Abstract
Resveratrol is reported to extend lifespan1,2 and provide cardio-neuro-protective3, anti-diabetic4, and anti-cancer effects3,5 by initiating a stress response2 that induces survival genes. Because human tyrosyl transfer-RNA (tRNA) synthetase (TyrRS) translocates to the nucleus under stress conditions6, we considered the possibility that the tyrosine-like phenolic ring of resveratrol might fit into the active site pocket to effect a nuclear role. Here we present a 2.1 Å co-crystal structure of resveratrol bound to the active site of TyrRS. Resveratrol nullifies the catalytic activity and redirects TyrRS to a nuclear function, stimulating NAD+-dependent auto-poly-ADP-ribosylation of poly(ADP-ribose) polymerase 1 (PARP1). Downstream activation of key stress signalling pathways are causally connected to TyrRS–PARP1–NAD+ collaboration. This collaboration is also demonstrated in the mouse, and is specifically blocked in vivo by a resveratrol-displacing tyrosyl adenylate analogue. In contrast to functionally diverse tRNA synthetase catalytic nulls created by alternative splicing events that ablate active sites7, here a non-spliced TyrRS catalytic null reveals a new PARP1- and NAD+-dependent dimension to the physiological mechanism of resveratrol.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Howitz, K. T. et al. Small molecule activators of sirtuins extend Saccharomyces cerevisiae lifespan. Nature 425, 191–196 (2003)
Viswanathan, M., Kim, S. K., Berdichevsky, A. & Guarente, L. A role for SIR-2.1 regulation of ER stress response genes in determining C. elegans life span. Dev. Cell 9, 605–615 (2005)
Baur, J. A. et al. Resveratrol improves health and survival of mice on a high-calorie diet. Nature 444, 337–342 (2006)
Milne, J. C. et al. Small molecule activators of SIRT1 as therapeutics for the treatment of type 2 diabetes. Nature 450, 712–716 (2007)
Jang, M. et al. Cancer chemopreventive activity of resveratrol, a natural product derived from grapes. Science 275, 218–220 (1997)
Fu, G., Xu, T., Shi, Y., Wei, N. & Yang, X. L. tRNA-controlled nuclear import of a human tRNA synthetase. J. Biol. Chem. 287, 9330–9334 (2012)
Lo, W. S. et al. Human tRNA synthetase catalytic nulls with diverse functions. Science 345, 328–332 (2014)
Wakasugi, K. & Schimmel, P. Two distinct cytokines released from a human aminoacyl-tRNA synthetase. Science 284, 147–151 (1999)
Guo, M. & Schimmel, P. Essential nontranslational functions of tRNA synthetases. Nature Chem. Biol. 9, 145–153 (2013)
Yang, X. L., Skene, R. J., McRee, D. E. & Schimmel, P. Crystal structure of a human aminoacyl-tRNA synthetase cytokine. Proc. Natl Acad. Sci. USA 99, 15369–15374 (2002)
Yang, X. L., Liu, F. M., Skene, R. J., McRee, D. E. & Schimmel, P. Crystal structure of an EMAP-II-like cytokine released from a human tRNA synthetase. Helv. Chim. Acta 86, 1246–1257 (2003)
Sajish, M. et al. Trp-tRNA synthetase bridges DNA-PKcs to PARP-1 to link IFN-γ and p53 signaling. Nature Chem. Biol. 8, 547–554 (2012)
Luo, X. & Kraus, W. L. On PAR with PARP: cellular stress signaling through poly(ADP-ribose) and PARP-1. Genes Dev. 26, 417–432 (2012)
Sinclair, D. A. & Guarente, L. Small-molecule allosteric activators of sirtuins. Annu. Rev. Pharmacol. Toxicol. 54, 363–380 (2014)
Kapoor, M., Otero, F. J., Slike, B. M., Ewalt, K. L. & Yang, X. L. Mutational separation of aminoacylation and cytokine activities of human tyrosyl-tRNA synthetase. Chem. Biol. 16, 531–539 (2009)
Lee, P. S., Zhang, H. M., Marshall, A. G., Yang, X. L. & Schimmel, P. Uncovering of a short internal peptide activates a tRNA synthetase procytokine. J. Biol. Chem. 287, 20504–20508 (2012)
