Abstract
Single particle electron cryomicroscopy (cryo-EM) has recently made significant progress in high-resolution structure determination of macromolecular complexes due to improvements in electron microscopic instrumentation and computational image analysis. However, cryo-EM structures can be highly non-uniform in local resolution1,2 and all structures available to date have been limited to resolutions above 3 Å3,4. Here we present the cryo-EM structure of the 70S ribosome from Escherichia coli in complex with elongation factor Tu, aminoacyl-tRNA and the antibiotic kirromycin at 2.65–2.9 Å resolution using spherical aberration (Cs)-corrected cryo-EM. Overall, the cryo-EM reconstruction at 2.9 Å resolution is comparable to the best-resolved X-ray structure of the E. coli 70S ribosome5 (2.8 Å), but provides more detailed information (2.65 Å) at the functionally important ribosomal core. The cryo-EM map elucidates for the first time the structure of all 35 rRNA modifications in the bacterial ribosome, explaining their roles in fine-tuning ribosome structure and function and modulating the action of antibiotics. We also obtained atomic models for flexible parts of the ribosome such as ribosomal proteins L9 and L31. The refined cryo-EM-based model presents the currently most complete high-resolution structure of the E. coli ribosome, which demonstrates the power of cryo-EM in structure determination of large and dynamic macromolecular complexes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
The 2.9 Å cryo-EM map of the E. coli ribosome–EF-Tu complex has been deposited in the Electron Microscopy Data Bank with accession code EMD-2847, the coordinates of the atomic model have been deposited in the Protein Data Bank under accession code 5AFI.
References
Leschziner, A. E. & Nogales, E. Visualizing flexibility at molecular resolution: analysis of heterogeneity in single-particle electron microscopy reconstructions. Annu. Rev. Biophys. Biomol. Struct. 36, 43–62 (2007)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nature Methods 11, 63–65 (2014)
Yu, X., Ge, P., Jiang, J. S., Atanasov, I. & Zhou, Z. H. Atomic model of CPV reveals the mechanism used by this single-shelled virus to economically carry out functions conserved in multishelled reoviruses. Structure 19, 652–661 (2011)
Wong, W. et al. Cryo-EM structure of the Plasmodium falciparum 80S ribosome bound to the anti-protozoan drug emetine. eLife 3, e03080 (2014)
Noeske, J. et al. Synergy of streptogramin antibiotics occurs independently of their effects on translation. Antimicrob. Agents Chemother. 58, 5269–5279 (2014)
Schmeing, T. M., Voorhees, R. M., Kelley, A. C. & Ramakrishnan, V. How mutations in tRNA distant from the anticodon affect the fidelity of decoding. Nature Struct. Mol. Biol. 18, 432–436 (2011)
Fischer, N., Konevega, A. L., Wintermeyer, W., Rodnina, M. V. & Stark, H. Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy. Nature 466, 329–333 (2010)
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012)
Polikanov, Y. S. et al. Amicoumacin A inhibits translation by stabilizing mRNA interaction with the ribosome. Mol. Cell 56, 531–540 (2014)
Schmeing, T. M., Huang, K. S., Kitchen, D. E., Strobel, S. A. & Steitz, T. A. Structural insights into the roles of water and the 2′ hydroxyl of the P site tRNA in the peptidyl transferase reaction. Mol. Cell 20, 437–448 (2005)
Sergiev, P. et al. in Ribosomes (eds Rodnina, M. V., Wintermeyer, W. & Green, R. ) Ch. 9 97–110 (Springer Vienna, 2011)
Burakovsky, D. E. et al. Impact of methylations of m2G966/m5C967 in 16S rRNA on bacterial fitness and translation initiation. Nucleic Acids Res. 40, 7885–7895 (2012)
Das, G. et al. Role of 16S ribosomal RNA methylations in translation initiation in Escherichia coli. EMBO J. 27, 840–851 (2008)
Kimura, S. & Suzuki, T. Fine-tuning of the ribosomal decoding center by conserved methyl-modifications in the Escherichia coli 16S rRNA. Nucleic Acids Res. 38, 1341–1352 (2010)
