Abstract
Immune checkpoint inhibitors1 result in impressive clinical responses2,3,4,5, but optimal results will require combination with each other6 and other therapies. This raises fundamental questions about mechanisms of non-redundancy and resistance. Here we report major tumour regressions in a subset of patients with metastatic melanoma treated with an anti-CTLA4 antibody (anti-CTLA4) and radiation, and reproduced this effect in mouse models. Although combined treatment improved responses in irradiated and unirradiated tumours, resistance was common. Unbiased analyses of mice revealed that resistance was due to upregulation of PD-L1 on melanoma cells and associated with T-cell exhaustion. Accordingly, optimal response in melanoma and other cancer types requires radiation, anti-CTLA4 and anti-PD-L1/PD-1. Anti-CTLA4 predominantly inhibits T-regulatory cells (Treg cells), thereby increasing the CD8 T-cell to Treg (CD8/Treg) ratio. Radiation enhances the diversity of the T-cell receptor (TCR) repertoire of intratumoral T cells. Together, anti-CTLA4 promotes expansion of T cells, while radiation shapes the TCR repertoire of the expanded peripheral clones. Addition of PD-L1 blockade reverses T-cell exhaustion to mitigate depression in the CD8/Treg ratio and further encourages oligoclonal T-cell expansion. Similarly to results from mice, patients on our clinical trial with melanoma showing high PD-L1 did not respond to radiation plus anti-CTLA4, demonstrated persistent T-cell exhaustion, and rapidly progressed. Thus, PD-L1 on melanoma cells allows tumours to escape anti-CTLA4-based therapy, and the combination of radiation, anti-CTLA4 and anti-PD-L1 promotes response and immunity through distinct mechanisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
The transcriptomic data are available at the GEO repository (http://www.ncbi.nlm.nih.gov/geo/) under accession GSE65503. Code for computational analysis is available upon request.
References
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nature Rev. Cancer 12, 252–264 (2012)
Hodi, F. S. et al. Improved survival with ipilimumab in patients with metastatic melanoma. N. Engl. J. Med. 363, 711–723 (2010)
Hamid, O. et al. Safety and tumor responses with lambrolizumab (anti-PD-1) in melanoma. N. Engl. J. Med. 369, 134–144 (2013)
Topalian, S. L. et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454 (2012)
Brahmer, J. R. et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N. Engl. J. Med. 366, 2455–2465 (2012)
Wolchok, J. D. et al. Nivolumab plus ipilimumab in advanced melanoma. N. Engl. J. Med. 369, 122–133 (2013)
Postow, M. A. et al. Immunologic correlates of the abscopal effect in a patient with melanoma. N. Engl. J. Med. 366, 925–931 (2012)
Ishwaran, H., Kogalur, U. B., Gorodeski, E. Z., Minn, A. J. & Lauer, M. S. High-dimensional variable selection for survival data. J. Am. Stat. Assoc. 105, 205–217 (2010)
Ishwaran, H., Kogalur, U. B., Chen, X. & Minn, A. J. Random survival forests for high-dimensional data. Statistical Analy Data Mining 4, 115–132 (2011)
Wilson, E. B. et al. Blockade of chronic type I interferon signaling to control persistent LCMV infection. Science 340, 202–207 (2013)
Teijaro, J. R. et al. Persistent LCMV infection is controlled by blockade of type I interferon signaling. Science 340, 207–211 (2013)
Jönsson, G. et al. Gene expression profiling-based identification of molecular subtypes in stage IV melanomas with different clinical outcome. Clin. Cancer Res. 16, 3356–3367 (2010)
Wherry, E. J. T cell exhaustion. Nature Immunol. 12, 492–499 (2011)
Paley, M. A. et al. Progenitor and terminal subsets of CD8+ T cells cooperate to contain chronic viral infection. Science 338, 1220–1225 (2012)
Thomas, N. et al. Tracking global changes induced in the CD4 T-cell receptor repertoire by immunization with a complex antigen using short stretches of CDR3 protein sequence. Bioinformatics 30, 3181–3188 (2014)
Atchley, W. R., Zhao, J., Fernandes, A. D. & Druke, T. Solving the protein sequence metric problem. Proc. Natl Acad. Sci. USA 102, 6395–6400 (2005)
Curran, M. A., Montalvo, W., Yagita, H. & Allison, J. P. PD-1 and CTLA-4 combination blockade expands infiltrating T cells and reduces regulatory T and myeloid cells within B16 melanoma tumors. Proc. Natl Acad. Sci. USA 107, 4275–4280 (2010)
Cha, E. et al. Improved survival with T cell clonotype stability after anti-CTLA-4 treatment in cancer patients. Sci. Transl. Med. 6, 238ra70–238ra70 (2014)
