Abstract
Mitochondria are involved in a variety of cellular functions, including ATP production, amino acid and lipid biogenesis and breakdown, signalling and apoptosis1,2,3. Mitochondrial dysfunction has been linked to neurodegenerative diseases, cancer and ageing4. Although transcriptional mechanisms that regulate mitochondrial abundance are known5, comparatively little is known about how mitochondrial function is regulated. Here we identify the metabolite stearic acid (C18:0) and human transferrin receptor 1 (TFR1; also known as TFRC) as mitochondrial regulators. We elucidate a signalling pathway whereby C18:0 stearoylates TFR1, thereby inhibiting its activation of JNK signalling. This leads to reduced ubiquitination of mitofusin via HUWE1, thereby promoting mitochondrial fusion and function. We find that animal cells are poised to respond to both increases and decreases in C18:0 levels, with increased C18:0 dietary intake boosting mitochondrial fusion in vivo. Intriguingly, dietary C18:0 supplementation can counteract the mitochondrial dysfunction caused by genetic defects such as loss of the Parkinson’s disease genes Pink or Parkin in Drosophila. This work identifies the metabolite C18:0 as a signalling molecule regulating mitochondrial function in response to diet.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
02 September 2015
A figure citation to ‘Extended Data Fig. 1h, i’ was corrected.
References
McBride, H. M., Neuspiel, M. & Wasiak, S. Mitochondria: more than just a powerhouse. Curr. Biol. 16, R551–R560 (2006)
Vafai, S. B. & Mootha, V. K. Mitochondrial disorders as windows into an ancient organelle. Nature 491, 374–383 (2012)
Suen, D. F., Norris, K. L. & Youle, R. J. Mitochondrial dynamics and apoptosis. Genes Dev. 22, 1577–1590 (2008)
Nunnari, J. & Suomalainen, A. Mitochondria: in sickness and in health. Cell 148, 1145–1159 (2012)
Jornayvaz, F. R. & Shulman, G. I. Regulation of mitochondrial biogenesis. Essays Biochem. 47, 69–84 (2010)
Moon, Y. A., Ochoa, C. R., Mitsche, M. A., Hammer, R. E. & Horton, J. D. Deletion of ELOVL6 blocks the synthesis of oleic acid but does not prevent the development of fatty liver or insulin resistance. J. Lipid Res. 55, 2597–2605 (2014)
Matsuzaka, T. et al. Elovl6 promotes nonalcoholic steatohepatitis. Hepatology 56, 2199–2208 (2012)
Jung, A., Hollmann, M. & Schafer, M. A. The fatty acid elongase NOA is necessary for viability and has a somatic role in Drosophila sperm development. J. Cell Sci. 120, 2924–2934 (2007)
Fernandez-Ayala, D. J. et al. Expression of the Ciona intestinalis alternative oxidase (AOX) in Drosophila complements defects in mitochondrial oxidative phosphorylation. Cell Metab. 9, 449–460 (2009)
Friedman, J. R. & Nunnari, J. Mitochondrial form and function. Nature 505, 335–343 (2014)
Okamoto, K. & Shaw, J. M. Mitochondrial morphology and dynamics in yeast and multicellular eukaryotes. Annu. Rev. Genet. 39, 503–536 (2005)
Cassidy-Stone, A. et al. Chemical inhibition of the mitochondrial division dynamin reveals its role in Bax/Bak-dependent mitochondrial outer membrane permeabilization. Dev. Cell 14, 193–204 (2008)
Twig, G. et al. Tagging and tracking individual networks within a complex mitochondrial web with photoactivatable GFP. Am. J. Physiol. Cell Physiol. 291, C176–C184 (2006)
Campello, S. & Scorrano, L. Mitochondrial shape changes: orchestrating cell pathophysiology. EMBO Rep. 11, 678–684 (2010)
