Abstract
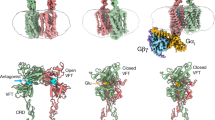
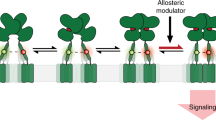
G-protein-coupled receptors (GPCRs) constitute the largest family of membrane receptors in eukaryotes. Crystal structures have provided insight into GPCR interactions with ligands and G proteins1,2, but our understanding of the conformational dynamics of activation is incomplete. Metabotropic glutamate receptors (mGluRs) are dimeric class C GPCRs that modulate neuronal excitability, synaptic plasticity, and serve as drug targets for neurological disorders3,4. A ‘clamshell’ ligand-binding domain (LBD), which contains the ligand-binding site, is coupled to the transmembrane domain via a cysteine-rich domain, and LBD closure seems to be the first step in activation5,6. Crystal structures of isolated mGluR LBD dimers led to the suggestion that activation also involves a reorientation of the dimer interface from a ‘relaxed’ to an ‘active’ state7,8, but the relationship between ligand binding, LBD closure and dimer interface rearrangement in activation remains unclear. Here we use single-molecule fluorescence resonance energy transfer to probe the activation mechanism of full-length mammalian group II mGluRs. We show that the LBDs interconvert between three conformations: resting, activated and a short-lived intermediate state. Orthosteric agonists induce transitions between these conformational states, with efficacy determined by occupancy of the active conformation. Unlike mGluR2, mGluR3 displays basal dynamics, which are Ca2+-dependent and lead to basal protein activation. Our results support a general mechanism for the activation of mGluRs in which agonist binding induces closure of the LBDs, followed by dimer interface reorientation. Our experimental strategy should be widely applicable to study conformational dynamics in GPCRs and other membrane proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Rasmussen, S. G. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011)
Katritch, V., Cherezov, V. & Stevens, R. C. Structure-function of the G protein-coupled receptor superfamily. Annu. Rev. Pharmacol. Toxicol. 53, 531–556 (2013)
Conn, P. J. & Pin, J. P. Pharmacology and functions of metabotropic glutamate receptors. Annu. Rev. Pharmacol. Toxicol. 37, 205–237 (1997)
Niswender, C. M. & Conn, P. J. Metabotropic glutamate receptors: physiology, pharmacology, and disease. Annu. Rev. Pharmacol. Toxicol. 50, 295–322 (2010)
Kniazeff, J. et al. Locking the dimeric GABAB G-protein-coupled receptor in its active state. J. Neurosci. 24, 370–377 (2004)
Kumar, J. & Mayer, M. L. Functional insights from glutamate receptor ion channel structures. Annu. Rev. Physiol. 75, 313–337 (2013)
Kunishima, N. et al. Structural basis of glutamate recognition by a dimeric metabotropic glutamate receptor. Nature 407, 971–977 (2000)
Tsuchiya, D., Kunishima, N., Kamiya, N., Jingami, H. & Morikawa, K. Structural views of the ligand-binding cores of a metabotropic glutamate receptor complexed with an antagonist and both glutamate and Gd3+ . Proc. Natl Acad. Sci. USA 99, 2660–2665 (2002)
Roy, R., Hohng, S. & Ha, T. A practical guide to single-molecule FRET. Nature Methods 5, 507–516 (2008)
Zhao, Y. et al. Single-molecule dynamics of gating in a neurotransmitter transporter homologue. Nature 465, 188–193 (2010)
Bockenhauer, S., Furstenberg, A., Yao, X. J., Kobilka, B. K. & Moerner, W. E. Conformational dynamics of single G protein-coupled receptors in solution. J. Phys. Chem. B 115, 13328–13338 (2011)
Morrison, E. A. et al. Antiparallel EmrE exports drugs by exchanging between asymmetric structures. Nature 481, 45–50 (2011)
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo . Nature Biotechnol. 21, 86–89 (2003)
Doumazane, E. et al. A new approach to analyze cell surface protein complexes reveals specific heterodimeric metabotropic glutamate receptors. FASEB J. 25, 66–77 (2011)
Doumazane, E. et al. Illuminating the activation mechanisms and allosteric properties of metabotropic glutamate receptors. Proc. Natl Acad. Sci. USA 110, E1416–E1425 (2013)
Jain, A. et al. Probing cellular protein complexes using single-molecule pull-down. Nature 473, 484–488 (2011)
Kniazeff, J. et al. Closed state of both binding domains of homodimeric mGlu receptors is required for full activity. Nature Struct. Mol. Biol. 11, 706–713 (2004)
