Abstract
Natural events present multiple types of sensory cues, each detected by a specialized sensory modality. Combining information from several modalities is essential for the selection of appropriate actions. Key to understanding multimodal computations is determining the structural patterns of multimodal convergence and how these patterns contribute to behaviour. Modalities could converge early, late or at multiple levels in the sensory processing hierarchy. Here we show that combining mechanosensory and nociceptive cues synergistically enhances the selection of the fastest mode of escape locomotion in Drosophila larvae. In an electron microscopy volume that spans the entire insect nervous system, we reconstructed the multisensory circuit supporting the synergy, spanning multiple levels of the sensory processing hierarchy. The wiring diagram revealed a complex multilevel multimodal convergence architecture. Using behavioural and physiological studies, we identified functionally connected circuit nodes that trigger the fastest locomotor mode, and others that facilitate it, and we provide evidence that multiple levels of multimodal integration contribute to escape mode selection. We propose that the multilevel multimodal convergence architecture may be a general feature of multisensory circuits enabling complex input–output functions and selective tuning to ecologically relevant combinations of cues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Stein, E. B. & Meredith, M. A. The Merging of the Senses (MIT press, 1993)
Fetsch, C. R., DeAngelis, G. C. & Angelaki, D. E. Bridging the gap between theories of sensory cue integration and the physiology of multisensory neurons. Nature Rev. Neurosci.. 14, 429–442 (2013)
van Atteveldt, N., Murray, M. M., Thut, G. & Schroeder, C. E. Multisensory integration: flexible use of general operations. Neuron 81, 1240–1253 (2014)
Stein, B. E., Meredith, M. A., Huneycutt, W. S. & McDade, L. Behavioral indices of multisensory integration: orientation to visual cues is affected by auditory stimuli. J. Cogn. Neurosci.. 1, 12–24 (1989)
McMeniman, C. J., Corfas, R. A., Matthews, B. J., Ritchie, S. A. & Vosshall, L. B. Multimodal integration of carbon dioxide and other sensory cues drives mosquito attraction to humans. Cell 156, 1060–1071 (2014)
Stein, B. E. & Stanford, T. R. Multisensory integration: current issues from the perspective of the single neuron. Nature Rev. Neurosci.. 9, 255–266 (2008)
Cybenko, G. Approximation by superpositions of a sigmoidal function. Math. Control Signals Syst. 2, 303–314 (1989)
Duda, R. O. E. H. P. &. Stork, D. G. Pattern Classification (John Wiley and Sons, 2001)
Hastie, T., Friedman, J. & Tibshirani, R. The Elements of Statistical Learning Vol. 2 (Springer, 2009)
Rigotti, M. et al. The importance of mixed selectivity in complex cognitive tasks. Nature 497, 585–590 (2013)
Fetsch, C. R., Pouget, A., DeAngelis, G. C. & Angelaki, D. E. Neural correlates of reliability-based cue weighting during multisensory integration. Nature Neurosci. 15, 146–154 (2012)
Meredith, M. A. & Stein, B. E. Interactions among converging sensory inputs in the superior colliculus. Science 221, 389–391 (1983)
Meredith, M. A. & Stein, B. E. Descending efferents from the superior colliculus relay integrated multisensory information. Science 227, 657–659 (1985)
Olcese, U., Iurilli, G. & Medini, P. Cellular and synaptic architecture of multisensory integration in the mouse neocortex. Neuron 79, 579–593 (2013)
Homberg, U., Christensen, T. A. & Hildebrand, J. G. Structure and function of the deutocerebrum in insects. Annu. Rev. Entomol.. 34, 477–501 (1989)
Milde, J. J. & Strausfeld, N. J. Cluster organization and response characteristics of the giant fiber pathway of the blowfly Calliphora erythrocephala. J. Comp. Neurol.. 294, 59–75 (1990)
Mu, L., Bacon, J. P., Ito, K. & Strausfeld, N. J. Responses of Drosophila giant descending neurons to visual and mechanical stimuli. J. Exp. Biol.. 217, 2121–2129 (2014)
Basura, G. J., Koehler, S. D. & Shore, S. E. Multi-sensory integration in brainstem and auditory cortex. Brain Res. 1485, 95–107 (2012)
Jain, R. & Shore, S. External inferior colliculus integrates trigeminal and acoustic information: unit responses to trigeminal nucleus and acoustic stimulation in the guinea pig. Neurosci. Lett.. 395, 71–75 (2006)
Lakatos, P., Chen, C. M., O’Connell, M. N., Mills, A. & Schroeder, C. E. Neuronal oscillations and multisensory interaction in primary auditory cortex. Neuron 53, 279–292 (2007)
Shore, S. E. Multisensory integration in the dorsal cochlear nucleus: unit responses to acoustic and trigeminal ganglion stimulation. Eur. J. Neurosci.. 21, 3334–3348 (2005)