Robert, L. et al. CTLA4 blockade broadens the peripheral T-cell receptor repertoire. Clin. Cancer Res. 20, 2424–2432 (2014)
Reits, E. A. Radiation modulates the peptide repertoire, enhances MHC class I expression, and induces successful antitumor immunotherapy. J. Exp. Med. 203, 1259–1271 (2006)
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009)
Bayne, L. J. et al. Tumor-derived granulocyte-macrophage colony-stimulating factor regulates myeloid inflammation and T cell immunity in pancreatic cancer. Cancer Cell 21, 822–835 (2012)
Lee, Y. et al. Therapeutic effects of ablative radiation on local tumor require CD8+ T cells: changing strategies for cancer treatment. Blood 114, 589–595 (2009)
Dewan, M. Z. et al. Fractionated but not single-dose radiotherapy induces an immune-mediated abscopal effect when combined with anti-CTLA-4 antibody. Clin. Cancer Res. 15, 5379–5388 (2009)
Green, M. R. et al. Integrative analysis reveals selective 9p24.1 amplification, increased PD-1 ligand expression, and further induction via JAK2 in nodular sclerosing Hodgkin lymphoma and primary mediastinal large B-cell lymphoma. Blood 116, 3268–3277 (2010)
Holets, L. M. Trophoblast CD274 (B7-H1) is differentially expressed across gestation: influence of oxygen concentration. Biol. Reprod. 74, 352–358 (2006)
Cheadle, C., Vawter, M. P., Freed, W. J. & Becker, K. G. Analysis of microarray data using Z score transformation. J. Mol. Diagn. 5, 73–81 (2003)
Tusher, V. G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl Acad. Sci. USA 98, 5116–5121 (2001)
Efron, B. & Tibshirani, R. On testing the significance of sets of genes. Ann. Appl. Stat. 1, 107–129 (2007)
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001)
Chen, X. & Ishwaran, H. Random forests for genomic data analysis. Genomics 99, 323–329 (2012)
Ishwaran, H. & Kogalur, U. B. Random forests for survival, regression and classification (RF-SRC) (2013)
Rempala, G. A. & Seweryn, M. Methods for diversity and overlap analysis in T-cell receptor populations. J. Math. Biol. 67, 1339–1368 (2013)
Acknowledgements
C.T. was supported by an NIH training grant and career development award (T32DK007066, KL2TR000139). K.E.P. was supported by a Robertson Foundation/Cancer Research Institute Irvington Fellowship, T.C.G. and R.R. by the Melanoma Research Alliance, and X.X. and M.D.F by a grant from the NIH (P50CA174523). B.X. and A.J.M. were supported by the Basser Research Center for BRCA. A.J.M. is a Department of Defense Era of Hope Scholar (W81XWH-09-1-0339) and was supported by funding from the NIH/NCI (R01CA172651). H.I. and A.J.M. were supported by a grant from the NIH (R01CA163739). R.H.V. was supported by grants from the NIH (R01CA158186, P30CA016520) and by the Abramson Cancer Center Translational Center of Excellence in Pancreatic Cancer. E.J.W. was supported by funding from the NIH (U19AI082630, R01AI105343, U01AI095608 and P01AI112521). The project was supported in part by the Institute for Translational Medicine and Therapeutics' Transdisciplinary Program in Translational Medicine and Therapeutics, and the National Center for Research Resources (UL1RR024134).
Author information
Authors and Affiliations
Contributions
C.T. and A.J.R. designed, performed, and/or analysed mouse and in vitro experiments. H.D. assisted with mouse experiments. J.L.B. performed genetic ablation experiments for PD-L1. B.X. assisted with breast cancer mouse model. K.E.P. and P.M.O. designed and/or performed mouse immune profiling studies. E.S., R.S.H. and K.D.M. designed and/or performed human immune profiling studies. R.R. designed the clinical trial and was principle investigator along with S.M.H. and A.M., who oversaw its completion. D.P. assisted in coordinating the trial. A.M. evaluated the final clinical trial data. R.K.A., T.C.G. and L.M.S. were investigators on the trial, D.A.P. evaluated all imaging response for trial patients and assisted in study design, and M.D.F. and X.X. evaluated pathological biomarkers. R.M. was responsible for the statistical design of the clinical trial, and R.M. and A.J.M. performed statistical analysis. H.I. and A.J.M. performed and interpreted statistical analysis of pre-clinical data. A.J.M. wrote the manuscript, and E.J.W., R.H.V., A.M., C.T. and A.J.R. edited the manuscript. A.J.M., E.J.W. and R.H.V. together designed, interpreted, and oversaw the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Patients and mice treated with radiation + anti-CTLA4 for melanoma.