Jin, R., Banke, T. G., Mayer, M. L., Traynelis, S. F. & Gouaux, E. Structural basis for partial agonist action at ionotropic glutamate receptors. Nature Neurosci. 6, 803–810 (2003)
Yamashita, T., Kai, T., Terakita, A. & Shichida, Y. A novel constitutively active mutation in the second cytoplasmic loop of metabotropic glutamate receptor. J. Neurochem. 91, 484–492 (2004)
Yanagawa, M., Yamashita, T. & Shichida, Y. Activation switch in the transmembrane domain of metabotropic glutamate receptor. Mol. Pharmacol. 76, 201–207 (2009)
Kubo, Y., Miyashita, T. & Murata, Y. Structural basis for a Ca2+-sensing function of the metabotropic glutamate receptors. Science 279, 1722–1725 (1998)
Nash, M. S., Saunders, R., Young, K. W., Challiss, R. A. & Nahorski, S. R. Reassessment of the Ca2+ sensing property of a type I metabotropic glutamate receptor by simultaneous measurement of inositol 1,4,5-trisphosphate and Ca2+ in single cells. J. Biol. Chem. 276, 19286–19293 (2001)
Tateyama, M., Abe, H., Nakata, H., Saito, O. & Kubo, Y. Ligand-induced rearrangement of the dimeric metabotropic glutamate receptor 1alpha. Nature Struct. Mol. Biol. 11, 637–642 (2004)
Huang, S. et al. Interdomain movements in metabotropic glutamate receptor activation. Proc. Natl Acad. Sci. USA 108, 15480–15485 (2011)
Hlavackova, V. et al. Sequential inter- and intrasubunit rearrangements during activation of dimeric metabotropic glutamate receptor 1. Sci. Signal. 5, ra59 (2012)
Xue, L. et al. Major ligand-induced rearrangement of the heptahelical domain interface in a GPCR dimer. Nature Chem. Biol. 11, 134–140 (2015)
Lohse, M. J., Maiellaro, I. & Calebiro, D. Kinetics and mechanism of G protein-coupled receptor activation. Curr. Opin. Cell Biol. 27, 87–93 (2014)
Olofsson, L. et al. Fine tuning of sub-millisecond conformational dynamics controls metabotropic glutamate receptors agonist efficacy. Nature Commun. 5, 5206 (2014)
Muto, T., Tsuchiya, D., Morikawa, K. & Jingami, H. Structures of the extracellular regions of the group II/III metabotropic glutamate receptors. Proc. Natl Acad. Sci. USA 104, 3759–3764 (2007)
Haran, G. Noise reduction in single-molecule fluorescence trajectories of folding proteins. Chem. Phys. 307, 137–145 (2004)
McKinney, S. A., Joo, C. & Ha, T. Analysis of single-molecule FRET trajectories using hidden Markov modeling. Biophys. J. 91, 1941–1951 (2006)
Zhou, R. et al. SSB functions as a sliding platform that migrates on DNA via reptation. Cell 146, 222–232 (2011)
Acknowledgements
We thank Z. Fu and H. Okada for technical assistance, J. P. Pin for generously providing the SNAP- and CLIP-tagged mGluRs and advice on their properties, and J. P. Pin, E. Margeat, P. Rondard, A. Jain, A. Reiner and members of the Isacoff laboratory for discussions. Funding was provided by the National Institutes of Health Nanomedicine Development Center for the Optical Control of Biological Function (2PN2EY018241) and the National Science Foundation (EAGER: IOS-1451027). R.V. is a Merck fellow of the Life Science Research Foundation.
Author information
Authors and Affiliations
Contributions
R.V., J.L. and E.Y.I. designed the research. R.V. set up, performed and analysed single-molecule FRET experiments. J.L. performed and analysed ensemble FRET and electrophysiology experiments and contributed to single-molecule FRET experiments. R.V., J.L. and E.IY. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Fluorophore labelling is specific and FRET constructs are functional in HEK293T cells.
a, SNAP–mGluR2 shows glutamate-induced currents in cells co-expressing GIRK channels. b, Treatment with 2.5 μM benzylguanine Alexa-647 (for SNAP) followed by 5 μM benzylcytosine DY-547 (for CLIP) produces specific and orthogonal labelling of SNAP and CLIP–mGluR2 constructs in HEK293T cells. All conditions were imaged with identical settings in both the red (excitation = 635 nm) and green (excitation = 561 nm) channels. c, Ensemble FRET measurements from HEK293T cells. Top, image of cells expressing SNAP and CLIP–mGluR2 labelled with donor (DY-547, green) and acceptor (Alexa-647, red). Bottom, representative trace showing dose-dependent, reversible decrease in FRET after glutamate application. Glutamate was washed out between applications. d, Ensemble FRET glutamate titration in HEK293T cells. Error bars are s.e.m.