Schuwirth, B. S. et al. Structural analysis of kasugamycin inhibition of translation. Nature Struct. Mol. Biol. 13, 879–886 (2006)
Gutierrez, B. et al. Fitness cost and interference of Arm/Rmt aminoglycoside resistance with the RsmF housekeeping methyltransferases. Antimicrob. Agents Chemother. 56, 2335–2341 (2012)
Green, R. & Noller, H. F. In vitro complementation analysis localizes 23S rRNA posttranscriptional modifications that are required for Escherichia coli 50S ribosomal subunit assembly and function. RNA 2, 1011–1021 (1996)
LaMarre, J. M., Howden, B. P. & Mankin, A. S. Inactivation of the indigenous methyltransferase RlmN in Staphylococcus aureus increases linezolid resistance. Antimicrob. Agents Chemother. 55, 2989–2991 (2011)
Toh, S.-M. & Mankin, A. S. An indigenous posttranscriptional modification in the ribosomal peptidyl transferase center confers resistance to an array of protein synthesis inhibitors. J. Mol. Biol. 380, 593–597 (2008)
Osterman, I. A. et al. Methylated 23S rRNA nucleotide m2G1835 of Escherichia coli ribosome facilitates subunit association. Biochimie 93, 725–729 (2011)
Sergiev, P. V., Serebryakova, M. V., Bogdanov, A. A. & Dontsova, O. A. The ybiN gene of Escherichia coli encodes adenine-N6 methyltransferase specific for modification of A1618 of 23S ribosomal RNA, a methylated residue located close to the ribosomal exit tunnel. J. Mol. Biol. 375, 291–300 (2008)
David-Eden, H., Mankin, A. S. & Mandel-Gutfreund, Y. Structural signatures of antibiotic binding sites on the ribosome. Nucleic Acids Res. 38, 5982–5994 (2010)
Burnley, B. T., Afonine, P. V., Adams, P. D. & Gros, P. Modelling dynamics in protein crystal structures by ensemble refinement. eLife 1, e00311 (2012)
Schröder, G. F., Levitt, M. & Brunger, A. T. Deformable elastic network refinement for low-resolution macromolecular crystallography. Acta Crystallogr. D 70, 2241–2255 (2014)
Kleywegt, G. J. Crystallographic refinement of ligand complexes. Acta Crystallogr. D 63, 94–100 (2007)
Schmeing, T. M. et al. The crystal structure of the ribosome bound to EF-Tu and aminoacyl-tRNA. Science 326, 688–694 (2009)
Brandt, F. et al. The native 3D organization of bacterial polysomes. Cell 136, 261–271 (2009)
Akanuma, G., Nanamiya, H., Natori, Y., Nomura, N. & Kawamura, F. Liberation of zinc-containing L31 (RpmE) from ribosomes by its paralogous gene product, YtiA, in Bacillus subtilis. J. Bacteriol. 188, 2715–2720 (2006)
Samaha, R. R., Green, R. & Noller, H. F. A base pair between tRNA and 23S rRNA in the peptidyl transferase centre of the ribosome. Nature 377, 309–314 (1995)
Ippolito, J. A. et al. Crystal structure of the oxazolidinone antibiotic linezolid bound to the 50S ribosomal subunit. J. Med. Chem. 51, 3353–3356 (2008)
Milon, P. et al. Transient kinetics, fluorescence, and FRET in studies of initiation of translation in bacteria. Methods Enzymol. 430, 1–30 (2007)
Rodnina, M. V. et al. Thiostrepton inhibits the turnover but not the GTPase of elongation factor G on the ribosome. Proc. Natl Acad. Sci. USA 96, 9586–9590 (1999)
Rodnina, M. V. & Wintermeyer, W. GTP consumption of elongation factor Tu during translation of heteropolymeric messenger-RNAs. Proc. Natl Acad. Sci. USA 92, 1945–1949 (1995)
Sander, B., Golas, M. M. & Stark, H. Automatic CTF correction for single particles based upon multivariate statistical analysis of individual power spectra. J. Struct. Biol. 142, 392–401 (2003)
Valle, M. et al. Cryo-EM reveals an active role for aminoacyl-tRNA in the accommodation process. EMBO J. 21, 3557–3567 (2002)
Stark, H. et al. Ribosome interactions of aminoacyl-tRNA and elongation factor Tu in the codon-recognition complex. Nature Struct. Mol. Biol. 9, 849–854 (2002)
Scheres, S. H. W. RELION: Implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Bock, L. V. et al. Energy barriers and driving forces in tRNA translocation through the ribosome. Nature Struct. Mol. Biol. 20, 1390–1396 (2013)
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013)
