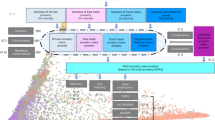
Abstract
Major depressive disorder (MDD), one of the most frequently encountered forms of mental illness and a leading cause of disability worldwide1, poses a major challenge to genetic analysis. To date, no robustly replicated genetic loci have been identified2, despite analysis of more than 9,000 cases3. Here, using low-coverage whole-genome sequencing of 5,303 Chinese women with recurrent MDD selected to reduce phenotypic heterogeneity, and 5,337 controls screened to exclude MDD, we identified, and subsequently replicated in an independent sample, two loci contributing to risk of MDD on chromosome 10: one near the SIRT1 gene (P = 2.53 × 10−10), the other in an intron of the LHPP gene (P = 6.45 × 10−12). Analysis of 4,509 cases with a severe subtype of MDD, melancholia, yielded an increased genetic signal at the SIRT1 locus. We attribute our success to the recruitment of relatively homogeneous cases with severe illness.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Change history
12 April 2019
Editors’ note: We would like to alert our readers to a change in the data availability for this manuscript. The genotype and whole genome sequence data used in this publication were made publicly available at the time of publication through the GigaDB database at http://gigadb.org/dataset/100155. In March 2019, Nature was notified that the whole genome sequence data files have been removed from GigaDB. The genotype data files remain available at GigaDB. The whole genome sequence data was also made available in 2016 at NCBI with accession number PRJNA289433 (https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA289433). In March 2019, Nature was notified that this dataset was subsequently withdrawn from NCBI by request of the submitter. We are currently investigating the data availability for the whole genome sequence data and will publish an update once our investigation is complete.
11 August 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41586-023-06276-1
References
Kessler, R. C. & Bromet, E. J. The epidemiology of depression across cultures. Annu. Rev. Public Health 34, 119–138 (2013)
Flint, J. & Kendler, K. S. The genetics of major depression. Neuron 81, 484–503 (2014)
Major Depressive Disorder Working Group of the Psychiatric GWAS Consortium et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol. Psychiatry 18, 497–511 (2013)
Foley, D. L. et al. Genetic and environmental risk factors for depression assessed by subject-rated symptom check list versus structured clinical interview. Psychol. Med. 31, 1413–1423 (2001)
Kendler, K. S. et al. Clinical indices of familial depression in the Swedish Twin Registry. Acta Psychiatr. Scand. 115, 214–220 (2007)
Sullivan, P. F. et al. Genetic epidemiology of major depression; review and meta-analysis. Am. J. Psychiatry 157, 1552–1562 (2000)
Li, Y. et al. Low-coverage sequencing; implications for design of complex trait association studies. Genome Res. 21, 940–951 (2011)
Lippert, C. et al. FaST linear mixed models for genome-wide association studies. Nature Methods 8, 833–835 (2011)
Widmer, C. et al. Further improvements to linear mixed models for genome-wide association studies. Sci. Rep. 4, 6874 (2014)
Angst, J. et al. Melancholia and atypical depression in the Zurich study: epidemiology, clinical characteristics, course, comorbidity and personality. Acta Psychiatr. Scand. Suppl. 433, 72–84 (2007)
Kendler, K. S. The diagnostic validity of melancholic major depression in a population-based sample of female twins. Arch. Gen. Psychiatry 54, 299–304 (1997)
Sun, N. et al. A comparison of melancholic and nonmelancholic recurrent major depression in Han Chinese women. Depress. Anxiety 29, 4–9 (2012)