Cardona, A. et al. An integrated micro- and macroarchitectural analysis of the Drosophila brain by computer-assisted serial section electron microscopy. PLoS Biol. 8, e1000502 (2010)
Saalfeld, S., Fetter, R., Cardona, A. & Tomancak, P. Elastic volume reconstruction from series of ultra-thin microscopy sections. Nature Methods 9, 717–720 (2012)
Pfeiffer, B. D. et al. Tools for neuroanatomy and neurogenetics in Drosophila. Proc. Natl Acad. Sci.. USA 105, 9715–9720 (2008)
Pfeiffer, B. D. et al. Refinement of tools for targeted gene expression in Drosophila. Genetics 186, 735–755 (2010)
Li, H. H. et al. A GAL4 driver resource for developmental and behavioral studies on the larval CNS of Drosophila. Cell Rep. 8, 897–908 (2014)
Hwang, R. Y. et al. Nociceptive neurons protect Drosophila larvae from parasitoid wasps. Curr. Biol. 17, 2105–2116 (2007)
Ohyama, T. et al. High-throughput analysis of stimulus-evoked behaviors in Drosophila larva reveals multiple modality-specific escape strategies. PLoS ONE 8, e71706 (2013)
Tsubouchi, A., Caldwell, J. C. & Tracey, W. D. Dendritic filopodia, Ripped Pocket, NOMPC, and NMDARs contribute to the sense of touch in Drosophila larvae. Curr. Biol. 22, 2124–2134 (2012)
Yan, Z. et al. Drosophila NOMPC is a mechanotransduction channel subunit for gentle-touch sensation. Nature 493, 221–225 (2013)
Zhang, W., Yan, Z., Jan, L. Y. & Jan, Y. N. Sound response mediated by the TRP channels NOMPC, NANCHUNG, and INACTIVE in chordotonal organs of Drosophila larvae. Proc. Natl Acad. Sci.. USA 110, 13612–13617 (2013)
Robertson, J. L., Tsubouchi, A. & Tracey, W. D. Larval defense against attack from parasitoid wasps requires nociceptive neurons. PLoS ONE 8, e78704 (2013)
Tracey, W. D., Jr, Wilson, R. I., Laurent, G. & Benzer, S. painless, a Drosophila gene essential for nociception. Cell 113, 261–273 (2003)
Vogelstein, J. T. et al. Discovery of brainwide neural–behavioral maps via multiscale unsupervised structure learning. Science 344, 386–392 (2014)
Xiang, Y. et al. Light-avoidance-mediating photoreceptors tile the Drosophila larval body wall. Nature 468, 921–926 (2010)
Zhong, L. et al. Thermosensory and nonthermosensory isoforms of Drosophila melanogaster TRPA1 reveal heat-sensor domains of a thermoTRP Channel. Cell Rep. 1, 43–55 (2012)
Schrader, S. & Merritt, D. J. Central projections of Drosophila sensory neurons in the transition from embryo to larva. J. Comp. Neurol. 425, 34–44 (2000)
Zlatic, M., Li, F., Strigini, M., Grueber, W. & Bate, M. Positional cues in the Drosophila nerve cord: semaphorins pattern the dorso-ventral axis. PLoS Biol. 7, e1000135 (2009)
Grueber, W. B. et al. Projections of Drosophila multidendritic neurons in the central nervous system: links with peripheral dendrite morphology. Development 134, 55–64 (2007)
Klapoetke, N. C. et al. Independent optical excitation of distinct neural populations. Nature Methods 11, 338–346 (2014)
Ernst, M. O. & Bulthoff, H. H. Merging the senses into a robust percept. Trends Cogn. Sci.. 8, 162–169 (2004)
Kenshalo, D. R., Iwata, K., Sholas, M. & Thomas, D. A. Response properties and organization of nociceptive neurons in area 1 of monkey primary somatosensory cortex. J. Neurophysiol. 84, 719–729 (2000)
Mendell, L. M. Physiological properties of unmyelinated fiber projection to the spinal cord. Exp. Neurol. 16, 316–332 (1966)
Ohshiro, T., Angelaki, D. E. & DeAngelis, G. C. A normalization model of multisensory integration. Nature Neurosci. 14, 775–782 (2011)
Bargmann, C. I. & Marder, E. From the connectome to brain function. Nature Methods 10, 483–490 (2013)
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Phil. Trans. R. Soc. Lond. B 314, 1–340 (1986)
Bock, D. D. et al. Network anatomy and in vivo physiology of visual cortical neurons. Nature 471, 177–182 (2011)
Briggman, K. L., Helmstaedter, M. & Denk, W. Wiring specificity in the direction-selectivity circuit of the retina. Nature 471, 183–188 (2011)
Kim, J. S. et al. Space-time wiring specificity supports direction selectivity in the retina. Nature 509, 331–336 (2014)
Takemura, S. Y. et al. A visual motion detection circuit suggested by Drosophila connectomics. Nature 500, 175–181 (2013)
Jenett, A. et al. A GAL4-driver line resource for Drosophila neurobiology. Cell Rep. 2, 991–1001 (2012)
Hamada, F. N. et al. An internal thermal sensor controlling temperature preference in Drosophila. Nature 454, 217–220 (2008)
Kang, K. et al. Modulation of TRPA1 thermal sensitivity enables sensory discrimination in Drosophila. Nature 481, 76–80 (2012)
Pfeiffer, B. D., Truman, J. W. & Rubin, G. M. Using translational enhancers to increase transgene expression in Drosophila. Proc. Natl Acad. Sci. USA 109, 6626–6631 (2012)