Kleywegt, G. J. & Jones, T. A. Software for handling macromolecular envelopes. Acta Crystallogr. D 55, 941–944 (1999)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Brunger, A. T. Version 1.2 of the Crystallography and NMR system. Nature Protocols 2, 2728–2733 (2007)
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Greber, B. J. et al. The complete structure of the large subunit of the mammalian mitochondrial ribosome. Nature 515, 283–286 (2014)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010)
DiMaio, F. et al. Improved molecular replacement by density-and energy-guided protein structure optimization. Nature 473, 540–543 (2011)
DiMaio, F., Tyka, M. D., Baker, M. L., Chiu, W. & Baker, D. Refinement of protein structures into low-resolution density maps using rosetta. J. Mol. Biol. 392, 181–190 (2009)
Söding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005)
Dunkle, J. A. et al. Structures of the bacterial ribosome in classical and hybrid states of tRNA binding. Science 332, 981–984 (2011)
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006)
Byrne, R. T., Konevega, A. L., Rodnina, M. V. & Antson, A. A. The crystal structure of unmodified tRNAPhe from Escherichia coli. Nucleic Acids Res. 38, 4154–4162 (2010)
Schröder, G. F., Levitt, M. & Brunger, A. T. Super-resolution biomolecular crystallography with low-resolution data. Nature 464, 1218–1222 (2010)
Emsley, P., Lohkamp, B., Scott, W. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Terwilliger, T. C. Maximum-likelihood density modification. Acta Crystallogr. D 56, 965–972 (2000)
The CCP4 Suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Chou, F.-C., Sripakdeevong, P., Dibrov, S. M., Hermann, T. & Das, R. Correcting pervasive errors in RNA crystallography through enumerative structure prediction. Nature Methods 10, 74–76 (2013)
Afonine, P. V., Headd, J. J., Terwilliger, T. C. & Adams, P. D. New tool: phenix.real_space_refine. Computational Crystallography Newsletter 4, 43–44 (2013)
Jenner, L., Demeshkina, N., Yusupova, G. & Yusupov, M. Structural rearrangements of the ribosome at the tRNA proofreading step. Nature Struct. Mol. Biol. 17, 1072–1078 (2010)
Jones, T. A. & Kjeldgaard, M. Electron-density map interpretation. Methods Enzymol. 277, 173–208 (1997)
Allegretti, M., Mills, D. J., McMullan, G., Kühlbrandt, W. & Vonck, J. Atomic model of the F420-reducing [NiFe] hydrogenase by electron cryo-microscopy using a direct electron detector. eLife 3, e01963 (2014)
Bartesaghi, A., Matthies, D., Banerjee, S., Merk, A. & Subramaniam, S. Structure of β-galactosidase at 3.2-Å resolution obtained by cryo-electron microscopy. Proc. Natl Acad. Sci. USA 111, 11709–11714 (2014)
Burmeister, W. P. Structural changes in a cryo-cooled protein crystal owing to radiation damage. Acta Crystallogr. D 56, 328–341 (2000)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Grosse-Kunstleve, R. W., Sauter, N. K., Moriarty, N. W. & Adams, P. D. The Computational Crystallography Toolbox: crystallographic algorithms in a reusable software framework. J. Appl. Crystallogr. 35, 126–136 (2002)
Demirci, H. et al. Modification of 16S ribosomal RNA by the KsgA methyltransferase restructures the 30S subunit to optimize ribosome function. RNA 16, 2319–2324 (2010)
Helser, T. L., Davies, J. E. & Dahlberg, J. E. Mechanism of kasugamycin resistance in Escherichia coli. Nature 235, 6–9 (1972)
O’Connor, M., Thomas, C. L., Zimmermann, R. A. & Dahlberg, A. E. Decoding fidelity at the ribosomal A and P sites: influence of mutations in three different regions of the decoding domain in 16S rRNA. Nucleic Acids Res. 25, 1185–1193 (1997)
François, B. et al. Crystal structures of complexes between aminoglycosides and decoding A site oligonucleotides: role of the number of rings and positive charges in the specific binding leading to miscoding. Nucleic Acids Res. 33, 5677–5690 (2005)
Jiang, J., Aduri, R., Chow, C. S. & SantaLucia, J. Structure modulation of helix 69 from Escherichia coli 23S ribosomal RNA by pseudouridylations. Nucleic Acids Res. 42, 3971–3981 (2013)