Tang, Y., Zhao, W. H., Chen, Y., Zhao, Y. M. & Gu, W. Acetylation is indispensable for p53 activation. Cell 133, 612–626 (2008)
Bitterman, K. J., Anderson, R. M., Cohen, H. Y., Latorre-Esteves, M. & Sinclair, D. A. Inhibition of silencing and accelerated aging by nicotinamide, a putative negative regulator of yeast sir2 and human SIRT1. J. Biol. Chem. 277, 45099–45107 (2002)
Petesch, S. J. & Lis, J. T. Activator-induced spread of poly(ADP-ribose) polymerase promotes nucleosome loss at Hsp70. Mol. Cell 45, 64–74 (2012)
Cohen-Armon, M. et al. DNA-independent PARP-1 activation by phosphorylated ERK2 increases Elk1 activity: a link to histone acetylation. Mol. Cell 25, 297–308 (2007)
Vyas, S. & Chang, P. Dual roles for PARP1 during heat shock: transcriptional activator and posttranscriptional inhibitor of gene expression. Mol. Cell 49, 1–3 (2013)
Hegan, D. C. et al. Inhibition of poly(ADP-ribose) polymerase down-regulates BRCA1 and RAD51 in a pathway mediated by E2F4 and p130. Proc. Natl Acad. Sci. USA 107, 2201–2206 (2010)
Orlando, G., Khoronenkova, S. V., Dianova, I. I., Parsons, J. L. & Dianov, G. L. ARF induction in response to DNA strand breaks is regulated by PARP1. Nucleic Acids Res. 42, 2320–2329 (2014)
Dasgupta, B. & Milbrandt, J. Resveratrol stimulates AMP kinase activity in neurons. Proc. Natl Acad. Sci. USA 104, 7217–7222 (2007)
Park, S. J. et al. Resveratrol ameliorates aging-related metabolic phenotypes by inhibiting cAMP phosphodiesterases. Cell 148, 421–433 (2012)
Butin-Israeli, V., Drayman, N. & Oppenheim, A. Simian virus 40 infection triggers a balanced network that includes apoptotic, survival, and stress pathways. J. Virol. 84, 3431–3442 (2010)
Bastide, B., Snoeckx, K. & Mounier, Y. ADP-ribose stimulates the calcium release channel RyR1 in skeletal muscle of rat. Biochem. Biophys. Res. Commun. 296, 1267–1271 (2002)
Shin, S. M., Cho, I. J. & Kim, S. G. Resveratrol protects mitochondria against oxidative stress through AMP-activated protein kinase-mediated glycogen synthase kinase-3β inhibition downstream of poly(ADP-ribose)polymerase-LKB1 pathway. Mol. Pharmacol. 76, 884–895 (2009)
Howitz, K. T. & Sinclair, D. A. Xenohormesis: sensing the chemical cues of other species. Cell 133, 387–391 (2008)
Nwachukwu, J. C. et al. Resveratrol modulates the inflammatory response via an estrogen receptor-signal integration network. Elife 3, e02057 (2014)
Wang, J. D. & Chen, J. J. SIRT1 regulates autoacetylation and histone acetyltransferase activity of TIP60. J. Biol. Chem. 285, 11458–11464 (2010)
Sun, Y., Jiang, X., Chen, S., Fernandes, N. & Price, B. D. A role for the Tip60 histone acetyltransferase in the acetylation and activation of ATM. Proc. Natl Acad. Sci. USA 102, 13182–13187 (2005)
Chen, W. M. et al. Direct interaction between Nrf2 and p21(Cip1/WAF1) upregulates the Nrf2-mediated antioxidant response. Mol. Cell 34, 663–673 (2009)
Zhang, P. et al. Tumor suppressor p53 cooperates with SIRT6 to regulate gluconeogenesis by promoting FoxO1 nuclear exclusion. Proc. Natl Acad. Sci. USA 111, 10684–10689 (2014)
Nemoto, S., Fergusson, M. M. & Finkel, T. Nutrient availability regulates SIRT1 through a forkhead-dependent pathway. Science 306, 2105–2108 (2004)
Budanov, A. V. & Karin, M. p53 target genes sestrin1 and sestrin2 connect genotoxic stress and mTOR signaling. Cell 134, 451–460 (2008)
Renault, V. M. et al. The pro-longevity gene FoxO3 is a direct target of the p53 tumor suppressor. Oncogene 30, 3207–3221 (2011)
Mao, Z. et al. SIRT6 promotes DNA repair under stress by activating PARP1. Science 332, 1443–1446 (2011)
