Abstract
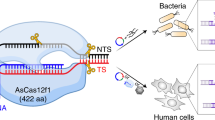
The RNA-guided endonuclease Cas9 has emerged as a versatile genome-editing platform. However, the size of the commonly used Cas9 from Streptococcus pyogenes (SpCas9) limits its utility for basic research and therapeutic applications that use the highly versatile adeno-associated virus (AAV) delivery vehicle. Here, we characterize six smaller Cas9 orthologues and show that Cas9 from Staphylococcus aureus (SaCas9) can edit the genome with efficiencies similar to those of SpCas9, while being more than 1 kilobase shorter. We packaged SaCas9 and its single guide RNA expression cassette into a single AAV vector and targeted the cholesterol regulatory gene Pcsk9 in the mouse liver. Within one week of injection, we observed >40% gene modification, accompanied by significant reductions in serum Pcsk9 and total cholesterol levels. We further assess the genome-wide targeting specificity of SaCas9 and SpCas9 using BLESS, and demonstrate that SaCas9-mediated in vivo genome editing has the potential to be efficient and specific.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
BioProject
Data deposits
All reagents described in this manuscript have been deposited with Addgene (plasmid IDs 61591, 61592 and 61593). Source data are available online and deep sequencing data are available at Sequence Read Archive under BioProject accession number PRJNA274149.
References
Bolotin, A., Quinquis, B., Sorokin, A. & Ehrlich, S. D. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151, 2551–2561 (2005)
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007)
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010)
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602–607 (2011)
Sapranauskas, R. et al. The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res. 39, 9275–9282 (2011)
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012)
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl Acad. Sci. USA 109, E2579–E2586 (2012)
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013)
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013)
Gaudet, D. et al. Review of the clinical development of alipogene tiparvovec gene therapy for lipoprotein lipase deficiency. Atheroscler. Suppl. 11, 55–60 (2010)
Vasileva, A. & Jessberger, R. Precise hit: adeno-associated virus in gene targeting. Nature Rev. Microbiol. 3, 837–847 (2005)
Mingozzi, F. & High, K. A. Therapeutic in vivo gene transfer for genetic disease using AAV: progress and challenges. Nature Rev. Genet. 12, 341–355 (2011)
Gao, G., Vandenberghe, L. H. & Wilson, J. M. New recombinant serotypes of AAV vectors. Curr. Gene Ther. 5, 285–297 (2005)
Kay, M. A. State-of-the-art gene-based therapies: the road ahead. Nature Rev. Genet. 12, 316–328 (2011)
Zincarelli, C., Soltys, S., Rengo, G. & Rabinowitz, J. E. Analysis of AAV serotypes 1–9 mediated gene expression and tropism in mice after systemic injection. Mol. Ther. 16, 1073–1080 (2008)
Swiech, L. et al. In vivo interrogation of gene function in the mammalian brain using CRISPR-Cas9. Nature Biotechnol. 33, 102–106 (2015)
Senís, E. et al. CRISPR/Cas9-mediated genome engineering: an adeno-associated viral (AAV) vector toolbox. Biotechnol. J. 9, 1402–1412 (2014)
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014)
Chylinski, K., Makarova, K. S., Charpentier, E. & Koonin, E. V. Classification and evolution of type II CRISPR-Cas systems. Nucleic Acids Res. 42, 6091–6105 (2014)
Chylinski, K., Le Rhun, A. & Charpentier, E. The tracrRNA and Cas9 families of type II CRISPR-Cas immunity systems. RNA Biol. 10, 726–737 (2013)
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014)
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nature Biotechnol. 31, 827–832 (2013)
Hou, Z. et al. Efficient genome engineering in human pluripotent stem cells using Cas9 from Neisseria meningitidis. Proc. Natl Acad. Sci. USA 110, 15644–15649 (2013)
Fu, Y., Sander, J. D., Reyon, D., Cascio, V. M. & Joung, J. K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nature Biotechnol. 32, 279–284 (2014)
Semenova, E. et al. Interference by clustered regularly interspaced short palindromic repeat (CRISPR) RNA is governed by a seed sequence. Proc. Natl Acad. Sci. USA 108, 10098–10103 (2011)
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nature Biotechnol. 31, 822–826 (2013)
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nature Biotechnol. 31, 833–838 (2013)
Pattanayak, V. et al. High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nature Biotechnol. 31, 839–843 (2013)
Lin, Y. et al. CRISPR/Cas9 systems have off-target activity with insertions or deletions between target DNA and guide RNA sequences. Nucleic Acids Res. 42, 7473–7485 (2014)
