Abstract
Obesity is an increasingly prevalent disease regulated by genetic and environmental factors. Emerging studies indicate that immune cells, including monocytes, granulocytes and lymphocytes, regulate metabolic homeostasis and are dysregulated in obesity1,2. Group 2 innate lymphoid cells (ILC2s) can regulate adaptive immunity3,4 and eosinophil and alternatively activated macrophage responses5, and were recently identified in murine white adipose tissue (WAT)5 where they may act to limit the development of obesity6. However, ILC2s have not been identified in human adipose tissue, and the mechanisms by which ILC2s regulate metabolic homeostasis remain unknown. Here we identify ILC2s in human WAT and demonstrate that decreased ILC2 responses in WAT are a conserved characteristic of obesity in humans and mice. Interleukin (IL)-33 was found to be critical for the maintenance of ILC2s in WAT and in limiting adiposity in mice by increasing caloric expenditure. This was associated with recruitment of uncoupling protein 1 (UCP1)+ beige adipocytes in WAT, a process known as beiging or browning that regulates caloric expenditure7,8,9. IL-33-induced beiging was dependent on ILC2s, and IL-33 treatment or transfer of IL-33-elicited ILC2s was sufficient to drive beiging independently of the adaptive immune system, eosinophils or IL-4 receptor signalling. We found that ILC2s produce methionine-enkephalin peptides that can act directly on adipocytes to upregulate Ucp1 expression in vitro and that promote beiging in vivo. Collectively, these studies indicate that, in addition to responding to infection or tissue damage, ILC2s can regulate adipose function and metabolic homeostasis in part via production of enkephalin peptides that elicit beiging.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lumeng, C. N. & Saltiel, A. R. Inflammatory links between obesity and metabolic disease. J. Clin. Invest. 121, 2111–2117 (2011)
Osborn, O. & Olefsky, J. M. The cellular and signaling networks linking the immune system and metabolism in disease. Nature Med. 18, 363–374 (2012)
Moro, K. et al. Innate production of TH2 cytokines by adipose tissue-associated c-Kit+Sca-1+ lymphoid cells. Nature 463, 540–544 (2010)
Halim, T. Y. et al. Group 2 innate lymphoid cells are critical for the initiation of adaptive T helper 2 cell-mediated allergic lung inflammation. Immunity 40, 425–435 (2014)
Molofsky, A. B. et al. Innate lymphoid type 2 cells sustain visceral adipose tissue eosinophils and alternatively activated macrophages. J. Exp. Med. 210, 535–549 (2013)
Hams, E., Locksley, R. M., McKenzie, A. N. & Fallon, P. G. Cutting edge: IL-25 elicits innate lymphoid type 2 and type II NKT cells that regulate obesity in mice. J. Immunol. 191, 5349–5353 (2013)
Harms, M. & Seale, P. Brown and beige fat: development, function and therapeutic potential. Nature Med. 19, 1252–1263 (2013)
Shabalina, I. G. et al. UCP1 in brite/beige adipose tissue mitochondria is functionally thermogenic. Cell Rep. 5, 1196–1203 (2013)
Cohen, P. et al. Ablation of PRDM16 and beige adipose causes metabolic dysfunction and a subcutaneous to visceral fat switch. Cell 156, 304–316 (2014)
Price, A. E. et al. Systemically dispersed innate IL-13-expressing cells in type 2 immunity. Proc. Natl Acad. Sci. USA 107, 11489–11494 (2010)
Neill, D. R. et al. Nuocytes represent a new innate effector leukocyte that mediates type-2 immunity. Nature 464, 1367–1370 (2010)
Miller, A. M. et al. Interleukin-33 induces protective effects in adipose tissue inflammation during obesity in mice. Circ. Res. 107, 650–658 (2010)
Mjösberg, J. M. et al. Human IL-25- and IL-33-responsive type 2 innate lymphoid cells are defined by expression of CRTH2 and CD161. Nature Immunol. 12, 1055–1062 (2011)
Monticelli, L. A. et al. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nature Immunol. 12, 1045–1054 (2011)
Yang, Q. et al. T cell factor 1 is required for group 2 innate lymphoid cell generation. Immunity 38, 694–704 (2013)
Miller, A. M. et al. IL-33 reduces the development of atherosclerosis. J. Exp. Med. 205, 339–346 (2008)
Wu, J. et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 150, 366–376 (2012)
Rosen, E. D. & Spiegelman, B. M. What we talk about when we talk about fat. Cell 156, 20–44 (2014)
Feldmann, H. M., Golozoubova, V., Cannon, B. & Nedergaard, J. UCP1 ablation induces obesity and abolishes diet-induced thermogenesis in mice exempt from thermal stress by living at thermoneutrality. Cell Metab. 9, 203–209 (2009)
