Abstract
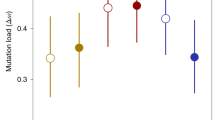
Reproduction through sex carries substantial costs, mainly because only half of sexual adults produce offspring1. It has been theorized that these costs could be countered if sex allows sexual selection to clear the universal fitness constraint of mutation load2,3,4. Under sexual selection, competition between (usually) males and mate choice by (usually) females create important intraspecific filters for reproductive success, so that only a subset of males gains paternity. If reproductive success under sexual selection is dependent on individual condition, which is contingent to mutation load, then sexually selected filtering through ‘genic capture’5 could offset the costs of sex because it provides genetic benefits to populations. Here we test this theory experimentally by comparing whether populations with histories of strong versus weak sexual selection purge mutation load and resist extinction differently. After evolving replicate populations of the flour beetle Tribolium castaneum for 6 to 7 years under conditions that differed solely in the strengths of sexual selection, we revealed mutation load using inbreeding. Lineages from populations that had previously experienced strong sexual selection were resilient to extinction and maintained fitness under inbreeding, with some families continuing to survive after 20 generations of sib × sib mating. By contrast, lineages derived from populations that experienced weak or non-existent sexual selection showed rapid fitness declines under inbreeding, and all were extinct after generation 10. Multiple mutations across the genome with individually small effects can be difficult to clear, yet sum to a significant fitness load; our findings reveal that sexual selection reduces this load, improving population viability in the face of genetic stress.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
24 June 2015
Minor changes were made to author affiliation number 4.
References
Maynard Smith, J. The Evolution of Sex (Cambridge Univ. Press, 1978)
Agrawal, A. F. Sexual selection and the maintenance of sexual reproduction. Nature 411, 692–695 (2001)
Siller, S. Sexual selection and the maintenance of sex. Nature 411, 689–692 (2001)
Whitlock, M. C. & Agrawal, A. F. Purging the genome with sexual selection: reducing mutation load through selection on males. Evolution 63, 569–582 (2009)
Tomkins, J. L., Radwan, J., Kotiaho, J. S. & Tregenza, T. Genic capture and resolving the lek paradox. Trends Ecol. Evol. 19, 323–328 (2004)
Andersson, M. B. Sexual Selection (Princeton Univ. Press, 1994)
Agrawal, A. F. & Whitlock, M. C. Mutation load: the fitness of individuals in populations where deleterious alleles are abundant. Annu. Rev. Ecol. Syst. 43, 115–135 (2012)
Holman, L. & Kokko, H. The consequences of polyandry for population viability, extinction risk and conservation. Phil. Trans. R. Soc. B 368, http://dx.doi.org/10.1098/rstb.2012.0053 (2013)
Crow, J. F. Mutation, mean fitness, and genetic load. Oxf. Surv. Evol. Biol. 9, 3–42 (1993)
Keightley, P. D. & Lynch, M. Toward a realistic model of mutations affecting fitness. Evolution 57, 683–685 (2003)
Haldane, J. B. S. The effect of variation of fitness. Am. Nat. 71, 337–349 (1937)
McVean, G. A. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012)
Darwin, C. On the Origin of Species by Means of Natural Selection, or the Preservation of Favoured Races in the Struggle for Life (John Murray, 1859)
Wigby, S. & Chapman, T. Female resistance to male harm evolves in response to manipulation of sexual conflict. Evolution 58, 1028–1037 (2004)
Michalczyk, Ł. et al. Experimental evolution exposes female and male responses to sexual selection and conflict in Tribolium castaneum . Evolution 65, 713–724 (2011)
Radwan, J. Effectiveness of sexual selection in removing mutations induced with ionizing radiation. Ecol. Lett. 7, 1149–1154 (2004)
Hollis, B., Fierst, J. L. & Houle, D. Sexual selection accelerates the elimination of a deleterious mutant in Drosophila melanogaster . Evolution 63, 324–333 (2009)
Almbro, M. & Simmons, L. W. Sexual selection can remove an experimentally induced mutation load. Evolution 68, 295–300 (2014)
Sharp, N. P. & Agrawal, A. F. Mating density and the strength of sexual selection against deleterious alleles in Drosphila melanogaster . Evolution 62, 857–867 (2008)
Arbuthnott, D. & Rundle, H. D. Sexual selection is ineffectual or inhibits the purging of deleterious mutations in Drosophila melanogaster . Evolution 66, 2127–2137 (2012)
Hollis, B. & Houle, D. Populations with elevated mutation load do not benefit from the operation of sexual selection. J. Evol. Biol. 24, 1918–1926 (2011)
Holland, B. & Rice, W. R. Experimental removal of sexual selection reverses intersexual antagonistic coevolution and removes a reproductive load. Proc. Natl Acad. Sci. USA 96, 5083–5088 (1999)
Charlesworth, B. & Charlesworth, D. The genetic basis of inbreeding depression. Genet. Res. 74, 329–340 (1999)
Rice, W. R. Sex chromosomes and the evolution of sexual dimorphism. Evolution 38, 735–742 (1984)
Kim, H. S. et al. BeetleBase in 2010: revisions to provide comprehensive genomic information for Tribolium castaneum . Nucleic Acids Res. 38, D437–D442 (2010)