Extended Data Figure 2 Control experiments verifying specificity of the smFRET assay.
a, Representative TIRF image showing single receptors in donor and acceptor channels during donor excitation with a 532 nm laser. b, Single-molecule pull-down of SNAP–mGluR2 on a passivated surface is specific. Left, representative images of individual molecules in the absence or presence of an anti-mGluR2 antibody. Right, quantification of the number of molecules pulled down for each condition. c, Photobleaching step analysis shows that mGluR2 remains a dimer in single-molecule pull-down. Left, representative single-molecule bleaching steps for mGluR2–GFP. Right, histogram of bleaching step counts for all molecules. Dotted red line shows the predicted proportions for an 80% GFP maturation rate. d, Pulldown of lysate from cells expressing only SNAP–mGluR2 (c) or CLIP–mGluR2 (d), and labelled with both donor and acceptor fluorophores confirms labelling specificity at the single-molecule level. e, f, Pulldown of SNAP– and CLIP–mGluR2 via an antibody against an N-terminal HA-tag (e) leads to very similar smFRET histograms (f, filled circles) in the absence (black) or presence of 1 mM glutamate (green) compared to pull-down with a C-terminal antibody (f, open circles). g, Application of either GTP, to remove any co-assembled G proteins, or apyrase, to lock any G proteins onto mGluR2, does not alter smFRET histograms.
Extended Data Figure 3 Further analysis of glutamate-induced smFRET and functional properties of mGluR2.
a, Representative smFRET traces for mGluR2 in the presence of 4 μM or 8 μM glutamate. b, Quantification of the percentage of single-molecule traces showing at least one transition to the active state at different glutamate concentrations c, In HEK293T cells co-expressing mGluR2 and GIRK, LY341495 prevents glutamate-induced inward currents without altering the baseline current. d, Glutamate titration curves produced from fitting FRET histograms to the sum of three Gaussian distributions. e, f, Glutamate induces inward currents via mGluR2 in a dose-dependent manner (n = 9 cells). g, The glutamate-insensitive mGluR2-YADA (Tyr216Ala, Asp295Ala) shows no smFRET response to 50 μM or 1 mM glutamate. h, smFRET histograms showing that application of the competitive antagonist LY341495 reverses the FRET change induced by glutamate. i, Dwell time analysis of mGluR2 for sub-saturating glutamate concentrations. Solid lines show single exponential fits to the data. j, FRET density plots constructed from synchronized transitions from the low to high FRET states show a short dwell at the medium FRET level (yellow box). Error bars are s.e.m.
Extended Data Figure 4 Further analysis of the effects of orthosteric agonists on mGluR2 smFRET.
a, Cross-correlation plot for mGluR2 in the presence of DCG-IV shows concentration-dependent dynamics. b, c, smFRET histogram (b) and cross-correlation plots (c) for mGluR2 in the presence of the full agonist LY379268. d, smFRET histograms in the presence of 1 μM glutamate, 100 nM DCG-IV and 2 nM LY379268 yields comparable occupancy of the active state. e, f, Representative smFRET traces for mGluR2 in the presence of 0.1 μM DCG-IV (e) or 2 nM LY379268 (f), which are the concentrations used for dwell-time analysis. g, h, FRET density plots constructed from the average transitions from the low to high states show a short dwell at the medium FRET level (yellow boxes) for mGluR2 in the presence of DCG-IV (g) or LY379268 (h).
Extended Data Figure 5 TMD mutations that introduce basal activity and a positive allosteric modulator increase the affinity and efficacy of a partial agonist.
a, Crystal structures of the mGluR1 TMD bound to a negative allosteric modulator (PDB code 4OR2) showing the location of conserved residues in TM4 and TM6 previously shown to be sensitive to mutations that induce basal activity. b, Ensemble FRET titrations showing that mutations Gln679Val and Cys770Ala increase the affinity and efficacy of DCG-IV compared to wild-type mGluR2. c, d, smFRET histograms for TMD mutants show population of the same three FRET states as the wild type, but with greater occupation of the low FRET state at either sub-saturating (c) or saturating (d) concentrations of DCG-IV. e, Binding of PAM LY487379 to the TMD of mGluR2 (top) increases the apparent affinity and efficacy of DCG-IV in ensemble FRET measurements in HEK293T cells. All values were normalized to the response to 1 mM glutamate. b, smFRET histograms showing a LY487379 -induced shift in the response to 1 μM DCG-IV. Error bars are s.e.m.
Extended Data Figure 6 Characterization of basic ensemble and smFRET properties of mGluR3 as compared to mGluR2.
a, Activation of GIRK by SNAP–mGluR3 in HEK293T cells. b, smFRET histograms for mGluR3 show glutamate-independent low FRET population. c, Cross-correlation plots for mGluR3 show glutamate-independent dynamics. d, Unlike mGluR3, mGluR2 shows zero or minimal current response to the antagonist LY341495 in the absence of glutamate. e, f, Ensemble FRET in HEK293T cells shows a robust antagonist LY341495-induced FRET increase in mGluR3 (e) but not in mGluR2 (f). g, smFRET histograms for mGluR3 with or without GTP treatment to dissociate any G proteins that may be coupled to the receptor. The time resolution for this data is 100 ms. Error bars are s.e.m.