Carey, A. L. et al. Ephedrine activates brown adipose tissue in lean but not obese humans. Diabetologia 56, 147–155 (2013)
Saito, M. et al. High incidence of metabolically active brown adipose tissue in healthy adult humans: effects of cold exposure and adiposity. Diabetes 58, 1526–1531 (2009)
Qiu, Y. et al. Eosinophils and type 2 cytokine signaling in macrophages orchestrate development of functional beige fat. Cell 157, 1292–1308 (2014)
Wu, D. et al. Eosinophils sustain adipose alternatively activated macrophages associated with glucose homeostasis. Science 332, 243–247 (2011)
Liu, P.-S. et al. Reducing RIP140 expression in macrophage alters ATM infiltration, facilitates white adipose tissue browning, and prevents high-fat diet-induced insulin resistance. Diabetes 63, 4021–4031 (2014)
Feuerer, M. et al. Lean, but not obese, fat is enriched for a unique population of regulatory T cells that affect metabolic parameters. Nature Med. 15, 930–939 (2009)
McCarthy, M. I. Genomics, type 2 diabetes, and obesity. N. Engl. J. Med. 363, 2339–2350 (2010)
Walley, A. J., Asher, J. E. & Froguel, P. The genetic contribution to non-syndromic human obesity. Nature Rev. Genet. 10, 431–442 (2009)
Seidah, N. G., Sadr, M. S., Chretien, M. & Mbikay, M. The multifaceted proprotein convertases: their unique, redundant, complementary, and opposite functions. J. Biol. Chem. 288, 21473–21481 (2013)
Lloyd, D. J., Bohan, S. & Gekakis, N. Obesity, hyperphagia and increased metabolic efficiency in Pc1 mutant mice. Hum. Mol. Genet. 15, 1884–1893 (2006)
Sharp, L. Z. et al. Human BAT possesses molecular signatures that resemble beige/brite cells. PLoS ONE 7, e49452 (2012)
Brestoff, J. R. & Van den Broeck, J. in Epidemiology: Principles and Practical Guidelines (eds Van den Broeck, J. & Brestoff, J. R. ) pp. 137–155 (Springer, 2013)
Zagon, I. S., Rahn, K. A., Bonneau, R. H., Turel, A. P. & McLaughlin, P. J. Opioid growth factor suppresses expression of experimental autoimmune encephalomyelitis. Brain Res. 1310, 154–161 (2010)
Matthews, D. R. et al. Homeostasis model assessment: insulin resistance and beta-cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia 28, 412–419 (1985)
Seale, P. et al. Prdm16 determines the thermogenic program of subcutaneous white adipose tissue in mice. J. Clin. Invest. 121, 96–105 (2011)
Acknowledgements
The authors wish to thank members of the Artis laboratory for the critical reading of this manuscript. Research in the Artis laboratory is supported by the National Institutes of Health (AI061570, AI074878, AI095466, AI095608, AI102942, and AI097333 to D.A.), the Burroughs Wellcome Fund Investigator in Pathogenesis of Infectious Disease Award (D.A.) and Crohn’s & Colitis Foundation of America (D.A.). Additional funding was provided by NIH F30-AI112023 (J.R.B.), T32-AI060516 (J.R.B.), T32-AI007532 (L.A.M.), KL2-RR024132 (B.S.K.), DP5OD012116 (G.F.S.), P01AI06697 (D.L.F.), F31AG047003 (J.J.T.) and DP2OD007288 (P.S.) and by the Searle Scholars Award (P.S.). We thank M. A. Lazar for scientific and technical advice, D. E. Smith for providing Il33−/− mice, A. Goldrath for providing Id2−/− chimaeras, and A. Bhandoola for providing Tcf7−/− mice. We also thank the Mouse Phenotyping, Physiology & Metabolism Core at the Diabetes Research Center (DRC) of the Institute for Diabetes, Obesity & Metabolism (IDOM) as well as the Penn Diabetes Endocrine Research Center Grant (P30DK19525). In addition, we thank the Matthew J. Ryan Veterinary Hospital Pathology Laboratory, the Penn Microarray Facility, and the Mucosal Immunology Studies Team (MIST) of the NIH NIAID for shared expertise and resources. The authors would also like to thank the Abramson Cancer Center Flow Cytometry and Cell Sorting Resource Laboratory for technical advice and support. The ACC Flow Cytometry and Cell Sorting Shared Resource is partially supported by NCI Comprehensive Cancer Center Support Grant (no. 2-P30 CA016520). This work was supported by the NIH/NIDDK P30 Center for Molecular Studies in Digestive and Liver Diseases (P30-DK050306), its pilot grant program and scientific core facilities (Molecular Pathology and Imaging, Molecular Biology, Cell Culture and Mouse), as well as the Joint CHOP-Penn Center in Digestive, Liver and Pancreatic Medicine and its pilot grant program. In addition, we would like to acknowledge and thank the New York Organ Donor Network, the Cooperative Human Tissue Network-Eastern Division and especially the donors and their families. We apologize to colleagues whose work we were unable to quote owing to space constraints.