Liesa, M., Palacin, M. & Zorzano, A. Mitochondrial dynamics in mammalian health and disease. Physiol. Rev. 89, 799–845 (2009)
Escobar-Henriques, M. & Langer, T. Dynamic survey of mitochondria by ubiquitin. EMBO Rep. 15, 231–243 (2014)
Livnat-Levanon, N. & Glickman, M. H. Ubiquitin–proteasome system and mitochondria—reciprocity. Biochim. Biophys. Acta 1809, 80–87 (2011)
Ziviani, E., Tao, R. N. & Whitworth, A. J. Drosophila Parkin requires PINK1 for mitochondrial translocation and ubiquitinates Mitofusin. Proc. Natl Acad. Sci. USA 107, 5018–5023 (2010)
Leboucher, G. P. et al. Stress-induced phosphorylation and proteasomal degradation of mitofusin 2 facilitates mitochondrial fragmentation and apoptosis. Mol. Cell 47, 547–557 (2012)
de Groot, R. E. et al. Huwe1-mediated ubiquitylation of Dishevelled defines a negative feedback loop in the Wnt signaling pathway. Sci. Signal. 7, ra26 (2014)
Lee, S. et al. Mitofusin 2 is necessary for striatal axonal projections of midbrain dopamine neurons. Hum. Mol. Genet. 21, 4827–4835 (2012)
de Brito, O. M. & Scorrano, L. Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature 456, 605–610 (2008)
Ozcan, U. et al. Chemical chaperones reduce ER stress and restore glucose homeostasis in a mouse model of type 2 diabetes. Science 313, 1137–1140 (2006)
Jian, J., Yang, Q. & Huang, X. Src regulates Tyr20 phosphorylation of transferrin receptor-1 and potentiates breast cancer cell survival. J. Biol. Chem. 286, 35708–35715 (2011)
Whitworth, A. J., Wes, P. D. & Pallanck, L. J. Drosophila models pioneer a new approach to drug discovery for Parkinson’s disease. Drug Discov. Today 11, 119–126 (2006)
Greene, J. C. et al. Mitochondrial pathology and apoptotic muscle degeneration in Drosophila parkin mutants. Proc. Natl Acad. Sci. USA 100, 4078–4083 (2003)
Park, J. et al. Mitochondrial dysfunction in Drosophila PINK1 mutants is complemented by parkin . Nature 441, 1157–1161 (2006)
Park, J., Kim, Y. & Chung, J. Mitochondrial dysfunction and Parkinson’s disease genes: insights from Drosophila . Dis. Model. Mech. 2, 336–340 (2009)
Guo, M. Drosophila as a model to study mitochondrial dysfunction in Parkinson’s disease. Cold Spring Harb. Perspect. Med. 2, a009944 (2012)
Jump, D. B. Mammalian fatty acid elongases. Methods Mol. Biol. 579, 375–389 (2009)
Acknowledgements
We thank A. Zorzano and J. Seco for photo-activatable mitoGFP, A. Zorzano and J. Pablot for Myc-tagged MFN2, H. Jacobs for UAS-AOX flies, C. Frei for UAS-Spargel flies, P. Verstreken for Parkinson’s flies, A. Whitworth for anti-dMfn, J. Chung for anti-dPorin, R. Lill for anti-IBA57, J. Sagalés for help with respirometry on the dMfn-knockout animals, F. Melchior for advice on detecting ubiquitination, K. Richter and the DKFZ EM Core Facility for help with electron microscopy, the DKFZ Light Microscopy Core Facility for help with live-cell imaging, M. Schnölzer and the DKFZ Proteomics Core Facility for mass spectrometry analysis of proteins, and M. del Sol for technical support. This work was supported in part by Deutsche Forschungsgemeinschaft (DFG), SFB1118, Helmholtz Portfolio Topic ‘Metabolic Dysfunction’, and European Research Council Starting Grants (#260602) to A.A.T., and Biotechnology and Biological Sciences Research Council, Medical Research Council programme and British Heart Foundation grants to A.V.-P.