Davis, D. R. Stabilization of RNA stacking by pseudouridine. Nucleic Acids Res. 23, 5020–5026 (1995)
Ortiz-Meoz, R. F. & Green, R. Helix 69 is key for uniformity during substrate selection on the ribosome. J. Biol. Chem. 286, 25604–25610 (2011)
Seidelt, B. et al. Structural insight into nascent polypeptide chain–mediated translational stalling. Science 326, 1412–1415 (2009)
Pulk, A. & Cate, J. H. Control of ribosomal subunit rotation by elongation factor G. Science 340, 1235970 (2013)
Vorstenbosch, E., Pape, T., Rodnina, M., Kraal, B. & Wintermeyer, W. The G222D mutation in elongation factor Tu inhibits the codon-induced conformational changes leading to GTPase activation on the ribosome. EMBO J. 15, 6766–6774 (1996)
Daviter, T., Wieden, H.-J. & Rodnina, M. V. Essential role of histidine 84 in elongation factor Tu for the chemical step of GTP hydrolysis on the ribosome. J. Mol. Biol. 332, 689–699 (2003)
Voorhees, R. M., Schmeing, T. M., Kelley, A. C. & Ramakrishnan, V. The mechanism for activation of GTP hydrolysis on the ribosome. Science 330, 835–838 (2010)
Pintilie, G. D., Zhang, J., Goddard, T. D., Chiu, W. & Gossard, D. C. Quantitative analysis of cryo-EM density map segmentation by watershed and scale-space filtering, and fitting of structures by alignment to regions. J. Struct. Biol. 170, 427–438 (2010)
Selmer, M., Gao, Y.-G., Weixlbaumer, A. & Ramakrishnan, V. Ribosome engineering to promote new crystal forms. Acta Crystallogr. D 68, 578–583 (2012)
Acknowledgements
We thank F. Würriehausen for expert technical assistance and M. Lüttich and B. Busche for support in high-performance computation and programming. The work was supported by the Deutsche Forschungsgemeinschaft Grant FOR 1805 (to H.S. and M.V.R.) and by the Sonderforschungsbereich 860 (to H.S., R. F., and M.V.R.).
Author information
Authors and Affiliations
Contributions
N.F. designed and performed cryo-EM experiments and data analysis. P.N. conceived and performed pseudo-crystallographic refinement and model validation and analysed data. A.L.K. prepared ribosome complexes. L.V.B. performed and analysed molecular dynamics simulations. All authors discussed the results. H.S. and N.F. conceived the project and wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Aberration-corrected cryo-EM.
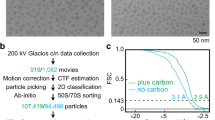
a, Exemplary Zemlin tableau (left) and phase diagram (right) as obtained for the present data set with the CEOS software by correcting electron optical aberrations using the Cs corrector. The resulting phase errors were less than 45° at ≤2.1 Å (that is, at scattering angles of 12 to 15 mrad) over up to 36 h of image acquisition. The main limiting aberration is axial coma (B2) and the next limiting aberration would be threefold astigmatism (A3). b, Local correction for the contrast transfer function. From micrographs (left) areas with individual ribosome particles (yellow frames) were extracted and local power spectra were computed for each of these areas by fast Fourier transform algorithms (FFT). Local power spectra were subjected to principal component analysis (PCA) and classification to average power spectra with similar contrast transfer function parameters that were obtained from different micrographs. Class averages of power spectra reveal an improved signal-to-noise ratio in Thon rings which are clearly visible up to 2.4 Å (right). c, Global power spectrum from a single micrograph showing Thon rings up to 3.5 Å.
Extended Data Figure 2 Hierarchical sorting of ribosome particle images.
Ribosome particles were sorted in three steps according to: (1) global ribosome conformation (C1), that is, 30S body rotation7; and (2) ligand occupancy (C2)35 and particle quality (C3)37(Methods). The asterisk denotes particles assigned to the largest 30S body rotations ≤−10° and ≥10° which contain particles with extreme 30S rotation angles, but also low-quality particle images.
