Abstract
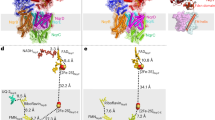
NADH oxidation in the respiratory chain is coupled to ion translocation across the membrane to build up an electrochemical gradient. The sodium-translocating NADH:quinone oxidoreductase (Na+-NQR), a membrane protein complex widespread among pathogenic bacteria, consists of six subunits, NqrA, B, C, D, E and F. To our knowledge, no structural information on the Na+-NQR complex has been available until now. Here we present the crystal structure of the Na+-NQR complex at 3.5 Å resolution. The arrangement of cofactors both at the cytoplasmic and the periplasmic side of the complex, together with a hitherto unknown iron centre in the midst of the membrane-embedded part, reveals an electron transfer pathway from the NADH-oxidizing cytoplasmic NqrF subunit across the membrane to the periplasmic NqrC, and back to the quinone reduction site on NqrA located in the cytoplasm. A sodium channel was localized in subunit NqrB, which represents the largest membrane subunit of the Na+-NQR and is structurally related to urea and ammonia transporters. On the basis of the structure we propose a mechanism of redox-driven Na+ translocation where the change in redox state of the flavin mononucleotide cofactor in NqrB triggers the transport of Na+ through the observed channel.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
Coordinates and structure factors for the entire complex of Na+-NQR and of individual subunits NqrA1–377, NqrC33–257 and NqrF129–408 have been deposited in the Protein Data Bank. The PDB accession codes are 4P6V (entire NQR complex), 4U9O (subunit NqrA, crystal 1), 4U9Q (subunit NqrA, crystal 2), 4U9S (subunit NqrC), and 4U9U (subunit NqrF).
References
Mitchell, P. Coupling of phosphorylation to electron and hydrogen transfer by a chemi-osmotic type of mechanism. Nature 191, 144–148 (1961)
Pfenninger-Li, X. D., Albracht, S. P., van Belzen, R. & Dimroth, P. NADH:ubiquinone oxidoreductase of Vibrio alginolyticus: purification, properties, and reconstitution of the Na+ pump. Biochemistry 35, 6233–6242 (1996)
Juárez, O., Nilges, M. J., Gillespie, P., Cotton, J. & Barquera, B. Riboflavin is an active redox cofactor in the Na+-pumping NADH: quinone oxidoreductase (Na+-NQR) from Vibrio cholerae. J. Biol. Chem. 283, 33162–33167 (2008)
Casutt, M. S. et al. Localization of ubiquinone-8 in the Na+-pumping NADH:quinone oxidoreductase from Vibrio cholerae. J. Biol. Chem. 286, 40075–40082 (2011)
Häse, C. C. & Mekalanos, J. J. Effects of changes in membrane sodium flux on virulence gene expression in Vibrio cholerae. Proc. Natl Acad. Sci. USA 96, 3183–3187 (1999)
Müller, V., Imkamp, F., Biegel, E., Schmidt, S. & Dilling, S. Discovery of a ferredoxin:NAD+-oxidoreductase (Rnf) in Acetobacterium woodii. Ann. NY Acad. Sci. 1125, 137–146 (2008)
Holm, L. & Rosenstrom, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010)
Baradaran, R., Berrisford, J. M., Minhas, G. S. & Sazanov, L. A. Crystal structure of the entire respiratory complex I. Nature 494, 443–448 (2013)
Reyes-Prieto, A., Barquera, B. & Juárez, O. Origin and evolution of the sodium-pumping NADH: ubiquinone oxidoreductase. PLoS ONE 9, e96696 (2014)
Nedielkov, R., Steffen, W., Steuber, J. & Möller, H. M. NMR reveals double occupancy of quinone-type ligands in the catalytic quinone binding site of the Na+-translocating NADH:quinone oxidoreductase from Vibrio cholerae. J. Biol. Chem. 288, 30597–30606 (2013)
Levin, E. J. et al. Structure and permeation mechanism of a mammalian urea transporter. Proc. Natl Acad. Sci. USA 109, 11194–11199 (2012)
Khademi, S. et al. Mechanism of ammonia transport by Amt/MEP/Rh: structure of AmtB at 1.35 Å. Science 305, 1587–1594 (2004)
Andrade, S. L., Dickmanns, A., Ficner, R. & Einsle, O. Crystal structure of the archaeal ammonium transporter Amt-1 from Archaeoglobus fulgidus. Proc. Natl Acad. Sci. USA 102, 14994–14999 (2005)