Hollis, B., Houle, D., Yan, Z., Kawecki, T. J. & Keller, L. Evolution under monogamy feminizes gene expression in Drosophila melanogaster . Nature Commun. 5, http://dx.doi.org/10.1038/ncomms4482 (2014)
Long, A. F. T., Agrawal, A. F. & Rowe, L. The effect of sexual selection on offspring fitness depends on the nature of genetic variation. Curr. Biol. 22, 204–208 (2012)
Michalczyk, Ł. et al. Inbreeding promotes female promiscuity. Science 333, 1739–1742 (2011)
Lande, R. Genetics and demography in biological conservation. Science 241, 1455–1460 (1988)
Gilpin, M. E. & Soulé, M. E. in Conservation Biology: The Science of Scarcity and Diversity (ed. Soulé, M. E.) 19–34 (Sinauer Associates, 1986)
Demont, M. et al. Experimental removal of sexual selection reveals adaptations to polyandry in both sexes. Evol. Biol. 41, 62–70 (2014)
Falconer, D. S. & Mackay, T. F. C. Introduction to Quantitative Genetics 4th edn (Pearson Education, 1996)
Therneau, T. A package for survival analysis in S. R package version 2.37-7. http://cran.r-project.org/web/packages/survival/index.html (2014)
R Core Team. R: a language and environment for statistical computing (R Foundation for Statistical Computing, 2013)
Fournier, D. A. et al. AD Model Builder: using automatic differentiation for statistical inference of highly parameterized complex nonlinear models. Optimiz. Meth. Software 27, 233–249 (2012)
Excoffier, L. & Lischer, H. E. L. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567 (2010)
Bates, D., Maechler, M., Bolker, B. & Walker, S. lme4: Linear mixed-effects models using Eigen and S4. R package version 1.1-6. http://CRAN.R-project.org/package=lme4 (2014)
Kuznetsova, A., Brockhoff, P. B. & Christensen, R. H. B. lmerTest: Tests for random and fixed effects for linear mixed effect models (lmer objects of lme4 package). R package version 2.0-6. http://CRAN.R-project.org/package=lmerTest (2014)
Acknowledgements
We thank the Natural Environment Research Council and the Leverhulme Trust for financial support, D. Edward for statistical advice and colleagues at the 2013 Biology of Sperm meeting for comments that improved analytical design and interpretation.
Author information
Authors and Affiliations
Contributions
Ł.M., O.Y.M. and M.J.G.G. initiated the experimental evolution lines used in this work in 2005 and, with A.J.L., have maintained them since. M.J.G.G., Ł.M. and A.J.L. conceived, designed, conducted and analysed the study, with input from B.C.E. and T.C. J.J.N.K. and L.G.S. ran the microsatellite analyses. J.L.G., M.E.D. and O.Y.M. helped with line maintenance and experimental data collection. C.A.M. performed the fitness analyses. M.J.G.G. and A.J.L. wrote the paper, with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Data sets for all experiments and assays have been deposited in the Dryad Digital Repository at http://dx.doi.org/10.5061/dryad.86750.
Extended data figures and tables
Extended Data Figure 1 Experimental rationale for purging and then exposing mutation load.
Having been changed by strong (+SS, red) versus weak (−SS, blue) histories of sexual selection, while under equal influences of natural selection (NS), variation in mutation load residing in the form of recessive alleles is exposed via inbreeding. Inbreeding was enforced through monogamous sib × sib pairings, also eliminating concurrent confounds of interlocus sexual conflict. Populations with reduced mutation load as a result of histories of strong sexual selection are predicted to resist extinction (survival, s) and maintain fitness (f) under continuous inbreeding (i).
Extended Data Figure 2 Experimental evolution protocols for regimes A and B.
Contrasting intensities of strong (red) versus weak (blue) sexual selection were imposed upon each generation of adult reproduction, while equalizing effective population size within a regime, and allowing full genetic mixing within the replicate lines at the egg/larval/pupal stages. From the start, each treatment was replicated to create three independent lines. Regime A (a) applied contrasting sexual selection by varying adult operational sex ratio, while regime B (b) enforced monogamy to compare against polyandry.
Extended Data Figure 3 Extinction and fitness decline protocols.
Inbreeding in family lines was performed via sib × sib crosses for up to 20 generations across 3 years. To measure extinction (a), a family was considered extinct when it failed to produce offspring, or offspring were of the same sex (which occurred in only 9 out of 216 family lines, indicating no sex-specific pre-adult mortality by treatment). In regime A, extinction data were collected from 28 initial families per line, three lines per sexual selection treatment, comparing both strong versus weak treatments (n = 168 total family lines). In regime B, extinction data were collected from eight initial families per line, three lines per sexual selection treatment, comparing both strong versus weak treatments (n = 48 total family lines). To measure fitness decline (b), two additional sib × sib pairs per family per generation were bred to estimate reproductive fitness in every generation by counting number of offspring produced (see Methods). In both regimes A and B, fitness data were collected from eight initial families per line, three lines per sexual selection treatment, and both strong versus weak treatment contrasts in each.