Extended Data Figure 3 Resolution curves and model validation of the E. coli 70S ribosome–EF-Tu cryo-EM structure.
a, Fourier-shell correlation (FSC) curve (black) for the 70S ribosome cryo-EM reconstruction computed between the masked independent half-maps (half1 and half2) that were obtained by so-called ‘gold-standard’ refinement in RELION37. The resolution of the cryo-EM reconstruction is ∼2.9 Å according to the 0.143 criterion63 (black dashed line). b, FSC curves computed between cryo-EM maps and model maps generated from refined atomic coordinates. The vertical black dashed line indicates the maximum resolution at which the full atomic models were refined. Black, the FSC curve between the final cryo-EM map (map) and the final model (model); blue, the FSC curve between half map 1 (half1) and the model obtained by refinement only against half map 1 (model1); red, the FSC curve between half map 1 and the model obtained by refinement only against half map 2 (model2). c, FSC curves (FSCwork) between reflections from solvent-flattened cryo-EM map and model as obtained by pseudo-crystallographic refinement of the complete ribosome model (mask1) and three sub-models using different masks corresponding to local variations in resolution (mask2–4; Methods) as shown in h. Coloured numbers indicate the highest resolution used in refinement with the respective mask as indicated by the colour code. For all refinements, the FSC is above the 0.5 threshold (black dashed line) in the highest-resolution shell. Differences to b result largely from solvent-flattening before Fourier transformation for refinement (Methods). d–g, CCwork and Rwork as obtained by refinement using the respective mask (see labels). For a reliable resolution estimate CCwork (ref. 8) is expected to be >0.2 and Rwork <0.51 in the highest-resolution shell. h, Isosurface representations of the mask used for local refinements; ‘%’ indicates the fraction of atoms of the complete model entailed in the refinement with the respective mask.
Extended Data Figure 4 Modifications in rRNA. Comparison between cryo-EM and X-ray crystallography.
a, Experimental densities. In each row density maps for the same type of rRNA modification are shown (from left to right): for the present cryo-EM map and for the current best resolved bacterial and archaeal ribosome maps determined by X-ray crystallography, that is, the bacterial 70S ribosome from E. coli (Eco70S) at 2.8 Å resolution5 (PDB IDs: 4TPA and 4TPB); the bacterial 70S ribosome from T. thermophilus (Tth70S) at 2.4 Å resolution9 (PDB IDs: 4RB5 and 4RB6); and the archaeal 50S subunit from Haloarcula marismortui (Hma70S) at 2.2 Å resolution10 (PDB ID: 1VQ0). E. coli numbering is used for bacterial ribosome structures. Locations of rRNA modifications as determined by biochemical data are marked by yellow circles, modifications not observed in the density maps are denoted by red arrows and the black arrow designates the non-planarity of dihydrouridine observed in the cryo-EM map. b, Model-based densities for m62A1518 and m62A1519 showing slight differences due to scattering properties. Densities were computed in CCTBX64 at 2.65 Å resolution from our final model with atomic-displacement factors kept unchanged using electron (e− scattering, purple) and X-ray scattering factors (X-ray scattering, blue), respectively. Map thresholds were normalized to show similar density levels for the electron-rich phosphate groups. Accordingly, the absence of densities for modifications in crystallographic maps also at higher resolutions may result from differences in electron and X-ray scattering and in data quality which is affected, for example, by local and global disorder.
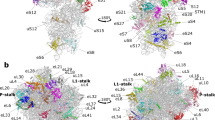
Extended Data Figure 5 rRNA modifications in the E. coli 70S ribosome.