Author information
Authors and Affiliations
Contributions
All authors designed experiments. D.S., S.V., X.X. and C.Y.T. performed experiments. All authors analysed data. D.S., S.V., A.K.M., A.V.-P. and A.A.T. wrote the manuscript and all authors read and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 noa/Elovl6 is the functional homologue of human ELOVL6.
a, C16:0 to C18:0 elongase activity is significantly blunted in Elovl6 mutants, whereas elongase activities on other fatty acids measured are not affected. Microsomal preparations from control or Elovl6 mutant animals were incubated with radioactive malonyl-CoA and the indicated fatty-acyl-CoA. Elongation was quantified by incorporation of the aqueous metabolite malonyl-CoA into lipid-soluble fatty acids, as described previously30. Values represent biological triplicates. b, Gas chromatography flame ionization detector (GC-FID) analysis reveals that Elovl6 mutant larvae have reduced levels of C18:0, the product of Elovl6 elongase activity. Values are the averages of technical duplicates on biological duplicates. Error bars represent standard error of the mean (s.e.m.). c, Lethality of Elovl6 mutants is fully rescued to expected Mendelian ratios by ubiquitous expression (with actin-GAL4) of Elovl6 from a UAS transgene (χ2 test, 1.149 < 3.841 = χ2, where P = 0.05, n = 141; ****P < 0.0001). d, Human ELOVL6 and Drosophila Elovl6 are functionally equivalent, since the lethality of Elovl6 mutant flies is fully rescued to expected Mendelian ratios by ubiquitous expression (with actin-GAL4) of human ELOVL6 from a UAS transgene (χ2 test, 2.38 < 3.841 = χ2, where P = 0.05, n = 76; **P < 0.01). e, The lethality of Elovl6 mutant flies is most strongly rescued by C18:0, the product of Elovl6. Synchronized 1st instar larvae of indicated genotypes were grown on standard food supplemented with indicated fatty acids (5%). The percentage of total animals surviving to pupation was calculated. Values represent average of biological triplicates. f, Elovl6 mutants are not hypersensitivite to drugs such as G418 (protein biosynthesis inhibitor) or etoposide (topoisomerase inhibitor). Thirty synchronized L1 larvae were grown in vials with food supplemented with either G418 (50 µg ml−1) or etoposite (25 µM). Percentage of animals that reach pupation was quantified. Values represent average of four biological replicates. g, Complex IV activity of Elovl6− larvae is not impaired. Complex IV activity of female pre-wandering larvae was measured with Oroboros high-resolution respirometry. Oxygen consumption was measured in the presence of only N,N,N′,N′-tetramethyl-p-phenylenediamine dihydrochloride (TMPD) as substrate, which can be directly oxidized by complex IV. The values were corrected for non-mitochondrial oxygen consumption (oxygen consumption in the presence of complex IV inhibitor potassium cyanide (KCN)) and normalized to tissue weight. n = 3. h, i, Overexpression of Spargel in Elovl6 mutant female pre-wandering larvae leads to increased mitochondrial abundance, assessed by porin levels (h; representative of six biological replicates) and citrate synthase activity (i; n = 4) in pre-wandering larvae. See Supplementary Fig. 12 for image of the uncropped full western blot. j, Drosophila Elovl6 (either N- or C-terminally tagged) localizes to the mitochondrial outer membrane. S2 cell lysates (‘total’) were successively fractionated to yield crude mitochondria (which include mitochondrial-associated membranes (MAMs)), pure mitochondria (lacking MAMs), and mitochondrial outer membranes (OM), inner membranes (IM) and inter-membrane space (IMS). Endogenous porin and ATPsyn-α were used as positive controls for outer membranes and inner membranes, respectively. 7.5 μg of protein from each fraction was loaded per lane. See Supplementary Fig. 13 for image of the uncropped full western blot. Representative of two biological replicates. k, Lipidomic analysis of standard fly food reveals low levels of C18:0 in the food. a, c, d, e, f, g, i, Error bars represent s.d. a, b, e–g, i, *P < 0.05, **P < 0.01, not significant (NS) P ≥ 0.05, two-tailed t-test.