Rajamohan, S. B. et al. SIRT1 promotes cell survival under stress by deacetylation-dependent deactivation of poly(ADP-ribose) polymerase 1. Mol. Cell. Biol. 29, 4116–4129 (2009)
Smith, B. C., Hallows, W. C. & Denu, J. M. A continuous microplate assay for sirtuins and nicotinamide-producing enzymes. Anal. Biochem. 394, 101–109 (2009)
Perraud, A. L. et al. ADP-ribose gating of the calcium-permeable LTRPC2 channel revealed by Nudix motif homology. Nature 411, 595–599 (2001)
Formentini, L. et al. Poly(ADP-ribose) catabolism triggers AMP-dependent mitochondrial energy failure. J. Biol. Chem. 284, 17668–17676 (2009)
Hawley, S. A. et al. Use of cells expressing γ subunit variants to identify diverse mechanisms of AMPK activation. Cell Metab. 11, 554–565 (2010)
Bungard, D. et al. Signaling kinase AMPK activates stress-promoted transcription via histone H2B phosphorylation. Science 329, 1201–1205 (2010)
Fulco, M. et al. Glucose restriction inhibits skeletal myoblast differentiation by activating SIRT1 through AMPK-mediated regulation of Nampt. Dev. Cell 14, 661–673 (2008)
Ramsey, K. M. et al. Circadian clock feedback cycle through NAMPT-mediated NAD+ biosynthesis. Science 324, 651–654 (2009)
Haigis, M. C. & Guarente, L. P. Mammalian sirtuins–emerging roles in physiology, aging, and calorie restriction. Genes Dev. 20, 2913–2921 (2006)
Michishita, E. et al. SIRT6 is a histone H3 lysine 9 deacetylase that modulates telomeric chromatin. Nature 452, 492–496 (2008)
Vyas, S. et al. Family-wide analysis of poly(ADP-ribose) polymerase activity. Nat. Commun. 5, 4426 (2014)
Yang, Y. et al. A cytosolic ATM/NEMO/RIP1 complex recruits TAK1 to mediate the NF-κB and p38 mitogen-activated protein kinase (MAPK)/MAPK-activated protein 2 responses to DNA damage. Mol. Cell. Biol. 31, 2774–2786 (2011)
Stilmann, M. et al. A nuclear poly(ADP-ribose)-dependent signalosome confers DNA damage-induced IκB kinase activation. Mol. Cell 36, 365–378 (2009)
Martin, N. et al. PARP-1 transcriptional activity is regulated by sumoylation upon heat shock. EMBO J. 28, 3534–3548 (2009)
Matheny, C. J. et al. Next-generation NAMPT inhibitors identified by sequential high-throughput phenotypic chemical and functional genomic screens. Chem. Biol. 20, 1352–1363 (2013)
Lambert, P. F., Kashanchi, F., Radonovich, M. F., Shiekhattar, R. & Brady, J. N. Phosphorylation of p53 serine 15 increases interaction with CBP. J. Biol. Chem. 273, 33048–33053 (1998)
Acknowledgements
This work was supported in part the National Cancer Institute grant CA92577, by a fellowship from the National Foundation for Cancer Research, and by aTyr Pharma through an agreement with The Scripps Research Institute. We thank the The Scripps Research Institute mouse facility for their efforts for this project. We also thank P. Chang for the ZZ-PARP1 clone, and the PARG and PARG-MT proteins, and Y. Shi for independently repeating some of the key experiments. We thank P. Chang, J. H. Chung, L. Guarente, L. Krauss, D. Sinclair, and C. Westphal for comments and suggestions on this work.
Author information
Authors and Affiliations
Contributions
M.S. and P.S. conceived the idea, designed the research, analysed the data, and wrote the manuscript. M.S. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Resveratrol inhibits TyrRS activation.
a, The ATP–PPi exchange assay as described in Methods demonstrated the inhibitory effect of resveratrol on TyrRS and b resveratrol shifts the Michaelis constant (Km) for tyrosine. c, Resveratrol binds TyrRS better than tyrosine. The apparent Ki for resveratrol was deduced by varying the concentration of RSV and plotting the slope of (1/v versus 1/[Tyr]) versus [RSV] as indicated.