Author information
Authors and Affiliations
Contributions
J.R.B., B.S.K., S.A.S., R.R.S., L.A.M., G.F.S., K.L., P.S. and D.A. designed and performed the research and/or provided advice and technical expertise. J.J.T. and D.L.F. provided human tissues. J.R.B. and D.A. analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Identification of human innate lymphoid cell (ILCs) in WAT and developmental and functional characterization of murine ILCs in WAT.
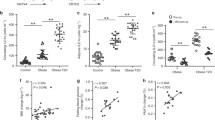
a, Gating strategy to identify human ILCs. Stromal vascular fraction (SVF) cells from human abdominal subcutaneous white adipose tissues (WAT) were isolated and subjected to flow cytometric analyses. First plot pre-gated on singlets. Lineage cocktail 1 (Lin1): CD3, CD5, TCRαβ. Lineage cocktail 2 (Lin2): CD19, CD56, CD11c, CD16. ILCs are identified as Lin-negative cells that are CD25+ CD127+. Plots shown are from an obese donor. b–e, SVF cells from murine epididymal (E)-WAT were isolated and subjected to flow cytometric analyses. ILCs were defined as live CD45+ Lin− CD25+ CD127+ cells. The lineage (Lin) cocktail included CD3, CD5, CD19, NK1.1, CD11c, CD11b and FcεRIα. Comparison of Lin− CD25+ CD127+ cells in E-WAT of b, Id2+/+ versus Id2−/− bone marrow chimaeras, c, Tcf7+/+ versus Tcf7−/− mice and d, Rag2−/− versus Rag2−/− γc−/− mice. n = 3–8 mice per group from 2 independent experiments. e, E-WAT SVF cells from C57BL/6 mice were treated with PMA (100 ng ml−1) and ionomycin (1 μg ml−1) in the presence of Brefeldin A (10 μg ml−1) for 4 h and stained for ILCs. Live CD45+ Lin− CD25+ CD127+ cells were pre-gated, and IL-5 and IL-13 protein levels were assessed. Plot shown is representative of n = 12 mice from 3 independent experiments. f, Human WAT ILC2 frequencies were compared in the 7 youngest donors (36.0 ± 3.5 years old) versus the 7 oldest donors (55.9 ± 1.9 years old). g, Human WAT ILC2 frequencies in female non-obese donors with body mass index (BMI) < 30.0 kg m−2 versus female obese donors with BMI ≥ 30.0 kg m−2. Student’s t-test. **P < 0.01, ***P < 0.001. Data are shown as mean ± standard error and are representative of 2 independent experiments. Sample sizes are biological replicates.
Extended Data Figure 2 IL-33-deficient mice exhibit dysregulated group 2 innate lymphoid cells (ILC2s) in association with increased adipocyte size and impaired glucose homeostasis.