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013)
Tian, L. et al. Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators. Nature Methods 6, 875–881 (2009)
Ainsley, J. A. et al. Enhanced locomotion caused by loss of the Drosophila DEG/ENaC protein Pick-pocket1. Curr. Biol. 13, 1557–1563 (2003)
Baines, R. A., Uhler, J. P., Thompson, A., Sweeney, S. T. & Bate, M. Altered electrical properties in Drosophila neurons developing without synaptic transmission. J. Neurosci. 21, 1523–1531 (2001)
Lima, S. Q. & Miesenbck, G. Remote control of behavior through genetically targeted photostimulation of neurons. Cell 121, 141–152 (2005)
von Philipsborn, A. C. et al. Neuronal control of Drosophila courtship song. Neuron 69, 509–522 (2011)
Calleja, M., Moreno, E., Pelaz, S. & Morata, G. Visualization of gene expression in living adult Drosophila. Science 274, 252–255 (1996)
Luan, H., Peabody, N. C., Vinson, C. R. & White, B. H. Refined spatial manipulation of neuronal function by combinatorial restriction of transgene expression. Neuron 52, 425–436 (2006)
Struhl, G. & Basler, K. Organizing activity of wingless protein in Drosophila. Cell 72, 527–540 (1993)
Swierczek, N. A., Giles, A. C., Rankin, C. H. & Kerr, R. A. High-throughput behavioral analysis in C. elegans. Nature Methods 8, 592–598 (2011)
Kabra, M., Robie, A. A., Rivera-Alba, M., Branson, S. & Branson, K. JAABA: interactive machine learning for automatic annotation of animal behavior. Nature Methods 10, 64–67 (2013)
Schneider, C. A. et al. NIH Image to ImageJ: 25 years of image analysis. Nature Methods 9, 671–675 (2012)
Verstreken, P., Ohyama, T. & Bellen, H. J. in Exocytosis and Endocytosis 349–369 (Springer, 2008)
Glasbey, C. A. An analysis of histogram-based thresholding algorithms. CVGIP. Graph. Models Image Proc. 55, 532–537 (1993)
Sato, T. A modified method for lead staining of thin sections. J. Electron Microsc. (Tokyo) 17, 158–159 (1968)
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005)
Saalfeld, S., Cardona, A., Hartenstein, V. & Tomancak, P. CATMAID: collaborative annotation toolkit for massive amounts of image data. Bioinformatics 25, 1984–1986 (2009)
Prokop, A. & Meinertzhagen, I. A. Development and structure of synaptic contacts in Drosophila. Semin. Cell Dev. Biol. 17, 20–30 (2006)
Landgraf, M., Snchez Soriano, N., Technau, G. M., Urban, J. & Prokop, A. Charting the Drosophila neuropile: a strategy for the standardised characterization of genetically amenable neurites. Dev. Biol. 260, 207–225 (2003)
Grueber, W. B., Jan, L. & Jan, Y. Tiling of the Drosophila epidermis by multidendritic sensory neurons. Development 129, 2867–2878 (2002)
Bastian, M. et al. Gephi: an open source software for exploring and manipulating networks (International AAAI Conference on Weblogs and Social Media, 2009)
Bate, M., Goodman, C. & Spitzer, N. Embryonic development of identified neurons: Segment-specific differences in the H cell homologues. J. Neurosci. 1, 103–106 (1981)
Jefferis, G. S. et al. Comprehensive maps of Drosophila higher olfactory centers: spatially segregated fruit and pheromone representation. Cell 128, 1187–1203 (2007)
Dayan, P. & Abbott, L. F. Theoretical Neuroscience (MIT Press, 2001)
Acknowledgements
We thank G. Rubin and the Janelia Fly EM Project for the gift of the comprehensive image dataset of the larval nervous system; S. Lauchie and A. Brothers for assistance with EM imaging; B. Arruda and T. Dang for assistance with behavioural screens; L. Herren, I. Andrade, K. Floria and A. Berthold van der Bourg for assistance with neuronal reconstruction; G. Rubin, H. Dionne and B. Pfeiffer for GAL4 and Split GAL4 lines; A. Nern and G. Rubin for the single cell FLP-out stocks; J.-M. Knapp for Tsh-LexA stock; H.-H. Li and Janelia Fly Light Project Team for images of neuronal lines; K. Hibbard, M. Mercer, T. Laverty and the rest of Janelia Fly Core for stock construction and fly crosses; G. Denisov for the roll and crawl detection LARA software; E. Trautman, R. Svirskas, C. Weaver and D. Olbris for data analysis pipelines; Y. Park, C. Priebe, D. Naiman and J.-B. Masson for advice on statistical analysis; V. Jayaraman for input on calcium imaging; and W. Denk, B. Dickson, S. Druckmann, B. Gerber, K. Svoboda and C. Zuker for helpful comments on the manuscript. We thank Janelia HHMI for funding. A.C. and C.M.S.-M. were also funded by the Institute of Neuroinformatics of the University of Zurich and ETH Zurich, the SNSF grant 31003A_132969, the Universität Zürich Forschungskredit, and the HHMI Visiting Scientist program at Janelia. The EM image data is available via the Open Connectome Project (http://www.openconnectomeproject.org).