Extended Data Figure 2 Resveratrol induces a distinct conformational change upon binding to active site of TyrRS.
a, Comparison of the overall conformational change induced by resveratrol at the active site of TyrRS by structure-based superposition (yellow, tyrosine-bound structure; magenta, resveratrol-bound structure). Note the conformational change near the helix region (P331–P342) that connects the linker region with the C-domain. b, Illustration of the extensive interactions of resveratrol with the active site. c, The trans-resveratrol (dark blue) docks (manual) into TyrRS active site without significant structural disturbances. d, Generation of a new pocket through a RSV-induced conformational change in TyrRS accommodates the dihydroxy phenolic ring of RSV (otherwise exposed to the destabilizing aqueous environment in the trans form) and hence facilitates the trans (dark blue) to cis (light blue) conversion of RSV.
Extended Data Figure 3 Resveratrol facilitates the TyrRS–PARP1 interaction in an active-site-dependent manner.
a, Both heat shock (42 °C for 30 min) and tunicamycin-treatment (10 μg ml−1, endoplasmic reticulum stress) facilitated the nuclear translocation of TyrRS and activation of PARP1. b, Resveratrol or serum starvation facilitate TyrRS interaction with PARP1, and Tyr-SA prevents this interaction. ZZ-PARP1 was immunoprecipiated with IgG from HeLa cells treated with RSV or serum starvation alone or in combination with Tyr-SA. c, Resveratrol- or serum-starvation-mediated PARP1 activation is blocked only by Tyr-SA and not by Gly-SA. d, TyrRS interacts directly with PARP1. HeLa cell lysate after RSV treatment (5 μM, 30 min) was divided into three parts and treated with PARG and catalytically inactive PARG-MT. PARP1 was immunoprecipitated and analysed for TyrRS interaction. e, Model illustrating the mechanism of RSV-mediated TyrRS interaction with PARP1 and subsequent release after auto-PARylation. f, Ni-NTA pull-down of N- and C-terminal fragments of PARP1 overexpressed in E. coli demonstrates that TyrRS interacts with the C-terminal region of PARP1. g, Only the full-length TyrRS (1–528), but none of the various fragments of TyrRS (mini-TyrRS (1–364), ΔN-TyrRS (228–528), or the C-domain (328–528)), interacts with PARP1. h, Coomassie blue staining of a gel showing the total protein input in for the experiment of Extended Data Fig. 3g.
Extended Data Figure 4 Tyrosyl-AMP analogue (Tyr-SA) does not affect DNA-dependent auto-PARylation of PARP1.
a, Silver-stained SDS–PAGE gel showing the purity and input of PARP1 and TyrRS in the in vitro PARylation study of Fig. 2. b, Quantitation (Image J software) of the band intensity of PARylated PARP1 in Fig. 2a, top. c, Tyrosyl-AMP analogue (Tyr-SA) does not affect DNA-dependent auto-PARylation of PARP1. d, Overexpression of nuclear-translocation-weakened mutant of TyrRS6 is less effective in activating PARP1. e, Y314A-TyrRS is more sensitive to RSV than is TyrRS in facilitating PARP1 activation.
Extended Data Figure 5 Resveratrol enhances the acetylation of Tip60 and modulates NAD+ concentration in a dose- and time-dependent manner.
a, Treatment of HeLa cells (1 h) with increasing concentration of resveratrol enhances the acetylation level of Tip60. Activation of Tip60 was monitored by histone acetylation status. b, Total NAD+ contents of serum-starved cells or RSV-treated samples were compared with untreated samples at 15 min using a commercially available BioVision NAD+/NADH quantitation colorimetric kit. c, Total nicotinamide or ADP-ribose produced was deduced from the difference in the amount of NAD+ in each sample with respect to the untreated sample (consumption of one mole of NAD+ would give rise one mole of nicotinamide and one mole of ADP-ribose). d, Total NAD+ content of the serum-starved cells or RSV-treated samples were compared with untreated samples at 1 h. (Although the experiments were done in biological triplicates (all samples showing similar results), the error bars in the figure represent the deviations from the mean of the technical triplicates from one representative biological sample.) e, f, Time course study of poly-ADP-ribosylation status and associated signalling events after (e) serum starvation (extended time course data of the same image shown in Fig. 3c) and (f) treatment with 5 μM RSV. Using the respective antibodies, Activation of p53 was monitored by the induction of p21 and SIRT6. Activation of NRF2 was monitored by HO-1 induction.
