Abstract
Long-standing evidence indicates that human immunodeficiency virus type 1 (HIV-1) preferentially integrates into a subset of transcriptionally active genes of the host cell genome1,2,3,4. However, the reason why the virus selects only certain genes among all transcriptionally active regions in a target cell remains largely unknown. Here we show that HIV-1 integration occurs in the outer shell of the nucleus in close correspondence with the nuclear pore. This region contains a series of cellular genes, which are preferentially targeted by the virus, and characterized by the presence of active transcription chromatin marks before viral infection. In contrast, the virus strongly disfavours the heterochromatic regions in the nuclear lamin-associated domains5 and other transcriptionally active regions located centrally in the nucleus. Functional viral integrase and the presence of the cellular Nup153 and LEDGF/p75 integration cofactors are indispensable for the peripheral integration of the virus. Once integrated at the nuclear pore, the HIV-1 DNA makes contact with various nucleoporins; this association takes part in the transcriptional regulation of the viral genome. These results indicate that nuclear topography is an essential determinant of the HIV-1 life cycle.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Wang, G. P., Ciuffi, A., Leipzig, J., Berry, C. C. & Bushman, F. D. HIV integration site selection: analysis by massively parallel pyrosequencing reveals association with epigenetic modifications. Genome Res. 17, 1186–1194 (2007)
Schroder, A. R. et al. HIV-1 integration in the human genome favors active genes and local hotspots. Cell 110, 521–529 (2002)
Brady, T. et al. HIV integration site distributions in resting and activated CD4+ T cells infected in culture. AIDS 23, 1461–1471 (2009)
Sherrill-Mix, S. et al. HIV latency and integration site placement in five cell-based models. Retrovirology 10, 90 (2013)
Guelen, L. et al. Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453, 948–951 (2008)
Cavalli, G. & Misteli, T. Functional implications of genome topology. Nature Struct. Mol. Biol. 20, 290–299 (2013)
Maldarelli, F. et al. HIV latency. Specific HIV integration sites are linked to clonal expansion and persistence of infected cells. Science 345, 179–183 (2014)
Wagner, T. A. et al. Proliferation of cells with HIV integrated into cancer genes contributes to persistent infection. Science 345, 570–573 (2014)
Kin, T. & Ono, Y. Idiographica: a general-purpose web application to build idiograms on-demand for human, mouse and rat. Bioinformatics 23, 2945–2946 (2007)
Nagai, S. et al. Functional targeting of DNA damage to a nuclear pore-associated SUMO-dependent ubiquitin ligase. Science 322, 597–602 (2008)
Hediger, F., Neumann, F. R., Van Houwe, G., Dubrana, K. & Gasser, S. M. Live imaging of telomeres: yKu and Sir proteins define redundant telomere-anchoring pathways in yeast. Curr. Biol. 12, 2076–2089 (2002)
Wu, C. et al. BioGPS: an extensible and customizable portal for querying and organizing gene annotation resources. Genome Biol. 10, R130 (2009)
Su, A. I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl Acad. Sci. USA 101, 6062–6067 (2004)
Lusic, M. et al. Proximity to PML nuclear bodies regulates HIV-1 latency in CD4+ T cells. Cell Host Microbe 13, 665–677 (2013)
Butler, S. L., Hansen, M. S. & Bushman, F. D. A quantitative assay for HIV DNA integration in vivo. Nature Med. 7, 631–634 (2001)
Manganaro, L. et al. Concerted action of cellular JNK and Pin1 restricts HIV-1 genome integration to activated CD4+ T lymphocytes. Nature Med. 16, 329–333 (2010)
Lu, R., Limon, A., Ghory, H. Z. & Engelman, A. Genetic analyses of DNA-binding mutants in the catalytic core domain of human immunodeficiency virus type 1 integrase. J. Virol. 79, 2493–2505 (2005)
Negri, D. R. et al. Successful immunization with a single injection of non-integrating lentiviral vector. Mol. Ther. 15, 1716–1723 (2007)
Shun, M. C. et al. LEDGF/p75 functions downstream from preintegration complex formation to effect gene-specific HIV-1 integration. Genes Dev. 21, 1767–1778 (2007)
Matreyek, K. A., Yucel, S. S., Li, X. & Engelman, A. Nucleoporin NUP153 phenylalanine-glycine motifs engage a common binding pocket within the HIV-1 capsid protein to mediate lentiviral infectivity. PLoS Pathog. 9, e1003693 (2013)
Cattoglio, C. et al. High-definition mapping of retroviral integration sites defines the fate of allogeneic T cells after donor lymphocyte infusion. PLoS ONE 5, e15688 (2010)
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007)
Capelson, M. et al. Chromatin-bound nuclear pore components regulate gene expression in higher eukaryotes. Cell 140, 372–383 (2010)