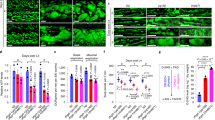
Extended Data Figure 2 C18:0 regulates mitochondrial morphology.
a, Lipidomic (GC-FID) profiles of purified mitochondria from Elovl6 mutant 3rd instar larvae do not show major differences compared with control animals. Mitochondrial membranes from both control and mutant animals have very low levels of C18:0. Controls for purity of mitochondrial prep are shown in Extended Data Fig. 6b. b, c, Elovl6 mutant larvae do not have reduced amounts of mitochondria, quantified via levels of porin (b; representative of three biological replicates) or citrate synthase activity (c; n = 3). See Supplementary Fig. 14 for image of the uncropped full western blot. d, Elovl6 mutant larvae have fragmented mitochondria, which is rescued by dietary C18:0 supplementation. Mitochondrial morphology from fat bodies of control or Elovl6 mutant female larvae, fed control or C18:0 (10%) supplemented food, visualized with mitoGFP. Images are representative of eight areas of four larvae from each genotype and food conditions. Equivalent pictures for body wall are shown in Fig. 2a. e, Only C18:0, and not shorter, longer or desaturated fatty acids, restores mitochondrial fragmentation to control levels in HeLa cells grown in medium containing delipidated serum. Mitochondria were visualized with MitoTracker (red) (top) and mitochondrial fragmentation was quantified by normalizing the number of mitochondrial particles to total mitochondrial area (bottom) (n = 15). f, Reduced lipoic acid (LA) levels do not lead to mitochondrial fragmentation. Lipoic acid synthase (LIAS) was knocked down by RNAi in HeLa cells, leading to significantly reduced lipoic acid levels, assayed by immunoblotting of total cell lysates with antibody detecting lipoic acid (bottom left). Unlike removal of C18:0, this does not lead to mitochondrial fragmentation. Representative images (top left) are quantified (top right) (n = 6). See Supplementary Fig. 14 for image of the uncropped full western blot. g, h, HeLa cells growing in medium containing delipidated serum do not display reduced levels of protein lipoylation (g) or reduced levels of lipoylated proteins (h). HeLa cells were grown in medium containing delipidated serum for either 24 h (the same time point used for all other experiments in which mitochondrial fragmentation was assessed) (g, h), or for an extended period of time: 4 days (h). Lipoic acid levels were assayed by immunobloting total cell lysates with an anti-lipoic acid antibody (g), and levels of lipoylated proteins were assessed with specific antibodies (h). See Supplementary Fig. 15 for image of the uncropped full western blot. c, e, f, *P < 0.05, **P < 0.01, not significant (NS) P ≥ 0.05, two-tailed t-test. Error bars represent s.d. Scale bars, 10 µm.
Extended Data Figure 3 Mitofusin loss of function phenocopies Elovl6 mutation or removal of C18:0.