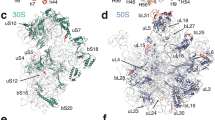
a, Stabilizing effects of 16S rRNA methyl groups in the P site of the decoding centre. Numbers in the overview (top left) mark the positions of close-ups (1–3), which show the interactions of the rRNA methyl groups with distances colour-encoded by dashed lines. Close-up 1: Stacking network of m2G966 and m5C967 stabilizing binding of initiator tRNA. Close-ups 2, 3: rRNA modifications impacting mRNA binding. The universally conserved bulky dimethylamine groups of m62A1518 and m62A1519 stabilize their direct environment by steric encumbrance explaining their requirement for correct packing of 16S rRNA helices 24a, 44 and 4565. In particular, the dimethylamine of m62A1519 is involved in medium and long-range repulsive interactions with the backbone of m3U1498 and the 2' O of C1520, while its conformation is mostly determined by the dimethylamine moiety of the adjacent m62A1518 which, in turn, is fixed by short repulsive interactions with O6 of G1517 and O4 of U793. The additional methyl groups of m62A1519 interact with A792, which provides part of the binding site for the antibiotic kasugamycin15, accounting for the resistance against kasugamycin upon demethylation of m62A151966. Furthermore, m62A1518 and m62A1519 impact initiation67 possibly via m3U1498 whose backbone interacts with m62A1519, while its modified base contacts the mRNA backbone. The methyl groups of m3U1498 and m4Cm1402 form part of the binding site for the initiation codon and modulate translation initiation13,14 by steric encumbrance and/or by preventing direct hydrogen bonds with the mRNA backbone. b, Constitutive rRNA modifications in the E. coli 70S ribosome (list adapted from ref. 11 and references therein). c, rRNA modifications in the A site of the decoding centre and helix 69 of 23S rRNA (H69). The binding site of aminoglycosides in helix 44 of 16S rRNA (h44)—including N4 of m5C1407—is indicated for neomycin B (magenta, superposition from PDB ID: 2ET4)68. The three pseudouridines stabilizing H6969 by enhancing base stacking70 are depicted in blue. The methyl group (yellow) on m3Ψ1917 in H69 prevents potential base-pairing with A1913, a residue important for uniform tRNA selection71. Note the flipped out conformation of A1913 facilitating interaction with the 2′ OH of m2s6iA37 of the distorted Phe–tRNAPhe (purple), which, in turn, stacks onto A36 of the tRNA anticodon. d, Methyl group on m2G1835 of 23S rRNA enhancing subunit association. The four helices of 23S rRNA that intersect around residue m2G1835 and form intersubunit bridges B2b and B2c with helices 24 (h24) and 45 (h45) of 16S rRNA (dark grey) are denoted in different colours: helix 67 (H67), light blue; helix 68 (H68), blue; helix 69 (H69), teal; helix 70 (H70), purple. Inset, contacts of the methyl group on m2G1835 with adjacent residues which, in turn, interact with 16S rRNA. e, Cluster of 23S rRNA modifications in the peptide exit tunnel. The modified rRNA residues, the functionally important nearby tip of protein L22 (teal), P-site fMet–tRNAfMet (green) and a model of the nascent peptide chain (pink, superposition from PDB ID: 2WWL)72 are indicated.
Extended Data Figure 6 Visualization of structural dynamics of the ribosome by different approaches.
a–d, In each panel, the ribosome is shown from the factor binding site on the left and in cut-away view on the right; h denotes the head and b the body of the 30S ribosomal subunit. a, Present cryo-EM map coloured according to local resolution as determined by Resmap2. b, Present cryo-EM map coloured according to the B factors obtained from the pseudo-crystallographic atomic model refinement (Methods). c, Model map of the 2.95 Å crystal structure of the E. coli 70S ribosome73 (PDB IDs: 4KJ1 and 4KJ2) coloured according to respective B factors. The black arrow denotes the stabilization of the 30S head region by crystal contacts, whereas the 30S body (white arrow) is less constrained by crystal contacts and shows higher B factors, indicating larger flexibility for this region. d, Snapshot from molecular dynamics trajectory of the E. coli 70S ribosome coloured according to root mean squared fluctuations (RMSFs) obtained from the full 2 µs explicit solvent molecular dynamics simulation (Methods). Note the large fluctuations of the 30S head and body (white arrows) of the ribosome in solution not constrained by crystal contacts.
Extended Data Figure 7 Structure of E. coli EF-Tu–Phe–tRNAPhe bound to the ribosome.
