Abstract
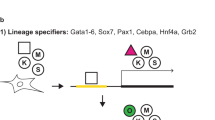
Pluripotency is defined by the ability of a cell to differentiate to the derivatives of all the three embryonic germ layers: ectoderm, mesoderm and endoderm. Pluripotent cells can be captured via the archetypal derivation of embryonic stem cells or via somatic cell reprogramming. Somatic cells are induced to acquire a pluripotent stem cell (iPSC) state through the forced expression of key transcription factors, and in the mouse these cells can fulfil the strictest of all developmental assays for pluripotent cells by generating completely iPSC-derived embryos and mice. However, it is not known whether there are additional classes of pluripotent cells, or what the spectrum of reprogrammed phenotypes encompasses. Here we explore alternative outcomes of somatic reprogramming by fully characterizing reprogrammed cells independent of preconceived definitions of iPSC states. We demonstrate that by maintaining elevated reprogramming factor expression levels, mouse embryonic fibroblasts go through unique epigenetic modifications to arrive at a stable, Nanog-positive, alternative pluripotent state. In doing so, we prove that the pluripotent spectrum can encompass multiple, unique cell states.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
Microarray data have been deposited on Stemformatics (http://www.stemformatics.org) and in the Gene Expression Omnibus database under accession number GSE49940.
Change history
10 December 2014
A minor addition was made to the Acknowledgements in the HTML and PDF versions.
References
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Nagy, A., Rossant, J., Nagy, R., Abramow-Newerly, W. & Roder, J. C. Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc. Natl Acad. Sci. USA 90, 8424–8428 (1993)
Zhao, X.-Y. et al. iPS cells produce viable mice through tetraploid complementation. Nature 461, 86–90 (2009)
Fussner, E. et al. Constitutive heterochromatin reorganization during somatic cell reprogramming. EMBO J. 30, 1778–1789 (2011)
Sridharan, R. et al. Role of the murine reprogramming factors in the induction of pluripotency. Cell 136, 364–377 (2009)
Chen, J. et al. H3K9 methylation is a barrier during somatic cell reprogramming into iPSCs. Nature Genet. 45, 34–42 (2013)
Kim, K. et al. Epigenetic memory in induced pluripotent stem cells. Nature 467, 285–290 (2010)
Polo, J. M. et al. Cell type of origin influences the molecular and functional properties of mouse induced pluripotent stem cells. Nature Biotechnol. 28, 848–855 (2010)
Hu, X. et al. Tet and TDG mediate DNA demethylation essential for mesenchymal-to-epithelial transition in somatic cell reprogramming. Cell Stem Cell 14, 512–522 (2014)
Mali, P. et al. Butyrate greatly enhances derivation of human induced pluripotent stem cells by promoting epigenetic remodeling and the expression of pluripotency-associated genes. Stem Cells 28, 713–720 (2010)
Wang, T. et al. The histone demethylases Jhdm1a/1b enhance somatic cell reprogramming in a vitamin-C-dependent manner. Cell Stem Cell 9, 575–587 (2011)
Mansour, A. A. et al. The H3K27 demethylase Utx regulates somatic and germ cell epigenetic reprogramming. Nature 488, 409–413 (2012)
Esteban, M. A. et al. Vitamin C enhances the generation of mouse and human induced pluripotent stem cells. Cell Stem Cell 6, 71–79 (2009)
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nature Genet. 24, 372–376 (2000)
Golipour, A. et al. A late transition in somatic cell reprogramming requires regulators distinct from the pluripotency network. Cell Stem Cell 11, 769–782 (2012)
Samavarchi-Tehrani, P. et al. Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell 7, 64–77 (2010)
Polo, J. M. et al. A molecular roadmap of reprogramming somatic cells into iPS cells. Cell 151, 1617–1632 (2012)
Woltjen, K. et al. piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Nature 458, 766–770 (2009)
