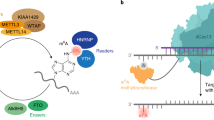
Abstract
RNA-binding proteins control many aspects of cellular biology through binding single-stranded RNA binding motifs (RBMs)1,2,3. However, RBMs can be buried within their local RNA structures4,5,6,7, thus inhibiting RNA–protein interactions. N6-methyladenosine (m6A), the most abundant and dynamic internal modification in eukaryotic messenger RNA8,9,10,11,12,13,14,15,16,17,18,19, can be selectively recognized by the YTHDF2 protein to affect the stability of cytoplasmic mRNAs15, but how m6A achieves its wide-ranging physiological role needs further exploration. Here we show in human cells that m6A controls the RNA-structure-dependent accessibility of RBMs to affect RNA–protein interactions for biological regulation; we term this mechanism ‘the m6A-switch’. We found that m6A alters the local structure in mRNA and long non-coding RNA (lncRNA) to facilitate binding of heterogeneous nuclear ribonucleoprotein C (HNRNPC), an abundant nuclear RNA-binding protein responsible for pre-mRNA processing20,21,22,23,24. Combining photoactivatable-ribonucleoside-enhanced crosslinking and immunoprecipitation (PAR-CLIP) and anti-m6A immunoprecipitation (MeRIP) approaches enabled us to identify 39,060 m6A-switches among HNRNPC-binding sites; and global m6A reduction decreased HNRNPC binding at 2,798 high-confidence m6A-switches. We determined that these m6A-switch-regulated HNRNPC-binding activities affect the abundance as well as alternative splicing of target mRNAs, demonstrating the regulatory role of m6A-switches on gene expression and RNA maturation. Our results illustrate how RNA-binding proteins gain regulated access to their RBMs through m6A-dependent RNA structural remodelling, and provide a new direction for investigating RNA-modification-coded cellular biology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
17 March 2015
The received date year was corrected.
References
Antson, A. A. Single-stranded-RNA binding proteins. Curr. Opin. Struct. Biol. 10, 87–94 (2000)
Dreyfuss, G., Kim, V. N. & Kataoka, N. Messenger-RNA-binding proteins and the messages they carry. Nature Rev. Mol. Cell Biol. 3, 195–205 (2002)
Ray, D. et al. A compendium of RNA-binding motifs for decoding gene regulation. Nature 499, 172–177 (2013)
Wan, Y. et al. Landscape and variation of RNA secondary structure across the human transcriptome. Nature 505, 706–709 (2014)
Ding, Y. et al. In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features. Nature 505, 696–700 (2014)
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010)
Rouskin, S., Zubradt, M., Washietl, S., Kellis, M. & Weissman, J. S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505, 701–705 (2014)
Bokar, J. A. in Fine-Tuning of RNA Functions by Modification and Editing (ed. Grosjean, H. ) 141–178 (Springer, 2005)
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nature Chem. Biol. 7, 885–887 (2011)
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013)
Liu, J. et al. A METTL3–METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nature Chem. Biol. 10, 93–95 (2014)
Wang, Y. et al. N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nature Cell Biol. 16, 191–198 (2014)
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012)
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012)
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014)
Fustin, J. M. et al. RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155, 793–806 (2013)
Schwartz, S. et al. High-resolution mapping reveals a conserved, widespread, dynamic mRNA methylation program in yeast meiosis. Cell 155, 1409–1421 (2013)
Batista, P. J. et al. m6A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014)
Zhao, X. et al. FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis. Cell Res. 24, 1403–1419 (2014)
König, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nature Struct. Mol. Biol. 17, 909–915 (2010)