Extended Data Figure 6 siRNA (siRNATyrRS or siRNAPARP1), with and without low RSV (5 μM), does not affect cell viability.
HeLa cells (1 × 106) were reverse-transfected with siRNA targeted to TyrRS or PARP1. An siRNACon (a scrambled sequence of siRNAPARP1) was used as a control. Viability was monitored using the RTCA iCELLigence System (ACEA Biosciences). Samples were treated with RSV (5 μM) at 60 h and monitoring was continued for another 2 h for siRNATyrRS (a total of 62 h of monitoring) and for another 16 h for siRNACon and siRNAPARP1 (total 76 h monitoring).
Extended Data Figure 7 siRNASIRT1 did not affect downstream signalling events at low RSV (5 μM).
HeLa cells were treated with siRNASIRT1 for 60 h to knockdown SIRT1. HeLa cells were treated with RSV (5 μM) for another 4 h and samples were collected intervals as indicated. Samples were analysed for downstream signalling markers using appropriate antibodies.
Extended Data Figure 8 siRNA- (siRNATyrRS or siRNAPARP1) treated cells did not upregulate the levels of NAD+ in response to RSV (5 μM) after 1 h.
HeLa cells (1 × 106) were reverse transfected, separately, with siRNA targeted to PARP1 or TyrRS. A scrambled sequence of target siRNA was used as a control. The total NAD+ content of RSV (5 μM)-treated samples was compared with untreated samples at 1 h, using a commercially available BioVision NAD+/NADH quantitation colorimetric kit. (Although the experiments were done in biological triplicates (all samples showing similar results), the error bars in the figure represent the deviations from the mean of the technical triplicates from one representative biological sample.) The comparator (shown as a dashed bar) is taken from Extended Data Fig. 5d.
Extended Data Figure 9 Resveratrol treatment activates PARP1 and associated signalling events in the mouse tissues.
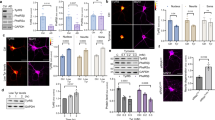
a, Activation of PARP1 in mouse muscle tissue treated with resveratrol monitored by increased PARylation and b by increased acetylation status. Activations of Tip60 and AMPK were monitored by using α-AcK16-H4 and α-pSer36-H2B, respectively. c, Activation of PARP1 in mouse heart tissue treated with resveratrol monitored by increased PARylation and d by increased acetylation status. e, Resveratrol treatment causes only a transient activation on PARP1. Immunoblotting of mouse muscle tissue samples after 24 h of RSV treatment showed no significant difference in the level of PARP1PAR compared with control. f, RSV treatment enhances TyrRS interaction with and activation of PARP1 in the muscle tissue. g and h, Resveratrol-mediated activation of PARP1 (g, monitored by PARylation status; h, monitored by acetylation status) is blocked by Tyr-SA in mouse heart tissues.
Rights and permissions
About this article
Cite this article
Sajish, M., Schimmel, P. A human tRNA synthetase is a potent PARP1-activating effector target for resveratrol. Nature 519, 370–373 (2015). https://doi.org/10.1038/nature14028
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14028
This article is cited by
-
Arg-tRNA synthetase links inflammatory metabolism to RNA splicing and nuclear trafficking via SRRM2
Nature Cell Biology (2023)
-
Resveratrol blocks retrotransposition of LINE-1 through PPAR α and sirtuin-6
Scientific Reports (2022)
-
Cis- and trans-resveratrol have opposite effects on histone serine-ADP-ribosylation and tyrosine induced neurodegeneration
Nature Communications (2022)
-
Effects of Glycoalkaloids from Solanum lycocarpum on Genomic Instability
Revista Brasileira de Farmacognosia (2022)
-
Modulation of Excitatory Synaptic Transmission During Cannabinoid Receptor Activation
Cellular and Molecular Neurobiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.