Il33+/+ (n = 6) or Il33−/− (n = 5) mice were fed a low-fat diet (10% kcal fat) for 12 weeks starting at 7 weeks of age. a, Representative plots and frequencies of live CD45+ Lin− CD25+ IL-33R+ ILC2s in epididymal (E)-WAT (data are from Fig. 2a) and iWAT. Plots pre-gated on CD45+ Lin− cells that lack CD3, CD5, CD19, NK1.1, CD11c, CD11b and FcεRIα. b, Frequencies of IL-5+ IL-13− and IL-5+ IL-13+ ILC2s in E-WAT and iWAT of wild-type and IL-33-deficient mice. E-WAT stromal vascular fraction cells were treated with PMA (100 ng ml−1) and ionomycin (1 μg ml−1) in the presence of brefeldin A (10 μg ml−1) for 4 h before staining for ILC2s and intracellular cytokines. Pre-gated on CD45+ Lin− CD25+ IL33R+ ILC2s. c, Inguinal white adipose tissue (iWAT) sections were haematoxylin and eosin stained and imaged at ×40 magnification. Adipocyte area was calculated from 25–40 adipocytes total from 2–3 images per mouse. d, 16-h fasting blood glucose concentrations. e, 16-h fasting serum insulin concentrations. f, Homeostatic model assessment of insulin resistance (HOMA-IR) index values. g, Glucose tolerance test (GTT) with 2 g per kg body weight glucose following a 16-h fast. h, Insulin tolerance test (ITT) with 0.5 U per kg body weight insulin following a 5-h fast. For panels a–f, groups were compared using Student’s t-test, *P < 0.05, ***P < 0.001. For panels g–h, a two-way ANOVA with repeated measures was performed followed by Tukey post-hoc test. *P < 0.05, **P < 0.01, ***P < 0.001. Data shown are from a single cohort and are representative of 2 independent experiment. Sample sizes are biological replicates.
Extended Data Figure 3 IL-33 increases E-WAT ILC2s and regulatory T cells (Tregs) and abrogates the development of obesity and glucose intolerance in mice fed a high-fat diet (HFD).
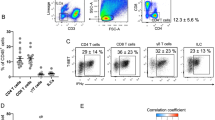
Male C57BL/6 mice were placed on a control diet (CD) or HFD (60% kcal fat) at age 8 weeks. On the first day of feeding, CD mice were treated with PBS and HFD mice were treated with PBS or recombinant murine (rm)IL-33 (12.5 μg per kg body weight) once every 4 days by intraperitoneal injection for 4 weeks. a, E-WAT ILC2 numbers per gram of adipose, b, body weight, c, relative E-WAT weight and d, relative iWAT weight at week 4. e, 16-h fasting blood glucose concentrations and f, glucose tolerance testing during week 3. g, Frequencies and representative plots of E-WAT Tregs defined as live CD45+ CD3+ CD4+ Foxp3+ cells. Plots are gated on live CD45+ CD3+ CD4+ cells, and numbers are the percentage of CD4+ T cells that are Foxp3+ Tregs. h, Numbers of Treg cells per gram of adipose. All panels include n = 10 mice per group from 2 independent cohorts, except panel A which includes n = 16 CD PBS and n = 18 HFD PBS from 4 independent cohorts. a–e, One-way ANOVA with Tukey post-hoc test, *P < 0.05, **P < 0.01, ***P < 0.001. f, Two-way ANOVA with repeated measures, ***P < 0.001 comparing CD PBS versus HFD PBS, ∧∧∧P < 0.001 comparing HFD PBS versus HFD IL-33. Data are shown as mean ± standard error. Sample sizes are biological replicates.
Extended Data Figure 4 Decreased ambulatory activity may limit hyperphagia following IL-33 treatment, but IL-33 does not appear to have direct suppressive effects on food intake or ambulatory activity.
a, Male C57BL/6 mice were treated with PBS or recombinant murine (rm)IL-33 (12.5 μg per kg body weight) daily for 7 days (PBS n = 10, rmIL-33, n = 12). Over a 24 h period between days 6 and 7, food intake and ambulatory activity were measured over 15-min intervals. The average difference in food intake or ambulatory activity between PBS- and rmIL-33-treated mice was calculated for each 15-min interval, and the differences in food intake and ambulatory activity were related by linear regression. Solid line, best-fit line. Dashed curves, upper and lower 95% confidence intervals around the best-fit line. Data are shown as mean differences for each interval and are representative of 2 independent experiments. b–d, Male C57BL/6 mice were treated with PBS or recombinant murine (rm)IL-33 (12.5 μg per kg body weight) once and monitored for the first 3 h post-treatment using CLAMS cages (n = 4 per group). b, Energy expenditure, c, food intake and d, ambulatory activity (beam breaks) were measured over of the first 3 h post-treatment. Student’s t-test. ***P < 0.001. Data are shown as mean ± standard error and are representative of 1 independent experiment. Sample sizes are biological replicates.