Kalverda, B., Pickersgill, H., Shloma, V. V. & Fornerod, M. Nucleoporins directly stimulate expression of developmental and cell-cycle genes inside the nucleoplasm. Cell 140, 360–371 (2010)
Vaquerizas, J. M. et al. Nuclear pore proteins nup153 and megator define transcriptionally active regions in the Drosophila genome. PLoS Genet. 6, e1000846 (2010)
Liang, Y., Franks, T. M., Marchetto, M. C., Gage, F. H. & Hetzer, M. W. Dynamic association of NUP98 with the human genome. PLoS Genet. 9, e1003308 (2013)
Light, W. H. et al. A conserved role for human Nup98 in altering chromatin structure and promoting epigenetic transcriptional memory. PLoS Biol. 11, e1001524 (2013)
Jordan, A., Bisgrove, D. & Verdin, E. HIV reproducibly establishes a latent infection after acute infection of T cells in vitro. EMBO J. 22, 1868–1877 (2003)
Brass, A. L. et al. Identification of host proteins required for HIV infection through a functional genomic screen. Science 319, 921–926 (2008)
Konig, R. et al. Global analysis of host-pathogen interactions that regulate early-stage HIV-1 replication. Cell 135, 49–60 (2008)
Connor, R. I., Chen, B. K., Choe, S. & Landau, N. R. Vpr is required for efficient replication of human immunodeficiency virus type-1 in mononuclear phagocytes. Virology 206, 935–944 (1995)
Wiznerowicz, M. & Trono, D. Conditional suppression of cellular genes: lentivirus vector-mediated drug-inducible RNA interference. J. Virol. 77, 8957–8961 (2003)
Dull, T. et al. A third-generation lentivirus vector with a conditional packaging system. J. Virol. 72, 8463–8471 (1998)
Cattoglio, C. et al. Hot spots of retroviral integration in human CD34+ hematopoietic cells. Blood 110, 1770–1778 (2007)
Repnik, U., Knezevic, M. & Jeras, M. Simple and cost-effective isolation of monocytes from buffy coats. J. Immunol. Methods 278, 283–292 (2003)
Daigle, N. et al. Nuclear pore complexes form immobile networks and have a very low turnover in live mammalian cells. J. Cell Biol. 154, 71–84 (2001)
Solovei, I. & Cremer, M. 3D-FISH on cultured cells combined with immunostaining. Methods Mol. Biol. 659, 117–126 (2010)
Han, Y., Wind-Rotolo, M., Yang, H. C., Siliciano, J. D. & Siliciano, R. F. Experimental approaches to the study of HIV-1 latency. Nature Rev. Microbiol. 5, 95–106 (2007)
Ikeda, T., Shibata, J., Yoshimura, K., Koito, A. & Matsushita, S. Recurrent HIV-1 integration at the BACH2 locus in resting CD4+ T cell populations during effective highly active antiretroviral therapy. J. Infect. Dis. 195, 716–725 (2007)
Liu, H. et al. Integration of human immunodeficiency virus type 1 in untreated infection occurs preferentially within genes. J. Virol. 80, 7765–7768 (2006)
van Helden, J. Regulatory sequence analysis tools. Nucleic Acids Res. 31, 3593–3596 (2003)
Wang, Z. et al. Genome-wide mapping of HATs and HDACs reveals distinct functions in active and inactive genes. Cell 138, 1019–1031 (2009)
Acknowledgements
This work was supported by grants from the Italian National Research Programme on AIDS of the Istituto Superiore di Sanità, Italy, to M.G. and M.L. and from the Young Investigator Grant RF2007-16 of the Italian Ministry of Health to M.L. The authors are grateful to S. Kerbavcic for editorial assistance.
Author information
Authors and Affiliations
Contributions
B.M., B.L., K.L. and M.L. performed the immuno-DNA FISH and ChIP experiments; A.K.-F., B.M., S.P., M.L. and M.G. analysed the data; A.K.-F., B.M. and S.P. performed the bioinformatics analysis; H.A. and M.L. performed the experiments using infectious virus; L.M. generated and analysed integrase-defective HIV-1 molecular clones; R.L. contributed to studies in primary cells from patients with HIV; A.R. and F.M. generated lentiviral vectors and analysed integration into CD4+ T cells and CD34+ bone marrow cells; M.L. and M.G. conceived and supervised the experiments and wrote the paper with help from the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 HIV-1 RIGs.
a, Probability of recurrence of a random set of genes in different lists of HIV-1 integration sites. The histogram shows the distribution of the number of genes present at least twice in the six HIV-1 integration site lists considered; 1 × 109 independent drawings were evaluated. The distribution peaks around 20 genes, with a maximum observed of 50. The number of RIGs detected experimentally in at least two lists was instead 156 (P < 1 × 10−9). b, Human chromosome map showing the localization of 156 HIV RIGs. Genes found in four, three and two HIV-1 integration lists are highlighted in red, orange and black, respectively. Hotter genomic regions, favoured for HIV-1 integration as described in ref. 1, are highlighted in blue and indicated by a star.
