Abstract
The current outbreak of Ebola virus in West Africa is unprecedented, causing more cases and fatalities than all previous outbreaks combined, and has yet to be controlled1. Several post-exposure interventions have been employed under compassionate use to treat patients repatriated to Europe and the United States2. However, the in vivo efficacy of these interventions against the new outbreak strain of Ebola virus is unknown. Here we show that lipid-nanoparticle-encapsulated short interfering RNAs (siRNAs) rapidly adapted to target the Makona outbreak strain of Ebola virus are able to protect 100% of rhesus monkeys against lethal challenge when treatment was initiated at 3 days after exposure while animals were viraemic and clinically ill. Although all infected animals showed evidence of advanced disease including abnormal haematology, blood chemistry and coagulopathy, siRNA-treated animals had milder clinical features and fully recovered, while the untreated control animals succumbed to the disease. These results represent the first, to our knowledge, successful demonstration of therapeutic anti-Ebola virus efficacy against the new outbreak strain in nonhuman primates and highlight the rapid development of lipid-nanoparticle-delivered siRNA as a countermeasure against this highly lethal human disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
World Health Organization. Ebola Data and Statistics; http://apps.who.int/gho/data/node.ebola-sitrep.quick-downloads?lang=en (2015)
Bishop, B. M. Potential and emerging treatment options for Ebola virus disease. Ann. Pharmacother. 49, 196–206 (2015)
Geisbert, T. W. et al. Postexposure protection of non-human primates against a lethal Ebola virus challenge with RNA interference: a proof-of-concept study. Lancet 375, 1896–1905 (2010)
Qiu, X. et al. Reversion of advanced Ebola virus disease in nonhuman primates with ZMapp. Nature 514, 47–53 (2014)
Qiu, X. et al. mAbs and Ad-vectored IFN-α therapy rescue Ebola-infected nonhuman primates when administered after the detection of viremia and symptoms. Sci. Transl. Med. 5, 207ra143 (2013)
Baize, S. et al. Emergence of Zaire Ebola virus disease in Guinea. N. Engl. J. Med. 371, 1418–1425 (2014)
Gire, S. K. et al. Genomic surveillance elucidates Ebola virus origin and transmission during the 2014 outbreak. Science 345, 1369–1372 (2014)
Kugelman, J. R. et al. Evaluation of the potential impact of ebola virus genomic drift on the efficacy of sequence-based candidate therapeutics. mBio 6, e02227–14 (2015)
Du, Q., Thonberg, H., Wang, J., Wahlestedt, C. & Liang, Z. A systematic analysis of the silencing effects of an active siRNA at all single-nucleotide mismatched target sites. Nucleic Acids Res. 33, 1671–1677 (2005)
Schwarz, D. S. et al. Designing siRNA that distinguish between genes that differ by a single nucleotide. PLoS Genet. 2, e140 (2006)
Huang, H. et al. Profiling of mismatch discrimination in RNAi enabled rational design of allele-specific siRNAs. Nucleic Acids Res. 37, 7560–7569 (2009)
Dallas, A. et al. Inhibition of hepatitis C virus in chimeric mice by short synthetic hairpin RNAs: sequence analysis of surviving virus shows added selective pressure of combination therapy. J. Virol. 88, 4647–4656 (2014)
Volchkova, V. A., Dolnik, O., Martinez, M. J., Reynard, O. & Volchkov, V. E. Genomic RNA editing and its impact on Ebola virus adaptation during serial passages in cell culture and infection of guinea pigs. J. Infect. Dis. 204 (suppl. 3). S941–S946 (2011)
Kugelman, J. R. et al. Ebola virus genome plasticity as a marker of its passaging history: a comparison of in vitro passaging to non-human primate infection. PLoS ONE 7, e50316 (2012)
Trefry, C. et al. In vivo pathological consequences of Ebola virus genome plasticity: is a 7U virus more lethal? ASM Biodefense Emerging Dis. Res. Meeting. Abstract 164. (2015)
Hirschberg, R. et al. Challenges, progress, and opportunities: proceedings of the filovirus medical countermeasures workshop. Viruses 6, 2673–2697 (2014)
Schieffelin, J. S. et al. Clinical illness and outcomes in patients with Ebola in Sierra Leone. N. Engl. J. Med. 371, 2092–2100 (2014)
Geisbert, T. W. et al. Treatment of Ebola virus infection with a recombinant inhibitor of factor VIIa/tissue factor: a study in rhesus monkeys. Lancet 362, 1953–1958 (2003)
Hensley, L. E. et al. Recombinant human activated protein C for the postexposure treatment of Ebola hemorrhagic fever. J. Infect. Dis. 196 (suppl. 2). S390–S399 (2007)
Honke, N. et al. Enforced viral replication activates adaptive immunity and is essential for the control of a cytopathic virus. Nature Immunol. 13, 51–57 (2011)
Ma, H. et al. Formulated minimal-length synthetic small hairpin RNAs are potent inhibitors of hepatitis C virus in mice with humanized livers. Gastroenterology 146, 63–66 (2014)
Acknowledgements
We thank V. Borisevich for assistance with clinical pathology assays performed in the GNL BSL-4 laboratory. We also thank S. Klassen for his assistance with siRNA-LNP preparation. This study was supported by the Department of Health and Human Services, National Institutes of Health grant U19AI109711 to T.W.G. and I.M.