Soncin, F. et al. Abrogation of E-cadherin-mediated cell–cell contact in mouse embryonic stem cells results in reversible LIF-independent self-renewal. Stem Cells 27, 2069–2080 (2009)
Soncin, F. et al. E-cadherin acts as a regulator of transcripts associated with a wide range of cellular processes in mouse embryonic stem cells. PLoS ONE 6, e21463 (2011)
Larue, L. et al. A role for cadherins in tissue formation. Development 122, 3185–3194 (1996)
Müller, F.-J. et al. Regulatory networks define phenotypic classes of human stem cell lines. Nature 455, 401–405 (2008)
Wernig, M. et al. A drug-inducible transgenic system for direct reprogramming of multiple somatic cell types. Nature Biotechnol. 26, 916–924 (2008)
Okita, K., Ichisaka, T. & Yamanaka, S. Generation of germline-competent induced pluripotent stem cells. Nature 448, 313–317 (2007)
Hussein, S. M. I. et al. Genome-wide characterization of the routes to induced pluripotency. Nature http://dx.doi.org/10.1038/nature14046 (2014)
Benevento, M. et al. Proteome adaptation in cell reprogramming proceeds via distinct transcriptional networks. Nature Commun. http://dx.doi.org/10.1038/ncomms6613 (2014)
Lee, D. S. et al. An epigenomic roadmap to induced pluripotency reveals DNA methylation as a reprogramming modulator. Nature Commun. http://dx.doi.org/10.1038/ncomms6619 (2014)
Clancy, J. L. et al. Small RNA changes en route to distinct cellular states of induced pluripotency. Nature Commun. http://dx.doi.org/10.1038/ncomms6522 (2014)
Li, R. et al. A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 7, 51–63 (2010)
Chan, E. M. et al. Live cell imaging distinguishes bona fide human iPS cells from partially reprogrammed cells. Nature Biotechnol. 27, 1033–1037 (2009)
Belteki, G. et al. Conditional and inducible transgene expression in mice through the combinatorial use of Cre-mediated recombination and tetracycline induction. Nucleic Acids Res. 33, e51 (2005)
Nagy, A., Gertsenstein, M., Vintersten, K. & Behringer, R. R. Manipulating the Mouse Embryo: A Laboratory Manual 3rd edn (Cold Spring Harbor Laboratory Press, 2003)
Gertsenstein, M. et al. Efficient generation of germ line transmitting chimeras from C57BL/6N ES cells by aggregation with outbred host embryos. PLoS ONE 5, e11260 (2010)
Hussein, S. M. et al. Copy number variation and selection during reprogramming to pluripotency. Nature 471, 58–62 (2011)
Imamura, M. et al. Transcriptional repression and DNA hypermethylation of a small set of ES cell marker genes in male germline stem cells. BMC Dev. Biol. 6, 34 (2006)
Han, D. W. et al. Epiblast stem cell subpopulations represent mouse embryos of distinct pregastrulation stages. Cell 143, 617–627 (2010)
Acknowledgements
We are grateful for A. Bang’s expertise and assistance regarding flow cytometry. We thank M. Gertsenstein and M. Pereira for chimaera production, and K. Harpal for teratoma sectioning. We would also like to acknowledge the assistance and support of lab colleagues, collaborators and all the members of the Project Grandiose Consortium who are too numerous to name individually but who made a positive impact on this research. A.N. is Tier 1 Canada Research Chair in Stem Cells and Regeneration. This work was supported by grants awarded to A.N. and I.M.R. from the Ontario Research Fund Global Leadership Round in Genomics and Life Sciences grants (GL2), to A.N. from the Canadian stem cell network (9/5254 (TR3)) and Canadian Institutes of Health Research (CIHR MOP102575), to J.-S.S. by the South Korean Ministry of Knowledge Economy (no. 10037410), SNUCM research fund (grant no. 0411-20100074), and Macrogen Inc. (no. MGR03-11 and 12), to S.G. from the Australian Research Council (no. SR110001002), and to C.A.W. by a Queensland government Smart Futures Fellowship and an ARC by Stem Cells Australia and to T.P. grants from NHMRC and ARC. S.M.I.H. received a fellowship from the McEwen Centre of Regenerative Medicine.