McCloskey, A., Taniguchi, I., Shinmyozu, K. & Ohno, M. hnRNP C tetramer measures RNA length to classify RNA polymerase II transcripts for export. Science 335, 1643–1646 (2012)
Rajagopalan, L. E., Westmark, C. J., Jarzembowski, J. A. & Malter, J. S. hnRNP C increases amyloid precursor protein (APP) production by stabilizing APP mRNA. Nucleic Acids Res. 26, 3418–3423 (1998)
Zarnack, K. et al. Direct competition between hnRNP C and U2AF65 protects the transcriptome from the exonization of Alu elements. Cell 152, 453–466 (2013)
Cieniková, Z. et al. Structural and mechanistic insights into poly(uridine) tract recognition by the hnRNP C RNA recognition motif. J. Am. Chem. Soc. 136, 14536–14544 (2014)
Liu, N. et al. Probing N6-methyladenosine RNA modification status at single nucleotide resolution in mRNA and long noncoding RNA. RNA 19, 1848–1856 (2013)
Krecic, A. M. & Swanson, M. S. hnRNP complexes: composition, structure, and function. Curr. Opin. Cell Biol. 11, 363–371 (1999)
Görlach, M., Burd, C. G. & Dreyfuss, G. The determinants of RNA-binding specificity of the heterogeneous nuclear ribonucleoprotein C proteins. J. Biol. Chem. 269, 23074–23078 (1994)
Kierzek, E. & Kierzek, R. The thermodynamic stability of RNA duplexes and hairpins containing N6-alkyladenosines and 2-methylthio-N6-alkyladenosines. Nucleic Acids Res. 31, 4472–4480 (2003)
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010)
Anders, S., Reyes, A. & Huber, W. Detecting differential usage of exons from RNA-seq data. Genome Res. 22, 2008–2017 (2012)
Ehresmann, C. et al. Probing the structure of RNAs in solution. Nucleic Acids Res. 15, 9109–9128 (1987)
Peterson, E. T., Pan, T., Coleman, J. & Uhlenbeck, O. C. In vitro selection of small RNAs that bind to Escherichia coli phenylalanyl-tRNA synthetase. J. Mol. Biol. 242, 186–192 (1994)
Corcoran, D. L. et al. PARalyzer: definition of RNA binding sites from PAR-CLIP short-read sequence data. Genome Biol. 12, R79 (2011)
Dominissini, D., Moshitch-Moshkovitz, S., Salmon-Divon, M., Amariglio, N. & Rechavi, G. Transcriptome-wide mapping of N6-methyladenosine by m6A-seq based on immunocapturing and massively parallel sequencing. Nature Protocols 8, 176–189 (2013)
Lohse, M. et al. RobiNA: a user-friendly, integrated software solution for RNA-Seq-based transcriptomics. Nucleic Acids Res. 40, W622–W627 (2012)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Elemento, O., Slonim, N. & Tavazoie, S. A universal framework for regulatory element discovery across all genomes and data types. Mol. Cell 28, 337–350 (2007)
Ouyang, Z., Snyder, M. P. & Chang, H. Y. SeqFold: genome-scale reconstruction of RNA secondary structure integrating high-throughput sequencing data. Genome Res. 23, 377–387 (2013)
Xiao, Y. et al. A novel significance score for gene selection and ranking. Bioinformatics 30, 801–807 (2014)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009)
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nature Protocols 7, 562–578 (2012)
Huelga, S. C. et al. Integrative genome-wide analysis reveals cooperative regulation of alternative splicing by hnRNP proteins. Cell Rep. 1, 167–178 (2012)
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009)
Pollard, K. S., Hubisz, M. J., Rosenbloom, K. R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 20, 110–121 (2010)
Yang, F., Yi, F., Han, X., Du, Q. & Liang, Z. MALAT-1 interacts with hnRNP C in cell cycle regulation. FEBS Lett. 587, 3175–3181 (2013)
Acknowledgements
This work is supported by National Institutes of Health EUREKA GM088599 (C.H. and T.P.), and by K01HG006699 (Q.D.). We thank all members of the Pan and He laboratories for comments and discussions. We also thank Y. C. Leung, G. Perdrizet, Y. Pigli, J. Yue, J. Liu, Y. Yue, K. Chen and M. Yu for technical assistance. C.H. is an Investigator of the Howard Hughes Medical Institute. M.P. was a Natural Sciences and Engineering Research Council of Canada postdoctoral fellow.