Bae, S., Park, J. & Kim, J.-S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014)
Wu, X. et al. Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells. Nature Biotechnol. 32, 670–676 (2014)
Kuscu, C., Arslan, S., Singh, R., Thorpe, J. & Adli, M. Genome-wide analysis reveals characteristics of off-target sites bound by the Cas9 endonuclease. Nature Biotechnol. 32, 677–683 (2014)
Crosetto, N. et al. Nucleotide-resolution DNA double-strand break mapping by next-generation sequencing. Nature Methods 10, 361–365 (2013)
Young, S. G. Recent progress in understanding apolipoprotein B. Circulation 82, 1574–1594 (1990)
Soutschek, J. et al. Therapeutic silencing of an endogenous gene by systemic administration of modified siRNAs. Nature 432, 173–178 (2004)
Rozema, D. B. et al. Dynamic PolyConjugates for targeted in vivo delivery of siRNA to hepatocytes. Proc. Natl Acad. Sci. USA 104, 12982–12987 (2007)
Wolfrum, C. et al. Mechanisms and optimization of in vivo delivery of lipophilic siRNAs. Nature Biotechnol. 25, 1149–1157 (2007)
Fitzgerald, K. et al. Effect of an RNA interference drug on the synthesis of proprotein convertase subtilisin/kexin type 9 (PCSK9) and the concentration of serum LDL cholesterol in healthy volunteers: a randomised, single-blind, placebo-controlled, phase 1 trial. Lancet 383, 60–68 (2014)
Abifadel, M. et al. Mutations in PCSK9 cause autosomal dominant hypercholesterolemia. Nature Genet. 34, 154–156 (2003)
Cohen, J. et al. Low LDL cholesterol in individuals of African descent resulting from frequent nonsense mutations in PCSK9. Nature Genet. 37, 161–165 (2005)
Horton, J. D., Cohen, J. C. & Hobbs, H. H. Molecular biology of PCSK9: its role in LDL metabolism. Trends Biochem. Sci. 32, 71–77 (2007)
Briner, A. E. et al. Guide RNA functional modules direct Cas9 activity and orthogonality. Mol. Cell 56, 333–339 (2014)
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nature Biotechnol. 33, 187–197 (2015)
Frock, R. L. et al. Genome-wide detection of DNA double-stranded breaks induced by engineered nucleases. Nature Biotechnol. 33, 179–186 (2015)
Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004)
Gautheret, D. & Lambert, A. Direct RNA motif definition and identification from multiple sequence alignments using secondary structure profiles. J. Mol. Biol. 313, 1003–1011 (2001)
Macke, T. J. et al. RNAMotif, an RNA secondary structure definition and search algorithm. Nucleic Acids Res. 29, 4724–4735 (2001)
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008)
Veldwijk, M. R. et al. Development and optimization of a real-time quantitative PCR-based method for the titration of AAV-2 vector stocks. Mol. Ther. 6, 272–278 (2002)
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003)
Acknowledgements
We thank E. Charpentier, I. Fonfara and K. Chylinski for discussions; A. Scherer-Hoock, B. Clear and the MIT Division of Comparative Medicine for assistance with animal experiments; Boston Children’s Hospital Viral Core and R. Xiao for assistance with AAV production; N. Crosetto for advice on BLESS; C.-Y. Lin and I. Slaymaker for experimental assistance; and the entire Zhang laboratory for support and advice. F.A.R. is a Junior Fellow at the Harvard Society of Fellows. W.X.Y. is supported by T32GM007753 from the National Institute of General Medical Sciences and a Paul and Daisy Soros Fellowship. J.S.G. is supported by a US Department of Energy Computational Science Graduate Fellowship. X.W. is a Howard Hughes Medical Institute International Student Research Fellow. P.A.S. is supported by United States Public Health Service grants RO1-GM34277, R01-CA133404 from the National Institutes of Health, and PO1-CA42063 from the National Cancer Institute, and partially by Cancer Center Support (core) grant P30-CA14051 from the National Cancer Institute. F.Z. is supported by the National Institutes of Health through NIMH (5DP1-MH100706) and NIDDK (5R01DK097768-03), a Waterman Award from the National Science Foundation, the Keck, New York Stem Cell, Damon Runyon, Searle Scholars, Merkin, and Vallee Foundations, and B. Metcalfe. F.Z. is a New York Stem Cell Foundation Robertson Investigator. The Children’s Hospital virus core is supported by an NIH core grant (5P30EY012196-17). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of General Medical Sciences or the National Institutes of Health. CRISPR reagents are available to the academic community through Addgene, and information about the protocols, plasmids, and reagents can be found at the Zhang laboratory website http://www.genome-engineering.org.