Author information
Authors and Affiliations
Contributions
P.D.T. and A.N. conceived and designed the experiments, and wrote the manuscript. P.D.T. and A.J.C. derived all iPSC lines, performed real-time PCR analysis and bisulphite sequencing analysis of DNA methylation. P.D.T., C.M. and I.P.M. performed the in vivo characterization of the iPS lines (teratomas and chimaera formation). O.K., J.L.C. and T.P. assisted in data analysis. P.D.T., J.C.M. and C.A.W. performed microarray analysis. M.L., M.C.P., S.M.I.H. and I.M.R. performed pull-downs for genome-wide ChiP-seq. D.-S.L., J.-Y.S., and J.-S.S. performed genome-wide MethylC-seq and ChiP-seq. N.C., D.L.W., M.E.G. and S.M.G. performed RNA-seq.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Expression profile of F-class cells.
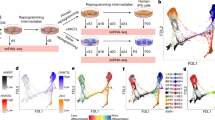
a, Quantitative RT–PCR analysis of total reprogramming factor expression in day 16 F-class (n = 6) and C-class (n = 22), non-parametric t-test. b, Differentially expressed genes (two-tailed Welch t-test P < 0.01, FDR < 0.01) between transgene-expressing reprogrammed lines (n = 28) and ESC-like lines (n = 3). c, Genes highly expressed in b compared against parental fibroblasts. Genes >twofold higher than fibroblasts classified as reprogramming induced. d, Scatter plot of differentially expressed genes (Welch’s t-test P < 0.01; FDR < 0.05). e, Quantitative RT–PCR profiling of cells in a. Non-parametric t-test between the F- and C-class lines (n = 28) ; *P < 0.05, **P < 0.01, ***P < 0.001. f, Expression of PluriNet genes were compared between ESC-like state and F-class state (P values < 0.05, adjusted using the Benjamini–Hochberg method). GeneMANIA interaction network of known gene co-expression and physical interactions. Black nodes represent input genelist, grey nodes represent connecting genes, red nodes represent non-PluriNet genes identified by GeneMANIA that are downregulated in F-class cells.
Extended Data Figure 2 Comparison to epiblast stem cells.
a, Flow cytometric analysis of Nanog expression in F- and C-class primary cell lines after 21 days of transgene expression. Graphs show one of n = 2 experiments. b, Immunofluorescent staining of F-class cells (clone 2) after 30 days of transgene expression. Blue represents Hoechst DNA stain. Scale bars, 100 μm. c, Unsupervised hierarchical clustering of gene expression. EpiSC and ESC populations from ref. 36, with all other cell lines described in Fig. 1b. d, Quantitative RT–PCR analysis of F-class cells (day 30) grown in EpiSC media for 7 days. Graphs show one of n = 2 biological replicates, with 3 technical replicates each. e, Proliferation of established F-class cells (day 30) plated in different media compositions, 1,000 cells plated per cm2, n = 3 technical replicates from one experiment.
Extended Data Figure 3 A stable stem-cell state.
a, Schematic of Cdh1 overexpressing sleeping beauty transposon. IR depicts sleeping beauty inverted repeats. Quantitative RT–PCR of gene expression after 7 days Cdh1 overexpression. n = 3 technical replicates from one experiment. b, Images of Cdh1 overexpressing F-class cells. Scale bars, 100 μm. c, Quantitative RT–PCR analysis of 12 sub-lines derived from clone 1 F-class cells. Average ± s.d. d, DPPA4 immunofluorescence of clone 1 (sub-line-1) after 30 days of transgene expression. Scale bars, 200 μm. e, G-banded karyotype on diploid metaphases of F-class clones. f, Ability to maintain a reprogrammed state in the absence of transgene expression, doxycycline removed after day 21, n = 3 technical replicates from one experiment. Clonal lines ordered as in Fig. 1b.