a, Twenty-two stage IV melanoma patients (M stage indicated) were stratified by treatment site of a single index metastasis, which was the irradiated tumour. Two dosing levels (DL) for stereotactic body radiation (SBRT) were in each stratum. b, Waterfall plot of the RECIST % change from baseline of unirradiated tumours annotated to indicate metabolic responses by PET/CT (hatches above plot) and response of the irradiated index tumour as measured by CT and PET/CT (hatches below plot). RECIST criteria do not include irradiated tumours. Legend shows colour-codes for response after CT or PET/CT (parenthesis). PMD, progressive metabolic disease; SMD, stable metabolic disease; PMR, partial metabolic response; CMR, complete metabolic response. White hatches indicate no imaging obtained. See Extended Data Table 2. c, Survival (right) and total tumour growth (bottom) after radiation treatment (RT) with either concurrent (n = 9) or sequential (n = 10) anti-CTLA4 compared to anti-CTLA4 (C4) (n = 10) or RT alone (n = 10). The regimens and the standard regimen used for all other melanoma experiments are illustrated (left). The P values for tumour growth are compared to anti-CTLA4 using a linear mixed-effects model. Overall P value for survival is by log-rank. d, Survival after RT and/or anti-CTLA4 with or without T-cell depletion (n = 5–10) using anti-CD8 (CD8). Shown are overall log-rank P values. The P value for RT + anti-CTLA4 with and without anti-CD8 is P = 0.005. Control is an isotype-matched antibody. e, Three mice with complete responses were rechallenged with B16-F10 tumours. Shown is a representative mouse. Arrow indicates location of regressed tumour and vitiligo-like condition represented by non-pigmented fur (observed in approximately 50% of mice with complete responses). Time line starts from original tumour implantation (day 0) and values above marks are days after first rechallenge. Recurrence occurred only after anti-CD8 treatment and second rechallenge.
Extended Data Figure 2 Tumour cells resistant to radiation + anti-CTLA4 upregulate PD-L1 but not other candidate inhibitory receptor pathways.
a, Unirradiated tumour growth (left, normalized; right, raw values) for mice implanted with Res 177 (n = 21), Res 499 (n = 25), and B16-F10 (n = 18) melanoma cells and treated with radiation treatment (RT) + anti-CTLA4 (C4). For normalization, volumes were divided by average of untreated controls (V/Vcont) to account for differences in growth between untreated tumour types. The P values are for comparisons with B16-F10 tumours using a linear mixed-effects model. b, Corresponding tumour volumes of unirradiated or irradiated index tumours at day 21 (blue line is mean). c, Clonogenic survival for Res 499 and B16-F10 cells (n = 2). d, Selection of immune variables that robustly predict resistance to RT + anti-CTLA4 using minimal depth (MD). A variable was selected if its MD was less than a threshold value for significance. Shown are bootstrap distributions of MD values (left) and % bootstrap models for which the MD for the indicated variable was significant (right). Bootstrap mean ± s.d. for the out-of-bag prediction error rate is listed on top. e, Volcano plot of differentially expressed genes from resistant tumours. Horizontal black line is 5% false-discovery rate and dotted green line is fold-change cut-off. Ligands for select inhibitory receptors are indicated. See Supplementary Table 1. f, Unirradiated tumour volumes (day 26–29) and g, survival after RT + anti-CTLA4 for mice with bilateral tumours from TSA breast cancer cells (n = 25) or from the Res 237 subline selected to be resistant (n = 21). h, Expression of candidate T cell inhibitory receptor ligands on B16-F10 and Res 499. Interferon-gamma (IFN-γ) responsiveness was tested. i, Boxplots show distribution of % positive CD8+CD44+ T cells for the indicated inhibitory receptor compared to IgG control. j, PD-L1 surface expression for CRISPR PD-L1 homozygous knockout Res 499 and wild type control cells. IFN-γ was used to induce PD-L1 and confirm abrogated response. Two-tailed t-test or Wilcoxon test was used for two-way comparisons. Log-rank test was used for survival analysis.