Author information
Authors and Affiliations
Contributions
N.L., G.Z. and M.P. designed and performed experiments, and analysed data. Q.D. synthesized all RNA oligonucleotides. N.L., M.P. and T.P. conceived the project. N.L. and T.P. wrote the paper with input from C.H. and M.P.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 m6A increases the accessibility of the U-tract to enhance HNRNPC binding.
a, Secondary structure of the MALAT1 hairpin with m6A methylation at the 2,577 site shown in red25. Nucleotide position numbers correspond to their locations along the human MALAT1 transcript (NCBI accession NR_002819). b, RNA pull down showing that HNRNPC preferably binds methylated RNA. c, The list of proteins with identified peptides by mass spectrometry in b. d, Recombinant HNRNPC1 binds more strongly with that MALAT1 2,577–m6A hairpin compared with the unmethylated hairpin, as determined by an in vitro ultraviolet crosslinking assay23. e, HNRNPC shows binding around the 2,577–A site along MALAT1 in vivo, as determined by previously published HNRNPC iCLIP data20. The underlying genomic sequence is shown at the bottom with a red square marking the 2,577–m6A site. The slight shift of the iCLIP signal to upstream of the U-tract-binding site is probably due to the steric hindrance of the peptide fragment remaining on RNA, which can cause reverse transcription to terminate more than one nucleotide upstream of the crosslink site20. f, Quantification of the RNase V1 cleavage signal for the U-tract region from the RNA structural mapping assay in Fig. 1e. To correct for sample loading difference, each band signal was normalized to the band signal of the immediate 3′ residue to the U-tract. Data are mean ± s.d.; n = 3, technical replicates. g, Quantitative analysis of the RNase T1 cleavage signal from the RNA structural mapping assay in Fig. 1e. An increased RNase T1 cleavage signal (single-strand specific and cleavage after guanosines) was observed due to the surrounding m6A residue. To correct for sample loading difference, the ratio for each band signal among all bands in each lane was calculated. Relative T1 cleavage = (m6Anative/m6Adenature)/(Anative/Adenature). n = 2, technical replicates. h, Quantitative CMCT mapping showing increased signals for the U-tract bases around the U base-pairing with m6A. Quantitation of band signals within the U-tract region is shown on the right. Data are mean ± s.d.; n = 4, technical replicates.
Extended Data Figure 2 Increased accessibility of U-tracts enhances HNRNPC binding.
a, Structure probing of the 2,577–A-to-U mutated MALAT1 hairpin (2,577–U). The annotation is the same as in Fig. 1d. b, Quantification of the RNase V1 cleavage signal for the U-tract region from RNA structural mapping assays as in a. To correct for sample loading difference, each band signal was normalized to the band signal of the 3′-most U of the U-tract. n = 2, technical replicates. c, Filter-binding curves displaying the binding affinities between recombinant HNRNPC1 and 2,577–U/A oligonucleotides. Data are mean ± s.d.; n = 3, technical replicates. d, Filter-binding results showing the binding affinities between recombinant HNRNPC1 and four mutated MALAT1 oligonucleotides. (1) Mutate G–C to C–C, 2,577–A: predicted to weaken the hairpin stem and increase HNRNPC binding. Results: binding improved from 722 nM Kd to 142 nM (fivefold). (2) Mutate G–C to C–C, 2,577–m6A: in this context of weaker stem, m6A is predicted to confer a smaller effect compared to wild-type hairpin. Result: improved binding only twofold instead of eightfold. (3) Restore C–C to C–G, 2,577–A: predicted to restore the hairpin stem and decrease HNRNPC binding compared to C–C mutant. Result: binding decreased by 6.4-fold. (4) Restore C–C to C–G, 2,577–m6A: in this context of restored stem, m6A is again predicted to confer increased binding compared to 2,577–A hairpin. Result: improved binding by 2.5-fold. Data are mean ± s.d.; n = 3 each, technical replicates. e, RNA alkaline hydrolysis terminal truncation assay showing recombinant HNRNPC1 binding to terminal truncated MALAT1 hairpin oligonucleotides (2,577 site m6A methylated or unmethylated). In this assay, 3′-radiolabelled MALAT1 2,577 hairpin oligonucleotides were terminal truncated by alkaline hydrolysis into RNA fragments that were then incubated with HNRNPC1 protein followed by filter binding wash steps. The remaining RNA on the filter paper was isolated and analysed by denaturing gel electrophoresis, as indicated in the lane ‘C1-bound or C1-B’. ‘Input’ refers to alkaline-hydrolysis-truncated RNA oligonucleotides used for incubation with hnRNP C1; ‘G-L or G-ladder’ was generated from RNase T1 digestion; ‘Ctrl’ refers to the intact MALAT1 hairpin without alkaline hydrolysis truncation. One pair of methylated/unmethylated truncated oligonucleotides (CUT1, marked by green arrows) was selected for subsequent biochemical analysis, due to their strong interaction with HNRNPC1. f, RNA terminal truncation assay as in e except 5′ 32P-labelled oligonucleotides were used. One pair of methylated/unmethylated truncated oligonucleotides (CUT2, marked by green arrows) was selected for subsequent biochemical analysis. g, Structure probing of the CUT1 oligonucleotides using RNase V1 and nuclease S1 digestion. Annotation is the same as in Fig. 1e. The red dot marks the m6A site and the red line marks the U-tract region. h, Structure probing of the CUT2 oligonucleotides using RNase V1 and nuclease S1 digestion. Annotation is the same as in g. i, Truncated oligonucleotides with exposed U-tracts increased HNRNPC binding regardless of m6A. Data are mean ± s.d.; n = 3, technical replicates.