a, dMfn knockout larvae (1st instar) have fragmented mitochondria, visualized with mitoGFP. Representative of ten images. b, Endogenous dMfn runs as a main band plus a laddering of apparently increasing molecular weights on an SDS–PAGE gel. Specificity is controlled by blotting lysates from control and Mfn-knockout female larvae with anti-Mfn antibody. c, Homozygous mutation of dMfn is lethal. Mfn-knockout larvae survive for several days as small L1/L2 larvae and eventually die. Synchronized 1st instar larvae were grown on standard fly food and examined every 24 h for developmental stage and per cent survival (n = 30). d, dMfn-knockout animals have impaired oxygen consumption. Oxygen consumption of inverted, digitonin permeabilized, female larval tissues was measured with an Oroboros oxygraph chamber and normalized to tissue weight. Oxygen consumption was measured in the presence of the following substrates: GMN (glutamate and malate), GMD (glutamate, malate and ADP), GMcD (glutamate, malate, cytochrome c and ADP), GMScD (glutamate, malate, succinate, cytochrome c and ADP), ETS (glutamate, malate, cytochrome c, ADP and uncoupling reagent), and Sc(Rot)u (glutamate, malate, cytochrome c, ADP and rotenone). n = 5. e, Endogenous dMfn is post-translationally modified in a C18:0-dependent manner in Drosophila. dMfn from Elovl6 female mutants migrates in an SDS–PAGE gel differently, compared with Mfn2 from control animals. This is reversed by supplementing the diet with C18:0. All indicated bands are dMfn, since they disappear in lysates from dMfn-knockout animals (see Extended Data Fig. 3b). Flies were grown on antifungal-free food. f, Endogenous MFN2 is post-translationally modified in a C18:0-dependent manner in human HeLa cells. MFN2 immunoprecipitated from HeLa cells treated for 24 h with medium containing standard or delipidated serum, and then for 2 h in the absence or presence of C18:0 (100 µM), lysed in 8M urea (see Methods). g, C18:0 affects ubiquitination of MFN2. MFN2 is more heavily ubiquitinated in cells treated with delipidated serum than in control cells and this is reversed by supplementing the medium with C18. HeLa cells were cotransfected with tagged versions of MFN2 (myc) and ubiquitin (HA). Tagged MFN2 was immunoprecipitated and blots were probed with HA antibody to detect ubiquitination. Quantification of ubiquitination, normalized to Myc–MFN2 in the immunoprecipitate (IP) is shown below the lane. h, C18:0 removal destabilizes MFN2 protein. A cyclohexamide (CHX) chase experiment was performed to block de novo synthesis of MFN2, thereby looking at turnover of existing MFN2 protein in vivo. HeLa cells treated with medium containing delipidated serum plus or minus C18:0 were treated with 100 µM CHX and then lysed at the indicated time points to compare MFN2 protein levels. Bottom, densitometric quantification of the blots normalized to loading control. i, dMfn expression is upregulated in Elovl6− flies compared with controls. dMfn transcript levels in 24 h female pre-wandering larvae were determined by quantitative polymerase chain reaction with reverse transcription (RT–PCR), normalized to rp49 (in triplicates). Scale bar, 10 μm. d, i, *P < 0.05, **P < 0.01, ***P < 0.001, not significant (NS) P ≥ 0.05, two-tailed t-test. Error bars represent s.d. See Supplementary Fig. 16 for images of the uncropped full western blots.
Extended Data Figure 4 HUWE1 is required for hyperubiquitination of MFN2 in response to C18:0 withdrawal.
a, Short interfering RNA (siRNA) depletion of other ubiquitin ligases targeting MFN (besides HUWE1, shown in Fig. 3) does not rescue the mitochondrial fragmentation induced by removal of C18:0 (top). Bottom, quantification. n = 15. b, siRNA depletion of PARK2 in HEK293 cells, as in HeLa cells (a), does not rescue the mitochondrial fragmentation induced by removal of C18:0 (top). Bottom, quantification. n = 15. c, HUWE1 knockdown efficiency controlled by detecting HUWE1 protein levels. See Supplementary Fig. 17 for image of the uncropped full western blot. d, Survival to pupation of Elovl6 mutants is fully rescued by ubiquitous expression (daughterless-GAL4) of RNAi targeting Huwe1 (CG8184). Elovl6 mutants expressing HUWE1 RNAi survive to pupation at expected Mendelian frequencies (χ2 test, 0.86 < 3.841 = χ2, where P = 0.05). Flies were grown on antifungal-free food. Values represent average of four biological replicates. Scale bars, 10 μm. a, b, d, *P < 0.05, **P < 0.01, ***P < 0.001, not significant (NS) P ≥ 0.05, two-tailed t-test. Error bars represent s.d.