Extended Data Figure 4 F-class expansion in absence of LIF signalling.
a, Nanog immunofluorescence of single-cell-derived colonies (Day 5). b, Clonal efficiency of F-class cells and ESCs treated with JAK inhibitor (data shown is the mean from n = 3 biological replicates, with 3 technical replicates each, average ± s.d.). c, 1B secondary fibroblast reprogramming25 initiated by doxycycline treatment of fibroblasts in either JAKi-supplemented media (no LIF) or LIF-supplemented media (standard serum-based ESC media). Scale bars, 200 μm. d, Cell expansion of c during 10 days of reprogramming (data shown is the mean from 3 technical replicates from one experiment). e, f, Gene expression analysis (qRT–PCR) of Day 16 reprogramming in JAKi and LIF media (c). Assessment of F-class markers (e) and ESC markers (f) (data shown is the mean from n = 2 biological replicates with 3 technical replicates each). g, DsRed ESCs were mixed with GFP F-class cells. Flow cytometric analysis of population composition before and after passaging. h, Proliferation of F-class and ESC cells grown as suspension culture. i, Phase contrast image of cells grown in suspension for 9 days. Scale bars, 200 μm.
Extended Data Figure 5 In vitro differentiation to three germ layers.
a, TUJ1-positive neurons generated by F-class cells upon doxycycline withdrawal in serum-free media (day 30). b, Multiple neuronal subtypes generated by F-class cells, (clone 1, sub-line 1). c, Quantitative RT–PCR analysis of gene expression during neural differentiation. Clone 1 F-class cells (black line) in comparison to ESC differentiation (grey line). 3 biological replicates (average ± s.d.). d, Doxycycline withdrawal induced differentiation of day 35 F-class cells in 15% serum-based media for 8 days. Immunofluorescent staining of cells representing endoderm (FoxA2) and mesoderm (α-SMA). Scale bars, 200 μm.
Extended Data Figure 6 Transgene expression levels direct reprogramming.
a, Schematic representation of three assessed reprogramming systems. b, Quantitative RT–PCR of factor expression of clonal lines, day 25–day 35. Each point represents a clonal reprogramming colony with n = 6 biological replicates and 3 technical replicates each. c, Representative images of transgene-expressing cells. The day 14 images are representative of 3F and low-expressing 4F reprogramming to highlight the appearance of ESC-like colonies. Scale bars, 200 μm. d, Nanog expression in day 30 colonies. High-expressing 4F cells (1B) exhibit the F-class cell morphology. Scale bars, 100 μm. e, Principal component analysis of quantitative RT–PCR values (32 genes). 3F, 4F low and 4F high cell lines are described in a and b. Cell-state landmarks are F-class clones 1 and 5 (red squares), and C-class clones 5, 10 and 23 (blue squares). f, Quantitative RT–PCR analysis of low-expressing 4F (Col1a1, grey line) and high-expressing 4F (2° 1B, black line) reprogramming, n = 1. g, Schematic model of proposed cell reprogramming routes.
Extended Data Figure 7 Adult tail tip derived F-class cells.
a, Tail-tip fibroblast-derived F-class cells. Scale bars, 200 μm. b, Quantitative RT–PCR analysis of gene expression (day 25 of transgene expression) in clonal tail tip fibroblast reprogrammed cell lines n = 7 biological replicates. c, Principal component analysis of gene expression profile (quantitative RT–PCR, 32 genes). d, Retroviral silencing during transposon mediated reprogramming to F-class state. Quantitative PCR analysis of retroviral copy number (genomic DNA levels) and RNA transcription (n = 3 technical replicates from one experiment). e, Retroviral silencing in established F-class cells (n = 3 technical replicates from one experiment).