Extended Data Figure 5 Brown adipose tissue (BAT) contains Lin− CD25+ IL-33R+ ILC2s that expand in response to IL-33 in association with decreased Ucp1 expression.
C57BL/6 male mice (10 weeks old) were treated with PBS (n = 8) or IL-33 (12.5 μg per kg body weight, n = 8) daily by intraperitoneal injection for 7 days. a, Representative plots and frequencies of Lin− CD25+ IL-33R+ ILC2s in interscapular BAT. Gated on live CD45+ Lin− cells. b, Numbers of ILC2s per gram of BAT. c, Ucp1 expression in BAT by real-time PCR. d, UCP1 immunohistochemistry of BAT at ×10 magnification. Scale bars, 100 μm. e, ×40 magnification of d. Scale bars, 100 μm. Student’s t-test, *P < 0.05, ***P < 0.001. Data are shown as mean ± standard error and are representative of 2 independent experiments. Sample sizes are biological replicates.
Extended Data Figure 6 ILC2s from E-WAT accumulate in white adipose tissue of recipient mice and expand in response to IL-33 to promote beiging.
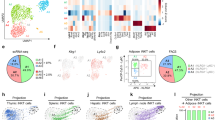
a, Experimental design for panels a, b. Live CD45+ Lin− CD25+ IL-33R+ ILC2s were sort-purified from E-WAT of CD45.1+ mice treated with 12.5 μg per kg body weight recombinant murine (rm)IL-33 daily for 7 days by intraperitoneal injection. PBS (n = 8) or ILC2s (1 × 105 total, n = 8) were transferred to CD45.2+ recipient mice daily for 4 days by subcutaneous injection near iWAT (5 × 104 ILC2s, split evenly bilaterally) and intraperitoneal injection (5 × 104 ILC2s). Tissues were harvested on day 5 for analyses. b, Donor and recipient ILC2s in E-WAT, brown adipose tissue (BAT), mesenteric lymph nodes (mLN) and lung. iWAT ILC2 plots from this experiment are shown in main Fig. 3g. Pre-gated on Live CD45+ Lin− CD25+ IL-33R+ ILC2s. Donor ILC2s are defined as CD45.1+ CD45.2−, whereas recipient ILC2s are defined as CD45.1− CD45.2+. Representative plots shown. Frequencies represent percent of ILC2s that are recipient or donor cells. Student’s t-test, ***P < 0.001. c, Experimental design for panels c–e. Sort-purified CD45.1+ ILC2s (×105) from E-WAT of IL-33-treated mice (as described above) were transferred into Rag2−/− γc−/− recipients by a single intraperitoneal injection. ILC2-sufficient Rag2−/− mice, ILC2-deficient Rag2−/− γc−/− mice and ILC2-reconstituted Rag2−/− γc−/− mice were treated with PBS or rmIL-33 (12.5 μg per kg body weight) by intraperitoneal injection daily for 7 days. There were n = 4 mice per group. This experimental design corresponds to main Fig. 3l-m. d, Representative plots of live CD45.1+ Lin− CD25+ IL33R+ ILC2s in E-WAT. Blue, recipient cells. Red, donor cells. Lineage cocktail includes CD3, CD5, CD19, NK1.1, CD11c, CD11b and FcεRIα. e, iWAT UCP1 IHC. Scale bars, 100 μm. ANOVA with Tukey post-hoc test, ***P < 0.001. Data are shown as mean ± standard error and are representative of 2 independent experiments. Sample sizes are biological replicates.