Extended Data Figure 5 Elovl6 mutants have other Mfn loss-of-function phenotypes such as reduced mitochondrial-associated membranes and abnormal cristae.
a, The mitochondrial associated membranes (MAM) band is strongly reduced or absent in Percoll gradients of crude mitochondrial fractions from Elovl6-mutant animals, compared with controls. b, Purity control of mitochondrial preparations show that pure mitochondrial fractions are lacking markers of other subcellular organelles such as calnexin (endoplasmic reticulum) and lamin (nuclei). See Supplementary Fig. 17 for image of the uncropped full western blot. Right, quantification shows that levels of the endoplasmic reticulum marker calnexin are reduced in crude mitochondrial fractions from Elovl6 mutants, compared with controls, in agreement with reduced MAMs in Elovl6 mutants. Values show densitometry ratios of calnexin levels in crude mitochondrial fractions, normalized to total lysate calnexin. c, Electron microscopy of Drosophila S2 cell mitochondria (left) reveals cristae abnormalities in Elovl6-depleted cells. Middle, quantification (n = 200). Significance of the difference was calculated with a Mann–Whitney test (*P < 0.05). Right, average circularity of mitochondria was calculated with ImageJ software. Scale bar, 1 µm. n = 200, ****P < 0.0001, two-tailed t-test. Error bars show s.d.
Extended Data Figure 6 C18:0 removal does not lead to endoplasmic reticulum stress, and inhibiting UPR does not inhibit mitochondrial fragmentation upon C18:0 removal.
a, C18:0 removal for 24 h does not induce expression of UPR target genes, quantified by quantitative RT–PCR, normalized to RPL19. BiP (also known as HSPA5) is a readout for IRE1 (also known as ERN1) activation, CHOP (also known as DDIT3) is a readout for ATF6 activation, and PERK (also known as EIF2AK3) is a readout of its own activation due to a positive transcriptional feedback loop. Tunicamycin serves as a positive control. y axis is displayed in a logarithmic scale to fit all data points on one graph. The experiment was done in triplicates. b, p-eIF2α, a UPR marker, does not increase upon removal of C18:0 whereas it is induced by tunicamycin, a positive control. See Supplementary Fig. 18 for image of the uncropped full western blot. c, Knocking down mediators of the UPR response does not inhibit mitochondrial fragmentation upon C18:0 removal. HeLa cells were transfected with either control siRNAs or siRNAs targeting UPR mediators as indicated. Left, the mitochondrial fragmentation index; right, representative images. n = 15. d, Inhibiting endoplasmic reticulum stress by means of a chemical chaperone, TUDCA, does not rescue mitochondrial fragmentation upon C18:0 removal. HeLa cells were pre-treated with 500 μg ml−1 TUDCA 30 min before delipidated serum treatment. Left, mitochondrial fragmentation index (n = 15); right, representative images. a, c, d, *P < 0.05, **P < 0.01, two-tailed t-test. Error bars show s.d.
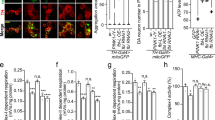
Extended Data Figure 7 TFR1 is the mediator of C18:0 signalling to mitochondrial morphology.