Author information
Authors and Affiliations
Contributions
E.P.T. and J.Z.Z. designed the siRNA and did preparative work for dual luciferase reporter studies. E.P.T., N.M.S., J.Z.Z. and A.C.H.L. designed the dual luciferase reporter studies. J.Z.Z., T.R.B. and N.M.S. conducted the dual luciferase reporter studies and analysed the data. E.P.T., C.E.M., A.C.H.L., I.M. and T.W.G. conceived and designed the NHP study. C.E.M., J.B.G., D.J.D. and T.W.G. performed the NHP challenge and treatment experiments and conducted clinical observations of the animals. J.B.G., K.N.A. and D.J.D. performed the clinical pathology assays. J.B.G. performed the EBOV Makona infectivity assays. C.E.M. and K.N.A. performed the PCR assays. E.P.T., C.E.M., J.B.G., K.N.A., D.J.D., K.A.F., A.C.H.L., I.M. and T.W.G. analysed the data. K.A.F. performed histological and immunohistochemical analysis of the data. E.P.T., C.E.M., A.C.H.L. and T.W.G. wrote the paper. All authors had access to all of the data and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.C.H.L., I.M. and T.W.G. claim intellectual property regarding RNA interference for the treatment of filovirus infections. I.M. and T.W.G. are co-inventors on US Patent 7,838,658 ‘siRNA silencing of filovirus gene expression’ and A.C.H.L., I.M. and T.W.G. are co-inventors on US Patent 8,716,464 ‘Compositions and methods for silencing Ebola virus gene expression’.
Additional information
Opinions, interpretations, conclusions and recommendations are those of the authors and are not necessarily endorsed by the University of Texas Medical Branch.
Extended data figures and tables
Extended Data Figure 1 Antiviral activity of siEbola-3 in cells infected with EBOV Makona.
For comparison, siEbola-3 activity was also assessed against the central African EBOV Kikwit strain and siEbola-2 activity was evaluated against both EBOV strains. Data are viral RNA copies per milliltre of each sample normalized to untreated infected cells. Results are mean ± s.e.m. from one biological replicate, conducted in technical triplicate.
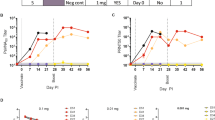
Extended Data Figure 2 siEbola-3 LNP treatment provides partial protection against EBOV Makona clinical pathologies, and infection with EBOV Makona infection induces a lesser degree of liver dysfunction compared to EBOV Kikwit infection.
a, No differences in viraemia levels were observed in untreated animals infected with EBOV Makona or Kikwit. b–e, Liver dysfunction markers. Normal values for uninfected NHPs ranges are GGT (40–115 U l−1), AST (20–45 U l−1), ALT (20–165 U l−1), ALP (130–500 U l−1). f, g, Protection against EBOV-Makona-induced CRE and BUN elevation was observed. Normal values for uninfected NHPs range from BUN (10–25 mg dl−1) and CRE (0.8–1.2 mg dl−1).
Extended Data Figure 3 Comparison of coagulation and haematology characteristics between untreated control animals infected with EBOV Makona or Kikwit.
a, b, Coagulopathies are not as marked in EBOV Makona infection when compared to historical EBOV Kikwit data. c, Lymphopenia is observed in all infected animals. d, Thrombocytopenia levels are similar between EBOV-Makona and EBOV-Kikwit-infected control animals.
Rights and permissions
About this article
Cite this article
Thi, E., Mire, C., Lee, A. et al. Lipid nanoparticle siRNA treatment of Ebola-virus-Makona-infected nonhuman primates. Nature 521, 362–365 (2015). https://doi.org/10.1038/nature14442
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14442
This article is cited by
-
Recent Advancements in the Therapeutic Development for Marburg Virus: Updates on Clinical Trials
Current Infectious Disease Reports (2024)
-
Natural history of nonhuman primates after conjunctival exposure to Ebola virus
Scientific Reports (2023)
-
Antiviral and protective effect of small interfering RNAs against rift valley fever virus in vitro
Molecular Biology Reports (2023)
-
Stimuli-Responsive Gene Delivery Nanocarriers for Cancer Therapy
Nano-Micro Letters (2023)
-
Meta-Analysis of Drug Delivery Approaches for Treating Intracellular Infections
Pharmaceutical Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.