Extended Data Figure 3 m6A is enriched in the vicinity of HNRNPC-binding sites.
a, Schematic diagram of the CLIP-2dTLC protocol. IP, immunoprecipitation; nt, nucleotide; UV, ultraviolet. The RNase T1 used in our 2dTLC assay cleaves single-stranded RNA after guanosines, so the m6A/A ratio determined here represents the m6A fraction of all adenosines following guanosines. b, Analysis of crosslinked RNA–HNRNPC complexes (CLIP RNP) using denaturing gel electrophoresis (lanes 1 and 2). Positions of the protein size standards are shown on the left. HNRNPC IP RNA region (RNA samples within RNA–HNRNPC crosslinked complexes) were extracted from the gel slices marked by the red rectangle. c, Denaturing gel analysing the size distribution for the HNRNPC PAR-CLIP RNA samples (lane 2). The RNA size standards were loaded in lanes 1 and 3.
Extended Data Figure 4 PAR-CLIP–MeRIP identifies transcriptome-wide m6A-switches in the vicinity of HNRNPC-binding sites.
a, Density plots illustrating the distribution of distance between the PAR-CLIP–MeRIP/input peaks and the nearest GRACH motif (top) or the nearest U-tracts (bottom). b, Definition and identification of HNRNPC m6A-switches based on the PAR-CLIP–MeRIP analysis. Approximately 89% of PAR-CLIP–MeRIP peaks harbouring both the U-tract and RRACH motifs have an RRACH–U-tract inter-motif distance within 50 nucleotides, significantly higher than the 64% of such coupling within the genomes. HNRNPC m6A-switches are identified as m6A-methylated RRACH–U-tract coupling events. c, Volcano plot depicting all coupling events (open circles) as defined in b, according to their P values36 (P; y axis) and fold-change values at RRACH sites (E; x axis). To identify HNRNPC m6A-switches, we generated the π value, π = E·(−log10P), as one comprehensive parameter to pick meaningful genomic loci37. HNRNPC m6A-switches identified from PAR-CLIP–MeRIP experiments should fulfil the following requirements: (1) read counts at both the control and IP sample ≥ 5; (2) π value ≥ 0.627, corresponding to FDR ≤ 5%. d, Pie chart depicting the region distribution of HNRNPC m6A-switches identified by PAR-CLIP–MeRIP. e, Pie chart depicting HNRNPC PAR-CLIP peaks. These are enriched in introns, consistent with previous reports that HNRNPC binds mainly nascent transcripts19,23,25.
Extended Data Figure 5 Validation of two identified m6A-switches.