Pasaniuc, B. et al. Extremely low-coverage sequencing and imputation increases power for genome-wide association studies. Nature Genet. 44, 631–635 (2012)
Abecasis, G. R. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012)
Liao, S. C. et al. Low prevalence of major depressive disorder in Taiwanese adults: possible explanations and implications. Psychol. Med. 42, 1227–1237 (2012)
Lee, S. et al. The epidemiology of depression in metropolitan China. Psychol. Med. 39, 735–747 (2009)
Kessler, R. C. et al. The epidemiology of major depressive disorder: results from the National Comorbidity Survey Replication (NCS-R). J. Am. Med. Assoc. 289, 3095–3105 (2003)
Gerhart-Hines, Z. et al. Metabolic control of muscle mitochondrial function and fatty acid oxidation through SIRT1/PGC-1α. EMBO J. 26, 1913–1923 (2007)
Cai, N. et al. Molecular signatures of major depression. Curr. Biol. 25, 1146–1156 (2015)
Pruim, R. J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010)
Association, A. P. Diagnostic and statistical manual of mental disorders 4th edn (American Psychiatric Association, 1994)
Lunter, G. & Goodson, M. Stampy: a statistical algorithm for sensitive and fast mapping of Illumina sequence reads. Genome Res. 21, 936–939 (2011)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
Wang, Y. et al. An integrative variant analysis pipeline for accurate genotype/haplotype inference in population NGS data. Genome Res. 23, 833–842 (2013)
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007)
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007)
R Development Core Team. A language and environment for statistical computing (R Foundation for Statistical Computing, 2004)
Dudbridge, F. Power and predictive accuracy of polygenic risk scores. PLoS Genet. 9, e1003348 (2013)
Acknowledgements
This work was funded by the Wellcome Trust (WT090532/Z/09/Z, WT083573/Z/07/Z, WT089269/Z/09/Z), NIH grant MH-100549 and the Brain and Behavior Research Foundation. All authors are part of the CONVERGE consortium (China, Oxford and VCU Experimental Research on Genetic Epidemiology) and gratefully acknowledge the support of all partners in hospitals across China. W. Kretzschmar is funded by the Wellcome Trust (WT097307). N. Cai is supported by the Agency of Science, Technology and Research (A*STAR) Graduate Academy. J. Marchini is funded by an ERC Consolidator Grant (617306). Q. Xu is funded by the 973 Program (2013CB531301) and NSFC (31430048, 31222031).
Author information
Authors and Affiliations
Consortia
Contributions
Manuscript preparation: N. Cai, T. B. Bigdeli, W. Kretzschmar, M. Reimers, T. Webb, B. Riley, S. Bacanu, R. E. Peterson, K. S. Kendler and J. Flint. Replication sample: Q. Xu CONVERGE sample collection: Yih. Li, Y. Chen, H. Deng, W. Sang, Ke. Li, J. Gao, B. Ha, S. Gao, J. Hu, C. Hu, G. Huang, G. Jiang, X. Zhou, You. Li, Kan Li, Q. Niu, Yi Li, G. Li, L. Liu, Z. Liu, Yi Li, X. Fang, R. Pan, G. Miao, Q. Zhang, F. Yu, G. Chen, M. Cai, D. Yang, X. Hong, Y. Song, C. Gao, J. Pan, Y. Zhang, T. Liu, J. Dong, X. Wang, L. Wang, Q. Mei, Z. Shen, X. Liu, W. Wu, D. Gu, Y. Chen, T. Liu, H. Rong, Yi. Liu, L. Lv, H. Meng, H. Sang, J. Shen, T. Tian, J. Shi, J. Sun, M. Tao, X. Wang, J. Xia, Q. He, G. Wang, X. Wang, Lina Yang, K. Zhang, N. Sun, J. Zhang, Z. Gan, Z. Zhang, W. Zhang, H. Zhong, F. Yang, E. Cong, S. Shi, G. Fu, J. Flint and K. S. Kendler. Genome sequencing and analysis: J. Liang, J. Hu, Q. Li, W. Jin, Z. Hu, G. Wang, Linm. Wang, P. Qian, Yu. Liu, T. Jiang, Y. Lu, X. Zhang, Y. Yin, Yin. Li, H. Yang, Jia. Wang, X. Gan, Yih. Li, N. Cai, R. Mott, J. Flint, Jun Wang and X. Xu. Genotype imputation: W. Kretzschmar, J. Hu, L. Song, Q. Li, N. Cai and J. Marchini. Genetic analysis: N. Cai, T. Bigdeli, Yih. Li, R. E. Peterson, S. Bacanu, T. Webb, B. Riley, K. S. Kendler, R. Mott and J. Flint.