a, Detailed comparison of the distorted A/T-site–tRNA interactions between the E. coli and T. thermophilus ribosome–EF-Tu–kirromycin complexes6,26 and the free E. coli tRNAPhe51. We found significant differences in tRNA conformation and interactions implicated in the GTPase activation mechanism26 that correlate with ribosome binding and differences in organism and tRNA species. Here and below residue numbers refer to E. coli. b, Overview of the E. coli ribosome–EF-Tu structure. The residues interacting with the EF-Tu ternary complex (depicted in stick representation) generally agree with those seen in the T. thermophilus structures6,26. rRNA helices are denoted as: h44, helix 44 of 16S rRNA; H69, helix 69; and SRL, sarcin–ricin loop of 23S rRNA. The dashed boxes indicate the parts of the structure magnified in c and d. c, Structural differences in an important ribosome–EF-Tu interaction. In the E. coli structure (Eco, left panel) residues A55 and A368 of 16S rRNA assume different conformations (cyan arrows) and interact differently with the ribosome and EF-Tu–tRNA complex than in the T. thermophilus structure (Tth, right panel, PDB ID: 2WRN)26. Furthermore, the differences in EF-Tu sequence result in slightly different ribosome–EF-Tu interactions, for example, the ε-amino group of lysine 282 in E. coli EF-Tu is within hydrogen-bonding distance of G382 in 16S rRNA, but not the serine at this position in T. thermophilus EF-Tu. Inset on left panel, overlay of the crucial β-turn26,74 in EF-Tu domain 2 in the free (PDB ID: 1OB2, R. C. Nielsen et al., unpublished data) and ribosome-bound state from E. coli and T. thermophilus. d, Dynamics of the catalytic histidine 84 (ref. 75) of EF-Tu. A split density for the side chain of histidine 84 (data not shown) indicates the presence of two rotamers (rot1 and rot2, panel 1) in the present E. coli complex. In d, the T. thermophilus ribosome-bound EF-Tu structures6,26,76 (dark grey, complex as indicated) are shown with the corresponding rotamer of the present structure (red). Residues valine 20 and isoleucine 60 of the hydrophobic gate76 are denoted; isoleucine 60 is not resolved in the kirromycin-stalled ribosome–EF-Tu complexes.
Extended Data Figure 8 Cryo-EM densities for mobile proteins L9 and L31 and the arrangement of L9 in polysomes.
Densities in a and b were obtained by semi-automatic segmentation using the ‘segger’ tool in UCSF CHIMERA41,77 and normalized and low-pass filtered according to local resolution estimates. a, Cryo-EM map and models of protein L9 (∼4 Å local resolution, rendered at 1σ). b, Cryo-EM maps and models of protein L31 in the ground-state of the ribosome (left, ∼4.3 Å local resolution) and in the rotated state (right, ∼6 Å local resolution); maps were rendered at 1.5σ. c, Model of protein L9 in the context of polysomes. Overviews show the arrangement of neighbouring ribosomes (i-1 and i) in the major t-t form of E. coli polysomes as obtained by fitting the present 70S ribosome structure into the cryo-electron tomography reconstruction27 in UCSF CHIMERA41. Left close-up, the conformer of L9 as seen in the present cryo-EM map (L9 cryo-EM, blue) is located close to protein S4 of the neighbouring 30S subunit (30S i) according to the polysome model. The purple arrow indicates the rearrangement of L9 from the cryo-EM conformation to that seen in crystals. The black arrow denotes the location of the mRNA entry channel in the 30S subunit i. Right close-up, the conformer of L9 as seen in the context of ribosome crystals (L9 X-ray, pink) reaches into the ribosomal A-site of the neighbouring 30S subunit and would be compatible with the simultaneous binding of elongation factors in the polysome model. In crystals, protein L9 precludes the binding of elongation factors due to the tighter packing of ribosomes78. The model of L9 was obtained by superposition of the E. coli 70S ribosome X-ray structure5 (PDB IDs: 4TP8 and 4TP9) onto ribosome i-1 in the polysome model using UCSF CHIMERA41.
Rights and permissions
About this article
Cite this article
Fischer, N., Neumann, P., Konevega, A. et al. Structure of the E. coli ribosome–EF-Tu complex at <3 Å resolution by Cs-corrected cryo-EM. Nature 520, 567–570 (2015). https://doi.org/10.1038/nature14275
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14275
This article is cited by
-
Geometric alignment of aminoacyl-tRNA relative to catalytic centers of the ribosome underpins accurate mRNA decoding
Nature Communications (2023)
-
The translating bacterial ribosome at 1.55 Å resolution generated by cryo-EM imaging services
Nature Communications (2023)
-
Cardiovascular Drugs: an Insight of In Silico Drug Design Tools
Journal of Pharmaceutical Innovation (2022)
-
In silico analysis of bacterial translation factors reveal distinct translation event specific pI values
BMC Genomics (2021)
-
Quantitative profiling of pseudouridylation dynamics in native RNAs with nanopore sequencing
Nature Biotechnology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.