Extended Data Figure 3 Addition of PD-L1/PD-1 blockade antagonizes resistance to radiation + anti-CTLA4, and optimal response to checkpoint blockade requires radiation.
a, Change in % CD8+CD44+ T cells after radiation treatment (RT) and checkpoint blockade vs change in the degree of reinvigoration of exhausted T cells measured by % PD-1+Eomes+ T cells that are Ki67+GzmB+. Values are subtracted from average of untreated control. b, Growth of Res 499 tumours after RT + anti-CTLA4 (C4) with and without addition of anti-PD-L1 (PDL1). Shown are index and unirradiated tumours from n = 25 mice in each group. The P value is for comparison to RT + anti-CTLA4. c, Proportion of complete responses (yellow) for mice with Res 499 tumours. d, Total tumour growth (index + unirradiated) for B16-F10 tumours after the indicated treatment that includes anti-PD-1 (PD1) or anti-PD-L1. The P values are for comparisons to RT + anti-CTLA4 (n = 18, n = 5 for others). Pie charts show % complete responses (yellow). e, Survival of mice after RT + anti-CTLA4 + anti-PD-1. Shown is the overall P value, and f, the two-way comparisons that include those from Fig. 2d. g, Proportion of mice with complete responses (yellow) after RT + anti-PD-L1 or anti-PD-1 that survived 90+ days after tumour rechallenge at day 60 (n = 12). h, Survival of mice with bilateral Res 237 breast cancer tumours treated with RT + anti-CTLA4 with (n = 16) or without (n = 21) anti-PD-L1. i, Proportion of complete responses (yellow) for mice with Res 237 or TSA breast cancer tumours. j, Survival of mice with pancreatic tumours from a cell line derived from KPC mice (KrasLSL-G12D/+;p53LSL-R172H/+;Pdx-1-Cre) (n = 5 for each group). Select treatment groups are labelled on the plot for clarity. Overall P value is shown. A linear mixed-effects model was used for tumour growth analysis. Log-rank test was used for survival analysis.
Extended Data Figure 4 TCR clonotypes associated with radiation are not observed in random clones from post-treatment blood and have distinct CDR3 features.
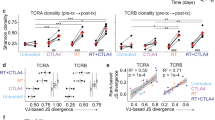
a, Boxplot of the bootstrap variance explained by multivariable random forest regression model for effect of RT, anti-CTLA4, and/or anti-PD-L1 on immune variables from TILs. b, k-means clustering (k = 2) was used on the average CDR3 amino acid features of randomly sampled clones from post-treatment blood after anti-CTLA4, anti-PD-L1, and/or radiation treatment (RT). Membership into each cluster was determined and the P value for separation into treatment groups with and without RT was calculated by Fisher’s exact test. Boxplot shows log10(P values) from 1,000 random iterations. Comparison to the P value from the observed data (red dotted line) gives a simulated P < 0.001. c, log10(P values) for separation into treatment groups with and without RT vs cut-off value used to select the most frequent clones. The 0.05 significance level is indicated (red dotted line). d, Average % occupancy in the CDR3 of the most frequent T cell clonotypes after RT ± checkpoint blockade (+RT, red line) or checkpoint blockade alone (NoRT, orange line) by contiguous short amino acid sequences of length three (3-tuples) belonging to e, subsets with distinct treatment-related amino acid properties. These properties are characterized by Atchley factors, which measure (1) PAH: accessibility, polarity, and hydrophobicity, (2) PSS: propensity for secondary structure, (3) MS: molecular size, (4) CC: codon composition, and (5) EC: electrostatic charge. Shown (right) are the average values of each Atchley factor for amino acids that comprise the 3-tuples from the indicated subset (red) compared to all unselected 3-tuples (blue). Boxplots (left) show the proportion of 3-tuples from each of these subsets that are found in the CDR3s of the five most frequent clones after treatment. Compared to pre-treatment samples (Pre-Tx), subset 6 is associated with RT ± checkpoint blockade (+RT) or checkpoint blockade alone (NoRT). Subset 1 is primarily associated with checkpoint blockade alone, and subset 16 is primarily associated with RT ± checkpoint blockade.
Extended Data Figure 5 Peripheral T-cell exhaustion, reinvigoration, CD8/Treg ratio, and tumour PD-L1 predict response to radiation + immune checkpoint blockade.