Extended Data Figure 2 Distribution of RIGs or individual integration sites all over the loci analysed by FISH.
The scheme describes the distribution of RIGs (bold) or simple integration sites (regular) around the locus analysed by FISH in Fig. 1; RIGs are in the left panel and hotter zones are in the right panel. As indicated on the side, the total number of RIGs/integrants was calculated within 1, 5 or 10 Mb from the locus analysed by FISH. In total, considering all the RIGs and hotter zones, there are 44 other RIGs/integrants within a window of 1 Mb, 116 within 5 Mb and 169 within 10 Mb around the analysed locus.
Extended Data Figure 3 FISH analysis.
a, Representative images of three-dimensional immuno-DNA FISH of HIV-1 DNA (green) in human primary HIV-1 macrophages stained for mAb414 (red), with relative distribution of FISH signals according to the three concentric zones. b, FISH of HIV-1 DNA (green) in the HIV-1 infected U937 monocytic cell line. c, Representative images of three-dimensional immuno-DNA FISH of lentiviral vector pLV-THM (green) in Jurkat cells. d, Representative images of three-dimensional immuno-DNA FISH of the promoter-less lentiviral vector pCCL–18GFP (green) in Jurkat cells. e, Representative images of three-dimensional immuno-DNA FISH of a gammaretroviral vector (green) in Jurkat cells. For all panels, the graphs are organized as described in the main text.
Extended Data Figure 4 Reconstitution of Nup153 by transfection of an siRNA-resistant plasmid coding for eGFP–Nup153.
a, Scheme of the experiment performed in Jurkat cells. eGFP–Nup153b, Western blot showing Nup153 protein level at the moment of infection. siNT, not targeting siRNA. c, Real-time Alu-PCR in Jurkat cells 2 days after infection with HIV-1NL4.3. Values are mean and s.e.m. of three experiments after normalization over Jurkat transfected with a control, non-targeting siRNA (siNT). d, Representative images of three-dimensional immuno-DNA FISH of HIV-1 DNA (red) in Jurkat cells transfected first with the eGFP–Nup153
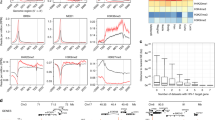
Extended Data Figure 5 ChIP-seq profiles for HIV RIGs, cold genes and controls.
a–e, Profiles of chromatin modifications around the TSS for HIV RIGs (red) and cold genes (green) compared with highly active (black) and silent (blue) genes in activated CD4+ T cells. Each panel reports results for a specific modification, as indicated.
Extended Data Figure 6 Association of HIV-1 provirus with nucleoporins.
a, Positions of primers used for ChIP on the HIV-LTR (numbering is according to the TSS and nucleosomes are shown), the NPLOC4 RIG, the PTPRD cold gene, B48 and B13 genomic controls for DNA standardization. B48 maps within the human lamin B2 origin of DNA replication. b, Control ChIP data in CD4+ T cells infected with HIV-1NL4-3/E-R-, using total immunoglobulin-γ and an antibody against the unrelated Mcm2 cellular protein. For each analysed region, the amount of immunoprecipitated chromatin using the indicated antibodies was normalized according to the input amount of chromatin. Mean and s.e.m. from at least three independent experiments. P < 0.01. c, ChIP results in CD4+ T cells, 4 days after HIV-1 infection, using the indicated antibodies. The amount of immunoprecipitated chromatin was normalized according to input. Mean and s.e.m. from at least three independent experiments. P < 0.01,P < 0.05. d, ChIP results in CD4+ T cells, 4 days after infection with wild-type HIV-1 or the IN(D64E) mutant virus. For the PPR1 region, corresponding to the viral promoter, the amount of immunoprecipitated chromatin using the indicated antibodies (mAb414, Nup153 and Pol2) was calculated according to the input amount of chromatin, and then normalized over the B13 control genomic region. The graphs show the mean and s.e.m. from three independent experiments. P < 0.001; P < 0.05.