Extended Data Figure 4 Estimated heterozygosity (±s.e.m.) does not differ between experimental evolution sexual selection treatments within regime A (left) and regime B (right).
Linear mixed effect modelling showed the estimated heterozygosity of the male-biased selection treatment (M: Hest = 0.312, t = 9.468) is not significantly different from that of female-biased (F: Hest = 0.318, t = 9.295, P = 0.863), but is significantly different from monogamous and polyandrous treatments (Mo: Hest = 0.199, t = 6.453, P = 0.003; Po: Hest = 0.197, t = 6.397, P = 0.003). The estimated heterozygosities of monogamous and polyandrous treatments are not significantly different (P = 0.956) (see Methods).
Extended Data Figure 5 Concordance between raw data and model fit in extinction analyses.
Survival curves of raw data (thick and dotted lines) overlaid on model fit (shaded areas with mean curves and 95% confidence intervals). Survival of families derived from strong (red, solid line) or weak (blue, dotted line) sexual selection treatment histories differed: (a) regime A, male-biased (red) versus female-biased (blue) sexual selection treatments; (b) regime B, polyandrous (red) versus monogamous (blue); (c) regimes A and B combined into a single analysis. See Fig. 1 and the main text for results of statistical analyses, and Methods and Extended Data Figs 2 and 3 for details of protocols, methods and experimental design.
Extended Data Figure 6 Regime A.
Boxplots of the relationships between fitness and inbreeding generation for the male-biased (a and c) versus the female-biased (b and d) treatments. Curves show the predicted relationships between reproductive fitness and inbreeding generation from the GLMMs, and the narrow red and blue shadows show the 95% confidence intervals predicted from the fixed effects. Horizontal bars indicate medians, boxes indicate interquartile ranges, whiskers indicate minimum and maximum values, and circles indicate outliers (values 1.5 times higher or lower than the first and third quartiles, respectively). Comparison of a versus b identifies the difference in total fitness declines between strong versus weak sexual selection histories in regime A, while c versus d identifies the same difference in decline for fitness but only for the sibling pairs that produced at least some offspring (that is, omitting zero fitness values that may have resulted from a failure to mate). See Fig. 1, main text and Extended Data Table 1 for results of statistical analyses.
Extended Data Figure 7 Regime B.
Boxplots of the relationships between fitness and inbreeding generation for the polyandrous (a and c) versus the monogamous (b and d) treatments. Curves show the predicted relationships between reproductive fitness and inbreeding generation from the GLMMs, and the narrow red and blue shadows show the 95% confidence intervals predicted from the fixed effects. Horizontal bars indicate medians, boxes indicate interquartile ranges, whiskers indicate minimum and maximum values, and circles indicate outliers (values 1.5 times higher or lower than first and third quartiles, respectively). Comparison of a versus b identifies the difference in total fitness declines between strong versus weak sexual selection histories in regime B, while c versus d identifies the same difference in decline for fitness but only for the sibling pairs that produced at least some offspring (that is, omitting zero fitness values that may have resulted from a failure to mate). See Fig. 1, main text and Extended Data Table 1 for results of statistical analyses.
Extended Data Figure 8 Across 7 days of mating opportunity, males successfully inseminated 50 females on average (±s.e.m.).
Six virgin females were allocated to individual GA1 control stock males (n = 11) every 12 h for 7 days, providing males with 84 potential mates. Over this 1 week period (replicating that applied within the experimental evolution protocols, Extended Data Fig. 2), males successfully inseminated and generated offspring from an average of 50 females (see Methods).
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data, Supplementary Tables 1-2 and additional references. (PDF 1015 kb)
Supplementary Data
This zipped file contains the following: Figure 1 R analysis script (Extinction Script.R); Figure 2 R analysis script (Fitness Script.R) and Extended data Figure 4 R analysis script (Heterozygosity Script.R). (ZIP 5 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Lumley, A., Michalczyk, Ł., Kitson, J. et al. Sexual selection protects against extinction. Nature 522, 470–473 (2015). https://doi.org/10.1038/nature14419
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14419
This article is cited by
-
Genomic evidence that a sexually selected trait captures genome-wide variation and facilitates the purging of genetic load
Nature Ecology & Evolution (2022)
-
Sexual selection for males with beneficial mutations
Scientific Reports (2022)
-
The origination events of gametic sexual reproduction and anisogamy
Journal of Ethology (2022)
-
Natural selection increases female fitness by reversing the exaggeration of a male sexually selected trait
Nature Communications (2021)
-
Tribolium beetles as a model system in evolution and ecology
Heredity (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.