Author information
Authors and Affiliations
Contributions
T.O., C.M.S.-M., A.C. and M.Z. conceived the project, analysed the data and wrote the manuscript. T.O. performed and analysed behavioural and functional imaging experiments. C.M.S.-M., J.V.A. and A.C. performed neuronal reconstructions. A.C. registered the L1 volume. J.W.T. analysed the expression patterns of all the GAL4 lines and intersections for targeting of single cell types. R.D.F. generated the EM image data. M.R.A. and K.B. wrote the JAABA roll detection pipeline and performed statistical analysis. J.H.S. supported the generation of the abd1.5 dataset. R.F. provided critical suggestions for functional imaging experiments. C.M.S.-M. built the model. B.D.M. provided critical input and helped with writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 This figure relates to Fig. 1a–d.
Chordotonal (Ch) neurons respond to vibration in a ‘dose-dependent’ manner, but MD IV neurons do not, consistent with previous reports. a, 1,000 Hz vibration evokes calcium transients (mean ± s.e.m. in this and all subsequent figures) in chordotonal (ch-GAL4>UAS-GCaMP6s), in a dose-dependent manner. b, c, Calcium transients of MD IV neurons in response to different vibration (1,000 Hz) intensities reveal no response to vibration (MD IV-GAL4>UAS-GCaMP6s). d, Quantification of calcium transients in mechanosensory (black) and nociceptive (orange) neurons from (a–c). n = 3 trials in each condition. e, A schematic of larval reactions to unimodal and multimodal stimuli. Line thickness qualitatively reflects the probability of the behaviour transition occurring in the absence of stimulation (bottom left) and after the onset of MD IV activation (orange triangle, bottom right), vibration (green triangle, top left), or their combination (black triangle, top right). Behaviours in grey occur with extremely low frequency. In response to nociceptive or combined stimulation animals can select one of two types of escapes responses: (1) roll-fast crawl; or (2) fast crawl alone. Even in response to combined stimulation the selection is probabilistic; the majority of animals select roll-fast crawl, but some select fast crawl without rolling. f, Crawling stride speed in the absence of stimulation (white) and following 1,000 Hz, 6.7 m s−2 vibration (green) (in MD IV-GAL4>+ larvae); thermogenetic nociceptive activation (32 °C, in MD IV-GAL4>UAS-dTRPA1 larvae) (orange); or their combination (grey). In the nociceptive and combined condition, we separately analysed the stride speed of animals that did not roll (non-rollers) and those that rolled (rollers) following stimulation. As previously described, vibration alone evoked an increase in stride speed. Nociceptive activation alone, or in combination with vibration, also evoked an increase in stride speed, both in animals that did not roll and in animals that did roll, after rolling. Combining vibration with nociceptive stimulation in animals that selected fast crawl significantly increased stride speed compared to either stimulation alone. Vibration combined with nociceptive stimulation therefore affects both aspects of the escapes response, facilitating the triggering of rolling (Fig. 1e, f) and increasing crawling stride speed in animals that do not roll. Wilcoxon rank sum test with Holm–Bonferroni correction. **P < 0.0001. g, The synergistic effect on rolling of combining vibration with nociceptive stimulation is observed across a range of frequencies. Rolling probabilities of MD IV-GAL4>UAS-dTrpA1 (nociceptive and combined conditions) or MD IV-GAL4>+ (no stimuli and mechanosensory conditions) animals during vibration stimulation at different frequencies, either alone (green) or in combination with thermogenetic activation of nociceptive neurons (32 °C) (grey). See Supplementary Table 3 for n (animals) in this and all other Extended Data Figures. See Supplementary Table 2 for a list of genotypes used all Figures.
Extended Data Figure 2 This figure relates to Fig. 2a, b.
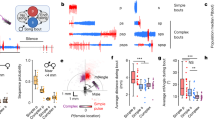
a, b, Confocal images of basins (green) and nociceptors (a) or chordotonals (b) (both in magenta), with cartoons to demonstrate their position within the nerve cord. c, Confocal images of individual Basins (in addition to those shown in Fig. 3a) visualized with a FLP-based method. Outline, neuropil boundary. Scale bar, 30 µm. d, Table of morphological features based on which Basins are uniquely distinguishable from other interneurons (blue) and individual Basin types from each other (purple). e, The table of morphological features based on which the mechanosensory chordotonal (Ch; green) and the nociceptive (orange) neuron projections are uniquely identifiable. f, Electron microscopy reconstructions of MD IV (orange) and chordotonal (green) axons in two abdominal segments, from two different animals. Top, reconstructions from the A3 segment in the smaller electron microscopy volume (comprising 1.5 abdominal segments: A3 and a part of A2) shown in Fig. 2. Bottom, reconstructions from the segment A1 in the comprehensive electron microscopy volume that spans the entire nervous system (L1 electron microscopy volume, see Fig. 5 and Supplementary Video 7). Outlines indicate approximate neuropil boundaries. Scale bar, 20 µm. g, Electron microscopy reconstructions of nociceptive (orange) and mechanosensory (green) axons and of the four Basin types (black) in the two abdominal segments, from two different animals. Top, A3 segment from the smaller 1.5 abdominal segment volume. Bottom, A1 segment in the comprehensive L1 electron microscopy volume. Outlines indicate approximate neuropil boundaries. Scale bar, 20 µm. The MD IV and chordotonal terminals span the ventromedial and ventrocentral nerve cord domains, respectively. Basin-2 and Basin-4 dendrites overlap with chordotonal and MD IV terminals. h, Detailed views of the neuronal arbors and synaptic locations of each Basin cell type from the two segments (from the two animals). Cells on the right side are mirrored for easier comparison. The location of synaptic domains is conserved across hemisegmental homologues and animals. The dendritic arbors of individual Basins across all four hemisegments are highly stereotyped and uniquely recognizable. i, Tables show the numbers of synapses that Basins in the A3 segment (from the smaller 1.5 segment electron microscopy dataset, top) and in the A1 segment (from the comprehensive L1 electron microscopy volume, bottom) receive from MD IV and chordotonal neurons in their own and in adjacent abdominal segments. See also Supplementary Tables 4a and 5a for more information. Both the left and right Basin-1 and Basin-3 homologues reproducibly received many synapses (each >25 synapses and >15% of total input, on average) from chordotonal in both animals, but very few (no more than 1% of total input synapses) from MD IV neurons. Both the left and right Basin-2 and Basin-4 homologues received many inputs from both chordotonal (on average >20 synapses and >10% total input) and MD IV (on average >20 synapses and >10% total input) neurons. j, Schematic representation of the distribution of inputs from MD IV and chordotonal from adjacent segments onto Basins.