a, b, PAR-CLIP–MeRIP data detected positive IP/input enrichment at the RRACH sites (red arrowheads) on the DNAJC25-GNG10 gene (a) and HNRNPH1 gene (b) in HEK293T cells. c, d, Quantification of RNase V1 cleavage signals around the U-tract region of m6A-switches on the DNAJC25-GNG10 (c) and HNRNPH1 (d) transcript, related to Fig. 2g, h. Data are mean ± s.d.; n = 3, technical replicates each. e, Quantitative CMCT mapping of DNAJC25-GNG10 m6A-switch shows increased band signals around the uridine base that pairs with m6A. The red vertical line marks the U-tract region. Quantitation of band signal for the U-tract region is shown on the right. Data are mean ± s.d.; n = 3, technical replicates. The HNRNPH1 m6A-switch hairpin is not suitable for CMCT probing, because its reverse transcription binding primer region is too short. f, g, In vivo DMS mapping of the DNAJC25-GNG10 hairpin (f) and HNRNPH1 (g); data are from ref. 7. A and C residues are marked with orange dots and the m6A residue is marked with a red dot. The hairpin loops are indicated by red bars. h, Transcriptome-wide S1/V1 mapping around the HNRNPH1 m6A-switch site. Blue bars represent V1 signal; magenta bars represent S1 signal. The hairpin loop is indicated by a red bar; data are from ref. 4. Not enough reads could be collected to make a plot for the DNAJC25-GNG10 m6A-switch region.
Extended Data Figure 6 Molecular features of high-confidence m6A-switches.
a, Western blot (WB) showing stable HNRNPC protein abundance upon METTL3/L14 knockdown. b, Volcano plot of the METTL3/L14 knockdown (KD) data depicting RRACH–U-tract coupling events (open red circles) as defined in Extended Data Fig. 4b, according to their P values38 (P; y axis) and fold-change values at the U-tracts (E; x axis). c, Overlap of RRACH–U-tract coupling events with decreased HNRNPC binding by METTL3 and METTL14 knockdown. d, The intron fraction of HCS m6A-switches in coding RNA and non-coding RNA. e, Density plot displaying the distribution of exonic m6A-switches/HNRNPC PAR-CLIP peaks according to exon length. f, Inter-motif (RRACH–U-tract) distance distributions suggest that m6A-switches have a preference for shorter distances between the RRACH and U-tract (>5×U) motifs. The distribution curves are from PAR-CLIP–MeRIP data (green), METTL3/L14 knockdown (red) and high-confidence (HCS) m6A-switches (black). g, Analysis of the inter-motif (U-tract–U-tract) distance patterns, previously identified by iCLIP20, in PAR-CLIP–MeRIP, METTL3/L14 knockdown and high-confidence m6A-switch data. The peaks at ∼165 and ∼300 nucleotides are clearly present. For the 2,798 high-confidence switches, we analysed those in which the other U-tract motif is also in a PAR-CLIP-identified sequence; the long-range peaks seem to have shifted to longer distances (∼220 and ∼370 nucleotides). h, METTL3/L14 knockdown does not affect the inter-motif (U-tract–U-tract) distance distributions for U-tracts (≥5× U) in HEK293T cells. i, EVOfold analysis for the 2,798 high-confidence m6A-switches. The chances for high-confidence m6A-switches to have EVOfold records are significantly higher than random genomic sequences. We first calculated the number of high-confidence sites in the EVO database if occurring in random to be ∼1.7. We found that 18 high-confidence sites are present in the EVO database, resulting in ∼11× enrichment. This result is further divided into intronic and exonic regions.
Extended Data Figure 7 m6A-switches regulate the abundance of target mRNAs.
a, HNRNPC, METTL3/L14 knockdown confirmed by western blots. b, HNRNPC knockdown (KD) and METTL3/L14 knockdown co-regulated the expression of a large number of genes. Gene expression changes between control (Ctrl) and HNRNPC, HNRNPU, METTL3/L14 knockdown HEK293T cells were analysed by Cuffdiff2 (refs 38, 39), and the absolute numbers of differentially expressed genes are shown. HCS-containing genes refers to the 1,815 genes containing high-confidence m6A-switches. The RNA-seq data from HNRNPU knockdown HEK293T cells (Gene Expression Omnibus accession GEO34995 data set40) were analysed for comparison with a different mRNA-binding protein. HNRNPU did not show preferential interaction with the 2,577–m6A modified MALAT1 hairpin (Fig. 1b, c). c, GO analysis of the m6A-switch-containing genes whose expression levels were co-differentially regulated by HNRNPC and METTL3/L14 knockdown, against all m6A-switch-containing genes as background. d, An example of an m6A-switch among co-regulated transcripts is the ARHGAP5 transcript (NCBI accession NM_001030055). Its proposed secondary structure with the m6A methylation site in red is shown with the opposing the U-tract in a stem. e, f, PAR-CLIP–MeRIP detected positive IP/input enrichment at the RRACH site (red arrowhead) of the ARHGAP5 m6A-switch (e), while METTL3/L14 knockdown decreased HNRNPC binding at the U-tract (red square) of this m6A-switch (f). g, The expression level of the ARHGAP5 gene was co-upregulated by HNRNPC, METTL3/L14 knockdown, as shown by the RNA-seq data from HEK293T cells. The vertical black line represents the m6A-switch site. h, HNRNPC, METTL3/L14 knockdown decreased the proliferation rates of HEK293T cells to a similar extent. Data are mean ± s.d.; n = 4, biological replicates.