Author information
Authors and Affiliations
Contributions
F.A.R. and F.Z. conceived this study. F.A.R., L.C., W.X.Y. and F.Z. designed and performed the experiments with help from all authors. F.A.R., J.S.G., O.S., K.S.M., E.V.K. and F.Z. contributed to analysis of Cas9 orthologues, crRNA and tracrRNA, and PAM. A.J.K., F.A.R., X.W., and P.A.S. led ChIP and computational analysis and validation. F.A.R., W.X.Y. and L.C. performed BLESS and targeted sequencing of BLESS-identified off-target sites, and D.A.S. contributed computational analysis of BLESS data. W.X.Y., F.A.R., L.C. and B.Z. contributed animal data. W.X.Y., F.A.R., L.C., J.S.G., and F.Z. wrote the manuscript with help from all authors.
Corresponding author
Ethics declarations
Competing interests
Patent applications have been filed as to the subject matter of the manuscript such as, for example, US patent 8,865,406 issued 21 October 2014 and US patent 8,895,308 issued 25 November 2014. F.Z. is a founder of Editas Medicine and scientific advisor for Editas Medicine and Horizon Discovery.
Extended data figures and tables
Extended Data Figure 1 Selection of Type II CRISPR-Cas loci from eight bacterial species.
a, Distribution of lengths for Cas9 > 600 Cas9 orthologues19. b, Schematic of Type II CRISPR-Cas loci and sgRNA from eight bacterial species. Spacer or ‘guide’ sequences are shown in blue, followed by direct repeats (grey). Predicted tracrRNAs are shown in red, and folded based on the Constraint Generation RNA folding model50.
Extended Data Figure 2 Cas9 orthologue cleavage pattern in vitro.
Stacked bar graph indicates the fraction of targets cleaved at 2, 3, 4, or 5 bp upstream of PAM for each Cas9 orthologue; most Cas9 enzymes cleave stereotypically at 3 bp upstream of PAM (red triangle).
Extended Data Figure 3 Test of Cas9 orthologue activity in 293FT cells.
a, SURVEYOR assays showing indel formation at human endogenous loci from co-transfection of Cas9 orthologues and sgRNA. PAM sequences for individual targets are shown above each lane, with the consensus region for each PAM highlighted in red. Red triangles indicate cleaved fragments. b, SaCas9 generates indels efficiently for a multiple targets. c, Box-whisker plot of indel formation as a function of SaCas9 guide length L, with unaltered guides (perfect match of L nucleotides, grey bars) or replacement of the 5′-most base of guide with guanine (G + L − 1 nucleotides, blue bars) (n = 8 guides).
Extended Data Figure 4 Optimization of SaCas9 sgRNA scaffold in mammalian cells.
a, Schematic of the Staphylococcus aureus subspecies aureus CRISPR locus. b, Schematic of SaCas9 sgRNA with 21-nucleotide guide, crRNA repeat (grey), tetraloop (black) and tracrRNA (red). The number of crRNA repeat to tracrRNA anti-repeat base-pairing is indicated above the grey boxes. SaCas9 cleaves targets with varying repeat:anti-repeat lengths in c, HEK 293FT and d, Hepa1-6 cell lines. (n = 3, error bars show s.e.m.)