Corresponding authors
Ethics declarations
Competing interests
The author declare no competing financial interests.
Additional information
All sequence data and MDD results are freely available at http://dx.doi.org/10.5524/100155. GWAS results are also available at http://www.med.unc.edu/pgc/downloads.
Extended data figures and tables
Extended Data Figure 1 Quantile–quantile plots for major depressive disorder.
Quantile–quantile plot of GWAS for MDD using the mixed linear model with exclusion of the chromosome that the marker is on (MLMe) method implemented in FastLMM on 10,640 samples (5,303 cases, 5,337 controls). Genomic inflation factor λ = 1.070, rescaled for an equivalent study of 1,000 cases and 1,000 controls (λ 1000) = 1.013.
Extended Data Figure 2 Forest plots of estimated SNP effects in CONVERGE and PGC studies.
This figure presents the association odds ratios (OR) at 12 SNPs in CONVERGE and the best available proxy SNPs in PGC-MDD (pairwise r 2 > 0.6, 500 kb window; the proxy SNP is marked by an asterisk). We present the alternative allele frequency (freq), odds ratio (or) with respect to the alternative allele, standard error of odds ratio (se) and P values of association (pval) for the following analyses (study): primary association analysis with a linear-mixed model using imputed allele dosages in 10,640 samples in CONVERGE (pri); validation analysis with logistic regression model with principal components (PCs) as covariates using genotypes from Sequenom on 9,921 samples in CONVERGE (sqnm); association with MDD with a logistic regression model in a replication cohort of 6,417 samples using genotypes from Sequenom (repli); joint association analysis with MDD with a logistic regression model using imputed allele dosages in CONVERGE and genotypes from Sequenom in a replication cohort (17,057 samples in total; joint).
Extended Data Figure 3 Manhattan and quantile quantile plots for melancholia.
a, Manhattan plot of GWAS for melancholia using the MLMe method implemented in FastLMM on 9,846 samples (4,509 cases, 5,337 controls). b, Quantile–quantile plot of GWAS for melancholia; λ = 1.069, λ 1000 = 1.014. c, Regional association plot of GWAS hits on chromosome 10, focusing on top SNP rs80309727 at 5′ of SIRT1 gene, generated with LocusZoom.
Extended Data Figure 4 Empirical estimation of the odds ratio increases due to the removal of cases not falling under the diagnostic class of melancholia from an association analysis with major depression.
The figures show the empirical distributions of the odds ratios for association with each of two SNPs (rs79804696, rs35936514), after removing a random set of 796 samples, equal to the number of cases of MDD not diagnosed as being melancholic. The horizontal axis is the odds ratio for each analysis, and the vertical axis the frequency of occurrence of the odds ratio in 10,000 analyses. The vertical red line is the observed odds ratio after removing cases of MDD not diagnosed as melancholic.
Supplementary information
Supplementary Information
This file contains Supplemental Notes 1-2, Supplementary References and Supplementary Tables 1-3. (PDF 478 kb)
Supplementary Information
This file contains Supplementary Table 4, a list of SNPs associated with MDD with P values < 10-5. (XLSX 60 kb)
Supplementary Information
This file contains Supplementary Table 5, a list of SNPs associated with Melancholia with P values < 10-5. (XLSX 109 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
CONVERGE consortium. Sparse whole-genome sequencing identifies two loci for major depressive disorder. Nature 523, 588–591 (2015). https://doi.org/10.1038/nature14659
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14659
This article is cited by
-
Emerging roles of SIRT1 activator, SRT2104, in disease treatment
Scientific Reports (2024)
-
Neuroimaging Genomics a Predictor of Major Depressive Disorder (MDD)
Molecular Neurobiology (2024)
-
Association between 14 candidate genes, PM2.5, and affective disorders: a study of the Taiwan Biobank
BMC Public Health (2023)
-
Similarity and diversity of genetic architecture for complex traits between East Asian and European populations
BMC Genomics (2023)
-
Reduced Vrk2 expression is associated with higher risk of depression in humans and mediates depressive-like behaviors in mice
BMC Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.