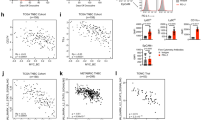
a, Heat map showing the relative proportions of PD-1+ CD8 T cells that are Ki67+GzmB+ or Eomes+ and the CD8/Treg ratio for each sample (columns) subtracted from the average values of untreated controls. Black hatches indicated complete response (CR) and radiation treatment (RT) + anti-CTLA4 (C4) ± anti-PD-L1 (P1). From these data, a multivariable random forest predictor for complete response was developed. Boxplot shows bootstrap distributions of variable importance scores (more predictive variables have higher values), and of b, minimal depth (MD), a statistic to measure predictiveness. Bar plot shows % bootstrap models for which the MD for the indicated variable was significant. Bootstrap mean ± s.d. for the out-of-bag prediction error rate is listed on top. c, Probability of complete response vs change (treated vs untreated control) in CD8/Treg ratio for mice with a high (blue dots) or low (red dots) change in % PD-1+ splenic CD8 T cells that are Eomes+. d, Heat map similar to a except using T cells from peripheral blood. e, Percent peripheral blood PD-1+ CD8 T cells that are Eomes+ vs Ki67+GzmB+ after RT + checkpoint blockade. Values are subtracted from average of untreated controls. Each circle represents a mouse. Probability of complete response (proportional to circle size), prediction error rate, and quadrant boundaries are estimated from the random forest model. f, Representative contour plots examining splenic CD8 T cells from B16-F10 or Res 499 tumours for PD-1 and Eomes (top), followed by examination of the PD-1+Eomes+ subset for Ki67 and GzmB (bottom). g, Ratios of PD-1+Eomes+ splenic CD8 T cells that are Ki67+GzmB+ (reinvigorated) compared to Ki67−GzmB− (exhausted) from mice with Res 499 tumours. P value by two-tailed t-test.
Extended Data Figure 6 Melanoma PD-L1 is associated with T-cell exhaustion, response, and survival for patients treated on clinical trial of radiation + anti-CTLA4.
a, Representative images (right) for patients with biopsies showing PD-L1 staining on tumour cells classified as PD-L1lo (top), 2+ (middle), or 3+ (bottom). Scores of 2+ and 3+ are classified as PD-L1hi. The arrow indicates PD-L1 staining on macrophages. An isotype antibody negative control and positive controls are shown (left). b, Changes in % Ki67+GzmB+ in PD-1+ CD8 T cells after radiation treatment (RT) + anti-CTLA4 vs PD-L1 status on melanoma cells from all patients with available pre- and post-treatment blood. c, Changes in % Ki67+GzmB+ in PD-1+Eomes+ CD8 T cells (left) or in PD-1+ CD8 T cells (right) vs macrophage PD-L1 status. d, Hazard ratio and 95% confidence interval for progression-free survival from a Cox regression model using PD-L1 status on tumour cells and macrophages. e, Model for non-redundant mechanisms and resistance to RT and immune checkpoint blockade. Two-tailed t-test was used for two-way comparisons.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, including a list of flow cytometry antibodies used for immune profiling in mouse and human experiments. (PDF 97 kb)
Supplementary Table 1
This file contains a list of genes differentially expressed in melanoma tumors resistant to RT + anti-CTLA4. (XLSX 54 kb)
Supplementary Table 2
This file contains normalized and adjusted transcriptomic data for melanoma tumors resistant or sensitive to RT + anti-CTLA4. (XLSX 4658 kb)
Rights and permissions
About this article
Cite this article
Twyman-Saint Victor, C., Rech, A., Maity, A. et al. Radiation and dual checkpoint blockade activate non-redundant immune mechanisms in cancer. Nature 520, 373–377 (2015). https://doi.org/10.1038/nature14292
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14292
This article is cited by
-
Roles of tumor-associated macrophages in anti-PD-1/PD-L1 immunotherapy for solid cancers
Molecular Cancer (2023)
-
Cancer cells resistant to immune checkpoint blockade acquire interferon-associated epigenetic memory to sustain T cell dysfunction
Nature Cancer (2023)
-
The optimization of postoperative radiotherapy in de novo stage IV breast cancer: evidence from real-world data to personalize treatment decisions
Scientific Reports (2023)
-
Clinical-mediated discovery of pyroptosis in CD8+ T cell and NK cell reveals melanoma heterogeneity by single-cell and bulk sequence
Cell Death & Disease (2023)
-
The mutual relationship between the host immune system and radiotherapy: stimulating the action of immune cells by irradiation
International Journal of Clinical Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.