Extended Data Figure 5 Genome-wide binding by Cas9-chromatin immunoprecipitation (dCas9-ChIP).
a, Unbiased identification of PAM motif for dSaCas9 and dSpCas9. Peaks were analysed for the best match by motif score to the guide region only within 50 nucleotides of the peak summit. The alignment was extended for 10 nucleotides at the 3′ end and visualized using Weblogo. Numbers in parentheses indicate the number of called peaks. b, Histograms show the distribution of the peak summit relative to motif for dSaCas9 and dSpCas9. Position 1 on x axis indicates the first base of PAM.
Extended Data Figure 6 Indel measurements at candidate off-target sites based on ChIP.
Indels at top off-target sites predicted by dCas9-ChIP for each Cas9 and sgRNA pair, based on ChIP peaks ranked by sequence similarity of the genomic loci to the guide motif (heat map in purple), or P value of ChIP enrichment over control (heat map in red). Lines connect the common targets (EMX1) and off-targets between the two Cas9 enzymes.
Extended Data Figure 7 Analysis pipeline of sequencing data from BLESS.
a, Overview of the data analysis pipeline starting from the raw sequencing reads. Representative sequencing read mappings and corresponding histograms of the pairwise distances between all the forward orientation (red) reads and reverse orientation (blue) reads, displayed for representative b, DSB hotspots and poorly defined DSB sites and c, Cas9-induced DSBs with detectable indels. Fraction of pairwise distances between reads overlapping by no more than 6 bp (dashed vertical line) are indicated over histogram plots.
Extended Data Figure 8 Indel measurements at off-target sites based on DSB scores.
List of top off-target sites ranked by DSB scores for each Cas9 and sgRNA pair. Indel levels are determined by targeted deep sequencing. Blue triangles indicate positions of peak BLESS signal, and where present, PAMs and targets with sequence homology to the guide are highlighted. Lines connect the common on-targets (EMX1) and off-targets between the two Cas9 enzymes. N.D., not determined.
Extended Data Figure 9 Indel measurements of top candidate off-target sites based on sequence similarity score.
Off-targets are predicted based on sequence similarity to on-target, accounting for number and position of Watson–Crick base-pairing mismatches as previously described22. NNGRR and NRG are used as potential PAMs for SaCas9 and SpCas9, respectively. Lines connect the common targets (EMX1) and off-targets between the two Cas9 enzymes. Correlation plots between indel percentages and b, prediction based on sequence similarity, c, ChIP peaks ranked by motif similarity, or d, DSB scores for top ranking off-target loci. Trendlines, r2, and P values are calculated using ordinary least squares.
Extended Data Figure 10 SaCas9 targeting Apob locus in the mouse liver.
a, Schematics illustrating the mouse Apob gene locus and the positions of the three guides tested. b, Experimental time course and c, SURVEYOR assay showing indel formation at target loci after intravenous injection of AAV2/8 carrying thyroxine-binding globulin (TBG) promoter-driven SaCas9 and U6-driven guide at 2 × 1011 total genome copies (n = 1 animal each). d, Oil-red staining of liver tissue from AAV- or saline-injected animals. Male C56BL/6 mice were injected at 8 weeks of age and analysed 4 weeks post injection.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary References, Supplementary Tables 1-9 and Supplementary Sequences. (PDF 769 kb)
Source data
Rights and permissions
About this article
Cite this article
Ran, F., Cong, L., Yan, W. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015). https://doi.org/10.1038/nature14299
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14299
This article is cited by
-
Deep learning models to predict the editing efficiencies and outcomes of diverse base editors
Nature Biotechnology (2024)
-
Domain-inlaid Nme2Cas9 adenine base editors with improved activity and targeting scope
Nature Communications (2024)
-
Expanding the genome-targeting scope of a compact Cas9 using phage-assisted evolution
Nature Chemical Biology (2024)
-
Improving prime editing with an endogenous small RNA-binding protein
Nature (2024)
-
Efficient prime editing in mouse brain, liver and heart with dual AAVs
Nature Biotechnology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.