Extended Data Figure 7 HIV-1 transcriptional activation is concomitant with, and requires, nucleoporins binding to the provirus.
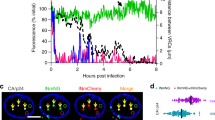
a, Quantitative reverse transcription PCR measurement of HIV-1 mRNA in mock- or TPA-treated J-Lat 15.4 cells. b, Three-dimensional immuno-DNA FISH of HIV-1 DNA (green) in J-Lat 15.4 cells stained for NPC (red) before and after TPA reactivation. c, Scheme of the experiment for the generation of a primary, cellular model of HIV-1 latency to study HIV-1 DNA localization in activated and resting primary CD4+ T cells. d, Quantitative reverse transcription PCR measurement of HIV-1 mRNA levels in primary infected CD4+ T cells before and after reactivation, normalized over the 18S housekeeping gene. Latent versus reactivated: P < 0.001. e, Three-dimensional immuno-DNA FISH of HIV-1 DNA (green) in latently infected CD4+ T cells stained for the NPC (red) before and after reactivation, with relative distribution of HIV-1 FISH signals according to the three concentric zones considered in this work. e, ChIP in control and TPA-stimulated J-Lat 15.4 cells, with the indicated antibodies. Mean and s.e.m. from at least three independent experiments. g, Immunoblot for Tpr (upper panel) and Nup153 (lower panel), 36 h after transfection of the indicated siRNAs (NT, non-targeting control). h, Levels of HIV-1 RNA in siRNA-treated J-Lat 15.4 cells after TPA activation. Mean and s.e.m. from three independent experiments. P < 0.05.
Extended Data Figure 8 Silencing of Tpr in HeLa cells and 15.4 J-Lat clones.
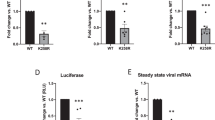
a, Scheme of the experiment to study HIV-1 integration in infected HeLa cells after Tpr silencing. b, Western blot showing Tpr protein level at the moment of infection, after treatment of HeLa cells with a non-targeting siRNA (siNT) or an siRNA targeting Tpr at two different doses. Values are mean and s.e.m. of three experiments after normalization over HeLa cells transfected with a control non-targeting siRNA. c, Real-time Alu PCR in HeLa cells infected with HIV-1NL4.3 and previously transfected with a non-targeting siRNA (siNT) or an siRNA targeting Tpr at two different doses. Values are mean and s.e.m. of three experiments after normalization over HeLa cells transfected with a control non-targeting siRNA. d, Luciferase activity assay in HeLa infected with HIV-1NL4.3 and previously transfected with a non-targeting siRNA (siNT) or an siRNA targeting Tpr at two different doses. Values are mean and s.e.m. of three experiments. Statistical significance: P < 0.001; P < 0.01. e, Real-time PCR quantification of IL-2 mRNA levels in J-Lat 15.4 cells. The following conditions were tested: untreated cells, plus TPA (4 h), transfection with non-targeting siRNA or an siRNA targeting Tpr for 24 h, followed by treatment with TPA (4 h). Values are mean and s.e.m. of three experiments after normalization over GAPDH. Transcription of interleukin-2 (IL-2) was not significantly altered upon Tpr downregulation.
Extended Data Figure 9 Model for HIV-1 integration site selection.
After entry into the nucleus through the nuclear pore, the viral DNA integrates into the active chromatin closest to the NPC (green zones), avoiding both LADs and the inner part of the nucleus (red zones).
Supplementary information
Supplementary Information
This file contains a list of HIV-1 recurrent integration genes (RIGs), hotter zones (HZs) and Controls. (PDF 99 kb)
Rights and permissions
About this article
Cite this article
Marini, B., Kertesz-Farkas, A., Ali, H. et al. Nuclear architecture dictates HIV-1 integration site selection. Nature 521, 227–231 (2015). https://doi.org/10.1038/nature14226
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14226
This article is cited by
-
Antiretroviral APOBEC3 cytidine deaminases alter HIV-1 provirus integration site profiles
Nature Communications (2023)
-
Longitudinal clonal tracking in humanized mice reveals sustained polyclonal repopulation of gene-modified human-HSPC despite vector integration bias
Stem Cell Research & Therapy (2021)
-
Transplantation of human embryonic stem cells alleviates motor dysfunction in AAV2-Htt171-82Q transfected rat model of Huntington’s disease
Stem Cell Research & Therapy (2021)
-
Functional mechanisms and abnormalities of the nuclear lamina
Nature Cell Biology (2021)
-
Light and shadow on the mechanisms of integration site selection in yeast Ty retrotransposon families
Current Genetics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.