Extended Data Figure 7 Calcium sensitivity of mGluR3.
a, b, Representative smFRET traces for mGluR3 in the absence (a) or presence (b) of 2 mM Ca2+. c, smFRET histograms for mGluR3 in the presence of various concentrations of calcium. d, Cross-correlation plots for mGluR3 in the presence of various concentrations of calcium. e, FRET density plot for showing that Ca2+-induced synchronized transitions show a similar intermediate in mGluR3, as observed in mGluR2. f, g, smFRET histograms (f) and cross-correlation plots (g) for mGluR2 in the presence of various concentrations of calcium.
Extended Data Figure 8 mGluR3(Ser152Asp) shows decreased basal FRET and calcium sensitivity.
a, Ensemble FRET glutamate titrations for mGluR3 and mGluR3(Ser152Asp). b, Representative ensemble FRET trace for mGluR3(Ser152Asp) shows no response to LY341495. c, Summary of basal FRET for mGluR3, mGluR2 and mGluR3-S152D. Basal FRET = [ΔFRETLY341495]/([ΔFRETLY341495] + [ΔFRETGlu]). d, Representative smFRET traces for mGluR3(Ser152Asp) in the absence (left) or presence (right) of 2 mM Ca2+. e, smFRET histograms for mGluR3(Ser152Asp) in the absence or presence of Ca2+ or saturating glutamate. f, Cross-correlation plots for mGluR3(Ser152Asp). Inset shows the percentage of traces showing dynamics in different ligand conditions. Error bars are s.e.m.
Extended Data Figure 9 Glutamate induced smFRET dynamics of mGluR3 in the absence of calcium.
a, Representative smFRET traces for mGluR3 in the absence of Ca2+ and the presence of sub-saturating glutamate. b, smFRET histogram showing dose-dependent response of mGluR3 to glutamate in the absence of Ca2+. c, Cross-correlation plots showing glutamate-induced smFRET dynamics. mGluR2 (green) at its maximum dynamics shows a smaller cross-correlation amplitude compared to mGluR3.
Extended Data Figure 10 Further characterization of mGluR2(Lys240Ala) and wild-type/YADA heterodimers.
a, Crystal structures of mGluR1 in the relaxed (top; PDB code 1EWT) and active states (bottom; PDB code IEWK) show a reorientation of the dimer interface that brings charged residues of the lower lobe in close proximity. Conserved negatively charged residues are shown in red, and Lys260 (Lys240 in mGluR2) is shown in blue. b, Glutamate titrations show a decreased apparent affinity for mGluR2(Lys240Ala) in ensemble FRET (top) and GIRK current activation (bottom). c, Cross-correlation plots in the presence of saturating glutamate show increased dynamics for mGluR2(Lys240Ala) relative to wild type. d, smFRET histogram showing distributions for wild-type/YADA heterodimers at a range of glutamate concentrations. e, Concentration-dependence of low FRET population in wild-type/YADA heterodimers produced from fitting FRET histograms to the sum of three Gaussian distributions. The EC50 for each phase of the distribution is shown. g, Concentration-dependence of medium FRET population in wild-type/YADA heterodimers and wild-type homodimers produced from fitting FRET histograms to the sum of three Gaussian distributions. g, Three-state fit to FRET histogram for wild-type/YADA heterodimers in the presence of 100 μM glutamate shows substantial population of the medium FRET (0.35) peak. h, Cross-correlation plots for wild-type/YADA heterodimers. Error bars are s.e.m.
Rights and permissions
About this article
Cite this article
Vafabakhsh, R., Levitz, J. & Isacoff, E. Conformational dynamics of a class C G-protein-coupled receptor. Nature 524, 497–501 (2015). https://doi.org/10.1038/nature14679
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14679
This article is cited by
-
Kinetic fingerprinting of metabotropic glutamate receptors
Communications Biology (2023)
-
Homo- and hetero-dimeric subunit interactions set affinity and efficacy in metabotropic glutamate receptors
Nature Communications (2023)
-
Single-molecule visualization of human A2A adenosine receptor activation by a G protein and constitutively activating mutations
Communications Biology (2023)
-
Mechanism of sensitivity modulation in the calcium-sensing receptor via electrostatic tuning
Nature Communications (2022)
-
Ligand modulation of the conformational dynamics of the A2A adenosine receptor revealed by single-molecule fluorescence
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.