Extended Data Figure 3 Reproducibility of the wiring diagram across animals.
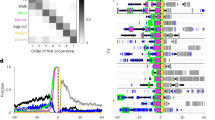
a, Electron microscopy reconstructions of the local interneurons downstream of both the MD IV and Basin neurons in both electron microscopy volumes; the volume comprising 1.5 abdominal (A2/A3) nerve cord segments of one animal (abd1.5, Fig. 2e), and the electron microscopy volume comprising the entire nervous system of another (L1, Fig. 5a and Supplementary Video 7). These local neurons (as well as the Basins shown in Fig. 2f and Extended Data Fig. 2h) with cell bodies and the majority of their arbor in the same segment as the sensory neurons are readily identifiable in both volumes and can be used to assess the reproducibility of sensory–interneuron and interneuron–interneuron connectivity across animals. Three distinct short-range interneuron types (A02m, A02n and A10a) received at least two synapses from nociceptive sensory neurons in both animals. A02m/n and A10a neurons also received inputs from Basin-2 or Basin-4 (Supplementary Information (Supplementary Atlas section)). The two interneurons A02m and A02n belong to the same lineage (lineage 2), have indistinguishable projections and connectivity, and can only be distinguished based on a small difference in the dorsoventral cell body location. In contrast, Basin-1, Basin-2, Basin-3 and Basin-4, even though they belong to the same lineage (lineage 9), all have distinct projections (and connectivity) and can each be considered a unique neuron type. A10a also have unique projections and belong to lineage 10. b, Total number of inputs each neuron in a receives. c, Reproducibility of connections between neuron types across individuals. Table shows class level connectivity between neurons in the abd1.5 and L1 data sets. Values given are the total sum of all synapses for all members of each class. 100% of connections with at least two synapses between homologous neuron types that were reproducible across the same individual were also reproducible across individuals; that is, the connection between those neurons existed in the other individual. d, Reproducibility of connections between individual neurons across individuals. Table shows connectivity between individual neurons in the abd1.5 and L1 data sets. Whenever two or more synapses were found between homologous neurons in both hemisegments in one animal, either two or more were also found in both hemisegments in the other animal (dark green cells); or at least one synapse per edge was found in both hemisegments in the other animal (light green cells); or two or more synapses were found in at least one hemisegment of the other animal (yellow cells). Thus, in 100% of cases in which a connection was present between homologues on both sides in one individual it was also present between homologues in the other individual, at least on one side.
Extended Data Figure 4 This figure relates to Figs 2 and 3.
a, c, Electron micrographs of representative synapses formed by MD IV and chordotonal (Ch) projections onto Basins containing a T-bar ribbon. Circles point to the postsynaptic neuron and arrows point to vesicle clouds in the presynaptic terminals. All synapses have small clear-core vesicles associated with fast chemical neurotransmitters such as acetylcholine, glutamate, or GABA. Scale bars are 200 nm. b, d, Calcium transients evoked in Basins by MD IV (b) and chordotonal (d) activation by local ATP injection. The ATP-gated channel, P2X2, is expressed in MD IV (b) and chordotonal (d) neurons and GCaMP6 is expressed in Basins (in MD IV-lexA>lexAop-P2X2; R72F11-GAL4>UAS-GCaMP6 (b) and Ch-lexA>lexAop-P2X2; R72F11-GAL4>UAS-GCaMP6 (d) animals). n = 30 trials (b), and 18 trials (d). Functional connectivity experiments suggest the connection from sensory neurons onto Basins is excitatory. Since chordotonal neurons are the only neurons that respond to vibration32,35 we used vibration in subsequent experiments so that we could activate chordotonal with vibration, and MD IV with P2X2 and ATP injection, in the same animal. e, R57F07 drives expression in Basin-4 neurons and in a few brain neurons. R57F07 was therefore used for generating the Basin-4 selective split-GAL4 line shown in g. g, h, Expression patterns of driver lines selective for Basin-1 (R20B01) and Basin-4 (R72F11 and R57F07 intersection) neurons (green). Scale bar, 25 μm. f, Activation of R57F07 is sufficient to evoke rolling in 45% of animals. Intersecting the parent line R57F07 with R72F11 (both drive expression in Basin-4 neurons and evoke rolling) results in a split-GAL4 line that drives expression selectively in Basin-4 neurons, shown in g. i, j, Negative control for Fig. 4d, f. i, Calcium transients in MD IV terminals observed in response to vibration (1,000 Hz, 6.8 m s−2), MD IV activation by local ATP injection or their combination (in MD IV-lexA>lexAop-P2X2; MD IV-GAL4>UAS-GCaMP6 animals). n = 14 trials. j, Maximum ΔF/F of calcium transients in response to mechanosensory, nociceptive, and combined simulation from i. Unlike Basin-4 (Fig. 3b, c), the MD IV neurons do not respond more to the combination of stimuli compared to MD IV activation alone.