a, C17:0–azide is a functional analogue of C18:0 in that it induces mitochondrial fusion in HeLa cells, whereas other C18:0 derivatives are not. Cn:0–azide = HO2C(CH2) n − 1N3; Cn:0–alkyne = HO2C(CH2) n − 1CCH. b, TFR1 is the most enriched protein in a C17:0–azide pulldown, and it regulates mitochondrial morphology. HeLa cells were treated with C17:0–azide for 2 h, and covalently bound proteins were precipitated by lysing cells under denaturing conditions (8 M urea), and linking the C17:0–azide to an alkyne-labelled resin via click chemistry (left). Precipitated proteins were identified by mass spectrometry, and peptide counts were normalized to peptide counts in a negative control pulldown from cells not treated with C17:0–azide (n = 3) (right; column 2). Indicated proteins were also tested by siRNA-mediated knockdown for effects on mitochondrial morphology (column 3). c, TFR1 is covalently bound to the C18:0 derivative C17:0–azide in HeLa cells in a ZDHHC6-dependent manner. HeLa cells were treated with C17:0–azide for 2 h, and subsequently lysed in denaturing conditions (8 M urea). Similar to b, the C17:0–azide was ‘clicked’ onto a biotinylated alkyne, and the labelled proteins were pulled-down with streptavidin beads. After washing, immunoprecipitated proteins were eluted off beads in Laemmli buffer containing biotin, and analysed by immunoblotting. The palmitic acid analogue C15:0–azide was used as a positive control since TFR1 is known to also be palmitoylated. C17:0–azide pulls down more TFR1 than equal amounts of C15:0–azide, indicating that TFR1 palmitoylation cannot account for the C17:0 signal. The C17:0–azide–TFR1 interaction is completely blunted upon ZDHHC6 knockdown. See Supplementary Fig. 18 for image of the uncropped full western blot. d, TFR1 is required for C18:0 removal to induce mitochondrial fragmentation. HeLa cells were transfected with either control or TFR1 targeting siRNAs before treatment with medium containing delipidated serum plus or minus C18:0. Representative images are shown here and quantification of mitochondrial fragmentation is shown in Fig. 3h. n = 15.
Extended Data Figure 8 C18:0 removal does not affect iron uptake or delivery.
a, HeLa cells cannot grow in the presence of deferoxamine (DFO), an iron chelator (top) whereas they grow in delipidated serum lacking C18:0 at a comparable rate to cells in control medium (bottom). n = 3. b, Treatment of HeLa cells with medium containing delipidated serum (lacking C18:0) for 24 h does not lead to transcriptional activation of iron deficiency response genes (bottom), which are activated by DFO-mediated iron chelation (24 h) as a positive control (top). n = 3. c, Treatment of HeLa cells with medium containing delipidated serum for 24 h or 4 days does not lead to a drop in levels of succinate dehydrogenase b (SDHB), which contains an Fe–S cluster. See Supplementary Fig. 19 for image of the uncropped full western blot. d–f, Treatment of HeLa cells with medium containing delipidated serum for 24 h or 4 days does not lead to a drop in activities of enzymes containing lipoylated subunits (PDH and OGDH) (d, e) or Fe–S-cluster-containing subunits (SDH) (f). DFO treatment to chelate iron from the medium, or siRNA-mediated depletion of the enzymes were used as positive controls (d–f, bottom). n = 4. g, Treatment of HeLa cells with medium containing delipidated serum (24 h) does not cause a reduction in transferrin uptake. Cells were treated with 25 μg ml−1 Alexa-488-coupled transferrin for 30 min. Representative images (left) and quantification of the amount of transferrin per cell (right) (n = 5). h, Treatment of HeLa cells with medium containing delipidated serum (24 h) does not reduce association of transferrin-containing vesicles with mitochondria. Crude mitochondria were fractionated from cells growing in medium containing or lacking C18:0, and the amount of transferrin that copurifies with mitochondria was analysed and quantified by immunoblotting. See Supplementary Fig. 19 for image of the uncropped full western blot. a, b, d–g, *P < 0.05, ***P < 0.001, ****P < 0.0001, not significant (NS) P ≥ 0.05, two-tailed t-test. Error bars show s.d.
