using Student’s t-test). The combination index for drug combinations was obtained using percentage inhibition (fraction affected, Fa) resulting from combined action of the two drugs versus effects of either drug alone.
Extended Data Figure 10 Expression analysis of Wnt-associated genes in primary AML patient samples.
a, Determination of JQ1 response profiles in 12 primary AML patient samples. Sensitivity was determined using 3H-thymidine uptake across different JQ1 concentrations. b, qRT–PCR analysis of mRNA levels of additional Wnt-associated genes in primary human leukaemia samples relative to GAPDH (continuation of Fig. 4g). Patient groups with low JQ1 IC50 (<200 nM, blue dots) were compared to patients with high IC50 (>500 nM, red dots). Statistical significance was determined using a Student’s t-test. c, Definition of a JQ1 resistance index. Expression of HOXB4, TCF4 and CCND2 in each primary AML patient sample was normalized to the geometric mean of all samples. The sum of these relative expression values of all three genes were added up to a resistance index, which was plotted in comparison to the JQ1 IC50 of each sensitive (IC50 < 200 nM, blue dots) and resistant (IC50 > 500 nM, red dots) AML patient sample. Two-tailed Pearson correlation coefficient r and P value are shown.
Supplementary information
Supplementary Figure
This file contains the uncropped scans of Western blot in Figure 1g and 3b with size marker indications. (PDF 816 kb)
Supplementary Tables
This file contains the following Supplementary Tables: shRNAs used in the described multiplexed shRNAmir screen, including sequences, as well as a summary of the primary screening data, all cell lines profiled for JQ1 sensitivity, shRNAs used in single assays, a list of primer sequences used in this study, gene names and accession numbers of identical genes, GSEA gene sets used in this study, STARR-seq BACs and primary patient samples. (XLSX 1432 kb)
Rights and permissions
About this article
Cite this article
Rathert, P., Roth, M., Neumann, T. et al. Transcriptional plasticity promotes primary and acquired resistance to BET inhibition. Nature 525, 543–547 (2015). https://doi.org/10.1038/nature14898
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14898
This article is cited by
-
RAPID resistance to BET inhibitors is mediated by FGFR1 in glioblastoma
Scientific Reports (2024)
-
Targeting bromodomain-containing proteins: research advances of drug discovery
Molecular Biomedicine (2023)
-
Bromodomain and extraterminal (BET) proteins: biological functions, diseases, and targeted therapy
Signal Transduction and Targeted Therapy (2023)
-
Rational combinations of targeted cancer therapies: background, advances and challenges
Nature Reviews Drug Discovery (2023)
-
Superenhancers as master gene regulators and novel therapeutic targets in brain tumors
Experimental & Molecular Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.