Extended Data Figure 5 This figure relates to Fig. 4.
a–c, The R16E11-LexA line faintly, but selectively, drives expression in the same pair of thoracic neurons (Goro neurons) as the R69F06 GAL4 line. a, Co-localization of R16E11-LexA and R69F06-GAL4 expression in the same nervous system (genotype: R16E11-lexA/LexAop-mry-tdTomato;R69F06-GAL4,UAS-mry-GFP). R16E11-lexA>lexAop-tdTomato (red) and R69F06-GAL4>UAS-GFP (green) co-localize in the two Goro cell bodies and their projections in the nerve cord (yellow). b, c, Single-channel images from a. Scale bar, 20 μm. d, Thermogenetic activation (32 °C) of the single pair of Goro neurons using the R16E11 line (R16E11-lexA>lexop-dTRPA1) evoked rolling in 36% of animals. Activating neurons in R69F06 evokes even more rolling (86%, Fig. 4a). R69F06 drives expression in Goro neurons more strongly that R16E11, but it also drives expression in more neurons. Using a FLP-based intersection between R16E11-lexA and R69F06 (see Methods), we found that 76% of animals with expression of dTRPA1 selectively in Goro rolled when exposed to restrictive temperature (32 °C). Thus, activating the single pair of command-like Goro neurons evokes rolling. e, Calcium transients in Goro dendrites in response to selective Basin-4 activation by local ATP injection (n = 12 trials). The ATP-gated P2X2 channel and GCaMP6 are expressed selectively in Basin-4 and Goro neurons, respectively (in Basin-4-Gal4>UAS-P2X2; Goro-lexA(R69F06-lexA)>lexAop-GCaMP6 animals). f, Basin (magenta) and Goro (green) projections do not co-localize (in 16E11-lexA>LexAop-tdTomato; R72F11-GAL4>UAS-GFP larvae), indicating that the two cell types that are functionally connected cannot be directly synaptically connected.
Extended Data Figure 6 This figure relates to Fig. 5.
a, The nerve cord Basin–Goro pathway is sufficient to activate the Goro neurons. Calcium transients in Goro dendrites evoked by Basin activation by local ATP injection with the brain lobes intact (n = 15 trials) or removed (n = 24 trials) (in R72F11-LexA>LexAop-P2X2; Goro-GAL4(R69F06)>GCaMP6 animals). b, Quantification of a. The difference is not significant. Error bars show s.e.m. c, Left, R47D07 drives expression in A05q neurons (that receive direct inputs from Basin-2 and synapse onto Goro neurons) and in other neurons in thorax and brain (green, visualized with anti-GFP). In abdominal segments A8 and A7, the only neurons labelled with this line are the A05q neurons (dashed circle). The rest of the nervous system (grey) is visualized with anti-N-cadherin. White square, A05q axon terminals that synapse onto Goro dendrites. Scale bar, 25 μm. Note that this line also drives expression in developing adult neurons in thorax and brain that are not functional in the larval nervous system (secondary lineages). Right, image of individual A05q cells (magenta) visualized with a FLP-based strategy in R47D07, in top-down (top) and cross-section (bottom) views. Arrowheads indicate midline. Thin white lines indicate neuropil boundary. d, f, i, Electron micrographs of representative synapses formed by Basin-2 onto A05q (d), A05q onto Goro (f) and Basins onto A00c (i) containing T-bar ribbons. Circles point to the postsynaptic neuron and arrows point to vesicle clouds in the presynaptic terminals. All synapses have small clear-core vesicles associated with fast chemical neurotransmitters such as acetylcholine, glutamate, or GABA. Scale bars, 200 nm. e, Calcium transients in A05q axon terminals (square in c) evoked by Basin activation using local ATP injection (R72F11-LexA>lexop-P2X2; R47D07-GAL4>UAS-GCaMP6, n = 18 trials) suggest a functionally excitatory connection. g, Calcium transients in Goro neurons evoked by A05q activation using local ATP injection specifically in the vicinity of A05q dendrites in A8 (dotted circle in c; R47D07-GAL4>UAS-P2X2; Goro(R69E06-LexA>lexop-GCaMP6, n = 18 trials) suggest a functionally excitatory connection. h, Thermogenetic activation of neurons in R47D07 triggers rolling (exposing R47D07-GAL4>UAS-dTrpA1 animals to 32 °C). Control, +>UAS-dTrpA1. j, Calcium transients in A00c evoked by Basin activation using local ATP injection (R72F11-LexA>lexop-P2X2; R71A10-GAL4>UAS-GCaMP6, n = 15 trials) suggest a functionally excitatory connection.