Extended Data Figure 8 m6A-switches regulate alternative splicing of target mRNAs.
a, Fold changes (knockdown (KD)/control (Ctrl), log2) in normalized exon expression against RNA-seq reads detect the exons in HNRNPC knockdown, METTL3 knockdown, METTL14 knockdown and control samples. Statistically significant differentially expressed exons (SSDEEs) called by DEXSeq are indicated in red. b, Proposed secondary structure of the CDS2 hairpin with the m6A methylation site shown in red, opposing the U-tract region. Nucleotide position numbers correspond to their locations along the human CDS2 transcript (NCBI accession NM_003818). c, Proposed secondary structure of the YTHDF2 hairpin with the m6A methylation site shown in red, opposing the U-tract region. Nucleotide position numbers correspond to their locations along the human YTHDF2 transcript (NM_001173128). d, e, PAR-CLIP–MeRIP detected a positive enrichment at the RRACH site (red arrowhead) (d), while METTL3/L14 knockdown decreased HNRNPC binding at the U-tract (red square) of this YTHDF2 m6A-switch (e). f, The inclusion level of one YTHDF2 exon is co-downregulated by HNRNPC knockdown, METTL3 knockdown and METTL14 knockdown, as validated by RT–PCR. Data are mean ± s.d.; n = 3, biological replicates. g, We analysed our polyA+ RNA-seq data to look for reads that span intron/exon junctions on CDS m6A-switch containing genes. We find that the control sample has significantly higher reads spanning intron/exon junctions than HNRNPC and METTL3/L14 knockdown samples. This result indicates that m6A depletion at the CDS m6A-switches promotes intron exclusion.
Extended Data Figure 9 Summary of the sequencing samples.
a, For PAR-CLIP–MeRIP and PAR-CLIP experiments from HEK293T cells, the number of mapped reads and ‘T-to-C’ mutation rates are given for each replicate. b, For RNA-seq experiments from HEK293T cells, the number of total reads, the number of mapped reads as well as the mapping rates is given for each replicate. c, Scatter plots comparing transcripts for all PAR-CLIP replicate experiments. The square of Spearman’s rank correlation value (r2) for each pair is shown in the top left corner of the respective panel. d, The detected expression level changes show a strong correlation between gene knockdown replicates. Scatter plots comparing the fold changes (log2) in normalized gene expression from replicates of HNRNPC, METTL3 and METTL14 knockdown. The square of Spearman’s rank correlation value (r2) for each pair is shown in the top left corner of the respective panel.
Rights and permissions
About this article
Cite this article
Liu, N., Dai, Q., Zheng, G. et al. N6-methyladenosine-dependent RNA structural switches regulate RNA–protein interactions. Nature 518, 560–564 (2015). https://doi.org/10.1038/nature14234
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14234
This article is cited by
-
The epigenetic downregulation of LncGHRLOS mediated by RNA m6A methylase ZCCHC4 promotes colorectal cancer tumorigenesis
Journal of Experimental & Clinical Cancer Research (2024)
-
The role of the methyltransferase METTL3 in prostate cancer: a potential therapeutic target
BMC Cancer (2024)
-
N6-methyladenosine-modified circ_104797 sustains cisplatin resistance in bladder cancer through acting as RNA sponges
Cellular & Molecular Biology Letters (2024)
-
Overcoming therapeutic resistance in oncolytic herpes virotherapy by targeting IGF2BP3-induced NETosis in malignant glioma
Nature Communications (2024)
-
LncRNA LY6E-DT and its encoded metastatic-related protein play oncogenic roles via different pathways and promote breast cancer progression
Cell Death & Differentiation (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.