Extended Data Figure 8 Requirement of four reprogramming factors.
a, Phase contrast images of F-class cells (CAG-3F + tetO Myc cells). Scale bars, 200 μm. b, Quantitative RT–PCR analysis of reprogramming factor expression, two independent cell lines (data are from n = 2 biological replicates with 3 technical replicates each). c, Genes exhibiting >twofold change upon doxycycline removal (Illumina BeadArray, two independent clones). d, Gene ontology term enrichment of differential gene expression. e, Reprogramming factor expression was activated in ESC-like cells (1B primary iPS cell line). f, Quantitative RT–PCR of reprogramming factor expression in cell lines (n = 10) established from F-class colonies picked in e. g, Quantitative RT–PCR expression of ESC and F-class gene identifiers in cell lines (n = 10) established from F-class colonies picked in e.
Extended Data Figure 9 HDACi-induced transition to ESC-like state.
a, Quantitative RT–PCR of gene expression in F-class cells (day 30, doxycycline-supplemented) that were either maintained in 2i media or exposed to HDAC inhibitors for 10 days. Two cell lines representative of six F-class lines. Data are from n = 3 technical replicates from one experiment. b, Quantitative RT–PCR of F-class (clone 1) sub-lines (n = 12) treated with 10 nM trichostatin A for 6 days. Line denotes average. c, Cell division rate of HDACi-treated cells, as determined directly by time-lapse analysis. d, Flow cytometric analysis of cell viability upon HDACi treatment (10 nM trichostatin A).
Extended Data Figure 10 Temporal effect of HDACi.
a, Schematic representation of HDACi treatment. b, Quantitative RT–PCR analysis of gene expression during HDACi treatment. Red bars depict time points of HDACi exposure (10 nM TSA). Data are from n = 3 biological replicates with 3 technical replicates each (average ± s.d.). c, Principal component analysis of gene expression (Illumina BeadArray). d, Gene ontology term enrichment analysis of genes during HDACi treatment. e, Unsupervised hierarchical clustering of gene expression (Illumina BeadArray) corresponding to primary reprogrammed clones after 16 days of transgene expression and day 16 cells from the 1B secondary reprogramming system (1BD16).
Supplementary information
Supplementary Information
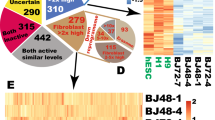
This file contains Supplementary Information sets 1-6. Supplementary Information 1 contains differentially expressed genes (twotailed Welch t-test p<0.01, FDR<0.01) between 28 transgene-expressing reprogrammed lines and 3 ESC-like lines. Supplementary Information 2 contains differentially expressed genes (Welchs t-test p<0.01 FDR<0.05) between F-class and C-class cells. Supplementary Information 3 contains plurinet genes of statistically significant differential expression between F-class samples and ESC samples. Supplementary Information 4 contains differential gene expression upon inactivation of c-Myc expression in F-class cells. Supplementary Information 5 contains differential gene expression upon treatment of F-class cells with HDAC inhibitors. Supplementary Information 6 contains epigenetic status of loci that exhibited differential gene expression (>5 fold) between F-class state and ESC-like cell state. The status of three epigenetic marks are examined; H3K4 trimethylation (H3K4me3), H3K27 trimethylation (H3K27me3) and CpG methylation. Supplementary Information 7 contains primer sequences for quantitative RT-PCR analysis of gene expression. (XLSX 8992 kb)
Rights and permissions
About this article
Cite this article
Tonge, P., Corso, A., Monetti, C. et al. Divergent reprogramming routes lead to alternative stem-cell states. Nature 516, 192–197 (2014). https://doi.org/10.1038/nature14047
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14047
This article is cited by
-
Cell-type dependent regulation of pluripotency and chromatin remodeling genes by hydralazine
Stem Cell Research & Therapy (2023)
-
Determining epigenetic memory in kidney proximal tubule cell derived induced pluripotent stem cells using a quadruple transgenic reprogrammable mouse
Scientific Reports (2022)
-
Imprinting fidelity in mouse iPSCs depends on sex of donor cell and medium formulation
Nature Communications (2022)
-
Here and there a trophoblast, a transcriptional evaluation of trophoblast cell models
Cellular and Molecular Life Sciences (2022)
-
Porcine Primordial Germ Cell-Like Cells Generated from Induced Pluripotent Stem Cells Under Different Culture Conditions
Stem Cell Reviews and Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.