Extended Data Figure 7 Distinct neurons differentially control distinct aspects of the escape sequence.
a, Graphs show the time course of rolling (green) and crawling (red and blue) behaviours before, during and after a 15 sec (indicated by the grey box) optogenetic (617 nm red light) activation period. The percentage of larvae that are rolling (green line) or crawling (red line) at each time point is shown. The normalized speed of crawling strides (normalized by speed prior to optogenetic stimulation) is also shown for those animals that are crawling at the time (blue line). Means (dark line) and s.e.m. (shaded areas) are shown. Data are from +>UAS-CsChrimson, basin1-GAL4>UAS-CsChrimson, basin4-GAL4>UAS-CsChrimson, Goro-GAL4(R69F06)>UAS-CsChrimson, or A00c-GAL4(R71A10)> UAS-CsChrimson animals, from top to bottom. b, Percentage of larvae that rolled at least once during the entire 15 sec activation window. Error bars are 95% confidence intervals. **P < 0.0001. Optogenetic activation of Basin-4 and Goro neurons evokes rolling, consistent with the results of thermogentic activation experiments shown in Figs 2a, g and 4e. Note that the instantaneous percentage of rolling shown in a is always lower than the percentage shown in b because not all larvae roll at exactly the same time. c, Comparison of the normalized speed of crawling strides in response to optogenetic activation. Boxes show medians (red) with 25th and 75th quantiles of the data. Whisker is 1.5 interquartile range (IQR) for the length of the whiskers. Red plus is outlier. **P < 0.0001. We observe that in control larvae, red light alone evoked a mild reduction in crawling speed, whereas Basin-1 activation evoked a mild, but significant, increase in stride speed and did not evoke rolling. Basin-4 activation evoked the entire escape sequence (roll followed by fast crawl). Note first an increase in instantaneous percentage of rolling larvae (peak in green line in a), followed by an increase in the instantaneous percentage of crawling larvae, and an increase in stride speed relative to the period prior to stimulation (peak in blue line in a). Goro neuron activation only evoked rolling and not crawling (note the percentage of crawling larvae is almost 0 during the optogenetic activation period). Furthermore there was no increase in relative stride speed following stimulation offset. Activation of the three A00c neurons with R71A10 was not sufficient to evoke rolling, but it was sufficient to evoke a mild, but significant, increase in stride speed. Thus, distinct neurons downstream of Basins appear to differentially control distinct aspects of the ‘roll-fast crawl’ escape sequence.
Extended Data Figure 8 Multilevel convergence can enhance sensitivity to weak multimodal stimuli.
A benefit of the multilevel architecture could be to amplify the effect of mechanosensory cues on nociceptive cues, increasing the sensitivity to relatively weak bimodal cues. To explore this idea further, we used a simple model to ask whether a two‐level network with two levels of convergence (multilevel convergence; MC) can be more sensitive to relatively weak bimodal events than a network with only early convergence (EC). a, Schematic for a simple model of early and multilevel convergence networks (see Methods for details). We considered an early convergence and a multilevel convergence two‐layer feed‐forward circuit with two inputs, corresponding to MD IV (ENoci) and chordotonal (EMech), and one output corresponding to rolling occurrence. We modelled steady‐state firing rates, where the output of each model neuron is a sigmoidal logistic function (with a lower threshold and upper saturation) of a weighted sum of its inputs. One pathway remains only chordotonal, while the other mixes modalities only early, or at both levels, depending on weights (wM and wB). b–d, Solutions for wM and wB are found using a constrained optimization procedure that maximizes sensitivity to weak bimodal inputs (b, c) within two experimentally observed unimodal target outputs (d): (1) that mechanosensory stimulus alone never evokes rolling; and (2) that the strongest MD IV stimulus alone evokes only 30% output (Fig. 1e, f and see Methods for details). e, Example deviation h from target outputs (see Methods) for multiple values of wM and wB for one set of thresholds (θp = 40; θM = 50; θB = 75). f, Values of wM and wB that maximize sensitivity S (dots) while also satisfying h < 3 (grey area). g, Early and multilevel solutions (if they exist) for other thresholds. Solutions exist if one or both θM or θB is high (keeping θp = 40). Note that although the optimal sensitivity multilevel convergence solution could have wM = 0 and thus effectively be an early convergence solution, this does not occur; multilevel convergence solutions are always more sensitive when they exist. Multilevel convergence solutions with wB = 0 are not shown since they do not exhibit multilevel convergence. h, Sensitivity of the optimal multilevel convergence solution as a function of the same thresholds as in Extended Data Fig. 8g, normalized by the most sensitive multilevel convergence solution found. Multilevel convergence solutions are more sensitive to relatively weak multimodal stimuli than early convergence solutions. Across all thresholds tested, the most sensitive circuits occur for θM= 50 and θB = 75 (Extended Data Fig. 8k). For such parameters, no early convergence solutions satisfy the unimodal constraints, hence multilevel convergence solutions are overall the most sensitive. i, j, Example early convergence and multilevel convergence solutions for wB and wB that satisfy the condition in d for one set of neuronal firing thresholds (θp = 40; θM = 50; θB = 75). k, Subtracting the output of early convergence from the multilevel convergence circuit. The multilevel convergence circuit triggers more rolling than the early convergence circuit in response to relatively weak bimodal cues and strong unimodal MD IV cues, but less rolling than the early convergence circuit to strong unimodal chordotonal cues (top left corner). The multilevel convergence circuit thus offers a more complex response function with greater sensitivity and selectivity for multimodal and strong unimodal MD IV cues, but not chordotonal cues, and could enable enhanced selection of rolling to more threatening events and reduced selection of rolling to less threatening events.