Extended Data Figure 9 JNK signalling is required for mitochondrial fragmentation induced by C18:0 removal.
a, Treatment of HeLa cells with 1 µM gambogic acid does not induce apoptosis. 10 μM gambogic acid was used as a positive control for apoptosis induction, assessed by cleaved caspase-3 levels. 1 µM gambogic acid neither induces caspase cleavage (shown here) nor causes cells to die (data not shown). Cells were treated with 10 µM gambogic acid for 1 h, or for all other concentrations for 3 h. See Supplementary Fig. 20 for image of the uncropped full western blot. b, Activation of TFR1 by treating cells with 1 µM gambogic acid leads to mitochondrial fragmentation that is reversed by 1 h C18:0 pre-treatment. Representative images are shown here and quantification of mitochondrial fragmentation is shown in Fig. 3i (n = 15). c, Treatment of HeLa cells with C18:0 to inhibit TFR1 causes reduced JNK signalling activity, assayed by p-JNK levels on an immunoblot. See Supplementary Fig. 20 for image of the uncropped full western blot. d, Removal of C18:0, as well as treatment with gambogic acid, induces shuttling of phosphorylated JNK into the nucleus. Cells were stained with phospho-JNK antibody (left) and relative levels of nuclear to cytosolic phospho-JNK signal was quantified (right) (n = 37 cells). ***P < 0.001, two-tailed t-test. Error bars show s.d. e, JNK signalling is required for TFR1 activation to induce mitochondrial fragmentation. HeLa cells were treated with the JNK inhibitor SP600125 30 min before gambogic acid treatment to activate TFR1. Representative images are shown here and quantification of mitochondrial fragmentation is shown in Fig. 3j (n = 15).
Extended Data Figure 10 Dietary C18:0 improves Parkinson’s disease phenotypes of Pink and Parkin mutant flies.
a, b, Dietary C18:0 supplementation (10%) significantly increases lifespan of male Parkin25 (a) and Pink1B9 (b) mutant flies. n = 8 × 10 animals. c, Dietary C18:0 supplementation rescues ATP levels of 1-week-old male Pink1B9 mutant adult flies. n = 3 × 3 animals. d, Dietary C18:0 supplementation significantly improves locomotor defects of 2-week-old male Pink1B9 mutant flies. Locomotion quantified as animals climbing up past a threshold in a given amount of time (technical duplicates, biological quadruplicates, ten animals per assay). e, Parkin loss of function in flies leads to mitochondrial fragmentation, which is rescued by dietary supplementation with C18:0. Guts from 14-day-old female control or park25 mutant adult flies expressing mitoGFP and grown on food supplemented with or without C18:0 (10%) were dissected and mitochondria were imaged. Quantification of mitochondrial fragmentation is shown (3 animals per condition, 6 optical areas per animal). b–d, Control flies are the revertant line Pink1RV . Error bars show s.d. Not significant (NS) P ≥ 0.05, *P < 0.05.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary Methods, Supplementary References and uncropped western blot images. (PDF 14269 kb)
Rights and permissions
About this article
Cite this article
Senyilmaz, D., Virtue, S., Xu, X. et al. Regulation of mitochondrial morphology and function by stearoylation of TFR1. Nature 525, 124–128 (2015). https://doi.org/10.1038/nature14601
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14601
This article is cited by
-
Proteomic analysis of mitochondria associated membranes in renal ischemic reperfusion injury
Journal of Translational Medicine (2024)
-
Mechanisms controlling cellular and systemic iron homeostasis
Nature Reviews Molecular Cell Biology (2024)
-
Identification and validation of a ferroptosis-related signature for prediction of the prognosis and tumor microenvironment in patients with chromophobe renal cell carcinoma
BMC Cancer (2023)
-
Low-carbohydrate diets containing plant-derived fat but not animal-derived fat ameliorate heart failure
Scientific Reports (2023)
-
Novel insights into the involvement of mitochondrial fission/fusion in heart failure: From molecular mechanisms to targeted therapies
Cell Stress and Chaperones (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.