Extended Data Figure 7 IL-33 treatment and ILC2 transfer can elicit beiging independently of eosinophils and IL-4Rα signalling.
a–f, Wild-type (Balb/c), DblGata1 mice that lack eosinophils or Il4ra−/− mice that have dysregulated alternatively activated macrophages (AAMacs) (both mutant strains on a Balb/c background) were treated with PBS or recombinant murine (rm)IL-33 (12.5 μg per kg body weight) daily by intraperitoneal injection for 7 days. a, iWAT ILC2 numbers per gram of adipose and b, iWAT UCP1 immunohistochemistry (IHC) in Balb/c mice (PBS, n = 4; rmIL-33, n = 3). c, iWAT ILC2 numbers per gram of adipose and d, iWAT UCP1 IHC in DblGata1 mice (PBS, n = 5; rmIL-33, n = 6). e, iWAT ILC2 numbers per gram of adipose and f, iWAT UCP1 IHC in Il4ra−/− mice (PBS, n = 4; rmIL-33, n = 6). g, h, Live CD45+ Lin− CD25+ IL-33R+ ILC2s were sort-purified from E-WAT of C57BL/6 mice treated with rmIL-33 (12.5 μg per kg body weight) daily for 5–7 days by intraperitoneal injection to Rag1−/− mice on a C57BL/6 background. ILC2s (1 × 105 total) were transferred to recipient mice daily for 4 days by subcutaneous injection (PBS, n = 8; ILC2, n = 8). g, iWAT ILC2 numbers per gram of adipose and h, iWAT UCP1 IHC. i–l, Live CD45+ Lin− CD25+ IL-33R+ ILC2s were sort-purified from E-WAT of Balb/c mice treated with rmIL-33 (12.5 μg per kg body weight) daily for 5–7 days by intraperitoneal injection. ILC2s (1 × 105 total) were transferred to recipient mice daily for 4 days by subcutaneous injection. i, iWAT ILC2 numbers per gram of adipose and j, iWAT UCP1 IHC in DblGata1 recipients (PBS, n = 6; ILC2, n = 6). k, iWAT ILC2 numbers per gram of adipose and l, iWAT UCP1 IHC in Il4ra−/− recipients (PBS, n = 3; ILC2, n = 4). Scale bars, 100 μm. Student’s t-test, *P < 0.05. Data are shown as mean ± standard error and are representative of 2 independent experiments. Sample sizes are biological replicates.
Extended Data Figure 8 Summary model linking the IL-33/ILC2/MetEnk pathway to the regulation of beiging and obesity.
Interleukin (IL)-33 acts on group 2 innate lymphoid cells (ILC2s) to upregulate production of the effector molecules IL-5, IL-13 and enkephalin peptides. ILC2-derived IL-5 promotes eosinophil homeostasis in WAT, and eosinophils in turn produce IL-4 to sustain alternatively activated macrophages (AAMacs) in WAT. ILC2-derived IL-13 can also promote AAMac responses. In the setting of chronic exposure to a cold environment, eosinophil-derived IL-4 stimulates AAMacs to produce catecholamines such as noradrenaline, which acts directly on beige adipocytes to upregulate uncoupling protein 1 (UCP1) expression and promote mitochondrial biogenesis. Although it remains unknown whether ILC2-derived IL-5 and IL-13 contribute to cold-stress-induced beiging, ILC2-derived enkephalin peptides can act directly on beige adipocytes to upregulate UCP1 and promote beiging. This results in increased energy expenditure and decreased adiposity that may counteract weight gain. In the setting of obesity, IL-33 expression in WAT is increased; however, WAT ILC2s are paradoxically decreased in both mice and humans, suggesting that the IL-33/ILC2 axis is dysregulated in obesity. This may impede the ability of ILC2s to contribute to the function of beige fat, resulting in a vicious cycle that promotes weight gain.
Rights and permissions
About this article
Cite this article
Brestoff, J., Kim, B., Saenz, S. et al. Group 2 innate lymphoid cells promote beiging of white adipose tissue and limit obesity. Nature 519, 242–246 (2015). https://doi.org/10.1038/nature14115
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14115
This article is cited by
-
Telmisartan and candesartan promote browning of white adipose tissue and reverse fatty liver changes in high fat diet fed male albino rats
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Characteristic and fate determination of adipose precursors during adipose tissue remodeling
Cell Regeneration (2023)
-
Role of thymic stromal lymphopoietin in allergy and beyond
Nature Reviews Immunology (2023)
-
Age-dependent Pdgfrβ signaling drives adipocyte progenitor dysfunction to alter the beige adipogenic niche in male mice
Nature Communications (2023)
-
Regulation and function of adiponectin in the intestinal epithelial cells in response to Trichinella spiralis infection
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.