Supplementary information
Supplementary Information
This file contains the Supplementary Atlas. (PDF 13456 kb)
Supplementary Information
This zipped file contains Supplementary Tables 1-6 and a Supplementary Table Guide. (ZIP 718 kb)
MD IV activation alone
Contours of larvae automatically tracked with the MWT software in the presence of continuous MD IV activation alone in MD IV>dTRPA1 larvae (32oC). The activation onset was at 0 seconds, the video shows the period from 5 seconds to 20 seconds. Automatic detection of rolling using JAABA is shown. In all videos of larval behavior: pink contours indicate JAABA detected larva is rolling at the time; green contours, LARA software detected larva is crawling at that time; blue contours, larvae are neither rolling, nor crawling – they may be pausing or turning. Note some false negatives – blue contours in larvae that are rolling or crawling. Some larvae roll in response to MD IV activation. The majority of larvae are crawling forward. (MP4 965 kb)
Vibration alone
Contours of larvae in the presence of continuous 1000 Hz, 6.7 m/s2 vibration alone in MD IV>+ larvae (32oC). The activation onset was at 0 sec, this video shows the period from 5 sec to 20 sec. Almost no rolling is observed in response to vibration. Instead most larvae are crawling forward or turning. Most larvae have blue contours indicating JAABA did not detect rolling events (correctly). Note an occasional pink contour in a non-rolling larva indicating a false positive. (MP4 1411 kb)
MD IV activation combined with vibration
Contours of larvae automatically tracked with the MWT software in the presence of continuous vibration (1000 Hz, 6.7 m/s2) and MD IV activation in MD IV>dTRPA1 larvae (32oC). The activation onset was at 0 sec, the video shows the period from 5 sec to 20 sec. A large number of larvae are rolling and the rolling events are correctly detected by JAABA (pink contours). (MP4 751 kb)
Basin activation
Contours of larvae automatically tracked with the MWT software in the presence of continuous thermogenetic activation of Basins in R72F11>dTRPA1 larvae (32oC). The activation onset was at 0 sec, the video shows the period from 5 sec to 20 sec. A large number of larvae are rolling and the rolling events are correctly detected by JAABA (pink contours). (MP4 1146 kb)
Effector control
Contours of larvae automatically tracked with the MWT software in control attp2>dTRPA1 larvae (32oC). The temperature was raised at 0 sec and this video shows the period from 5 sec to 20 sec. No larvae are rolling and no contours are pink. (MP4 1177 kb)
Goro activation
Contours of larvae automatically tracked with the MWT software in the presence of continuous thermogenetic activation of Goro in R69F06>dTRPA1 larvae (32oC). The activation onset was at 0 sec, this video shows the period from 5 sec to 20 sec. A large number of larvae are rolling and the rolling events are correctly detected by JAABA (pink contours). (MP4 1075 kb)
EM volume spanning the entire nervous system of Drosophila larva.
This video illustrates the complete central nervous system of the first instar larva of Drosophila melanogaster, imaged with transmission electron microscopy. The volume consists of 4840 serial sections of 50 nm in thickness, prepared by Rick D. Fetter and imaged at a resolution of 3.8 x 3.8 nanometers/pixel with the semi-automatic EM imaging software Leginon by Rick D. Fetter and two assistants, Shirley Launchie and Andrea Brothers. The resulting 144,953 image tiles of 4096x4096 pixels each were registered by Albert Cardona and Stephan Saalfeld using the elastic serial section registration methods available in the software TrakEM2. (MP4 21230 kb)
Rotation of the EM reconstruction of the Goro cell shown in Figure 5b
Black lines indicate arbor, cyan marks show input synapses, red marks show presynaptic sites and the ball indicates the cell body. Dorsal is up. (MP4 2065 kb)
Rights and permissions
About this article
Cite this article
Ohyama, T., Schneider-Mizell, C., Fetter, R. et al. A multilevel multimodal circuit enhances action selection in Drosophila. Nature 520, 633–639 (2015). https://doi.org/10.1038/nature14297
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14297
This article is cited by
-
Descending GABAergic pathway links brain sugar-sensing to peripheral nociceptive gating in Drosophila
Nature Communications (2023)
-
A neuromechanical model for Drosophila larval crawling based on physical measurements
BMC Biology (2022)
-
A single-cell transcriptomic atlas of complete insect nervous systems across multiple life stages
Neural Development (2022)
-
Automated synapse-level reconstruction of neural circuits in the larval zebrafish brain
Nature Methods (2022)
-
Mapping of the zebrafish brain takes shape
Nature Methods (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.