Wacker, T., Garcia-Celma, J. J., Lewe, P. & Andrade, S. L. Direct observation of electrogenic NH4+ transport in ammonium transport (Amt) proteins. Proc. Natl Acad. Sci. USA 111, 9995–10000 (2014)
Juárez, O., Athearn, K., Gillespie, P. & Barquera, B. Acid residues in the transmembrane helices of the Na+-pumping NADH:quinone oxidoreductase from Vibrio cholerae involved in sodium translocation. Biochemistry 48, 9516–9524 (2009)
Juárez, O., Morgan, J. E., Nilges, M. J. & Barquera, B. Energy transducing redox steps of the Na+-pumping NADH:quinone oxidoreductase from Vibrio cholerae. Proc. Natl Acad. Sci. USA 107, 12505–12510 (2010)
Casutt, M. S., Schlosser, A., Buckel, W. & Steuber, J. The single NqrB and NqrC subunits in the Na+-translocating NADH:quinone oxidoreductase (Na+-NQR) from Vibrio cholerae each carry one covalently attached FMN. Biochim. Biophys. Acta 1817, 1817–1822 (2012)
Duffy, E. B. & Barquera, B. Membrane topology mapping of the Na+-pumping NADH:quinone oxidoreductase from Vibrio cholerae by PhoA/GFP fusion analysis. J. Bacteriol. 188, 8343–8351 (2006)
Bertsova, Y. V. et al. Alternative pyrimidine biosynthesis protein ApbE is a flavin transferase catalyzing covalent attachment of FMN to a threonine residue in bacterial flavoproteins. J. Biol. Chem. 288, 14276–14286 (2013)
Casutt, M. S. et al. Localization and function of the membrane-bound riboflavin in the Na+-translocating NADH:quinone oxidoreductase (Na+-NQR) from Vibrio cholerae. J. Biol. Chem. 285, 27088–27099 (2010)
Terwilliger, T. C., Klei, H., Adams, P. D., Moriarty, N. W. & Cohn, J. D. Automated ligand fitting by core-fragment fitting and extension into density. Acta Crystallogr. D 62, 915–922 (2006)
Page, C. C., Moser, C. C., Chen, X. & Dutton, P. L. Natural engineering principles of electron tunnelling in biological oxidation-reduction. Nature 402, 47–52 (1999)
Tao, M., Casutt, M. S., Fritz, G. & Steuber, J. Oxidant-induced formation of a neutral flavosemiquinone in the Na+-translocating NADH:quinone oxidoreductase (Na+-NQR) from Vibrio cholerae. Biochim. Biophys. Acta 1777, 696–702 (2008)
Zhang, Z. et al. Electron transfer by domain movement in cytochrome bc1 . Nature 392, 677–684 (1998)
Verkhovsky, M. I. et al. Sodium-dependent movement of covalently bound FMN residue(s) in Na+-translocating NADH:quinone oxidoreductase. Biochemistry 51, 5414–5421 (2012)
Steuber, J. et al. Central role of the Na+-translocating NADH:quinone oxidoreductase (Na+-NQR) in sodium bioenergetics of Vibrio cholerae. Biol. Chem. 395, 1389–1399 (2014)
Rich, P. R., Meunier, B. & Ward, F. B. Predicted structure and possible ionmotive mechanism of the sodium-linked NADH-ubiquinone oxidoreductase of Vibrio alginolyticus. FEBS Lett. 375, 5–10 (1995)
Bogachev, A. V., Bertsova, Y. V., Aitio, O., Permi, P. & Verkhovsky, M. I. Redox-dependent sodium binding by the Na+-translocating NADH:quinone oxidoreductase from Vibrio harveyi. Biochemistry 46, 10186–10191 (2007)
Verkhovsky, M. I. & Bogachev, A. V. Sodium-translocating NADH:quinone oxidoreductase as a redox-driven ion pump. Biochim. Biophys. Acta 1797, 738–746 (2010)
Bogachev, A. V., Belevich, N. P., Bertsova, Y. V. & Verkhovsky, M. I. Primary steps of the Na+-translocating NADH:ubiquinone oxidoreductase catalytic cycle resolved by the ultrafast freeze-quench approach. J. Biol. Chem. 284, 5533–5538 (2009)
Bogachev, A. V., Bertsova, Y. V., Ruuge, E. K., Wikstrom, M. & Verkhovsky, M. I. Kinetics of the spectral changes during reduction of the Na+-motive NADH:quinone oxidoreductase from Vibrio harveyi. Biochim. Biophys. Acta 1556, 113–120 (2002)
Zhu, W. & Becker, D. F. Flavin redox state triggers conformational changes in the PutA protein from Escherichia coli. Biochemistry 42, 5469–5477 (2003)
Leung, K. K. & Shilton, B. H. Chloroquine binding reveals flavin redox switch function of quinone reductase 2. J. Biol. Chem. 288, 11242–11251 (2013)
Bogachev, A. V., Murtazina, R. A. & Skulachev, V. P. The Na+/e− stoichiometry of the Na+-motive NADH:quinone oxidoreductase in Vibrio alginolyticus. FEBS Lett. 409, 475–477 (1997)
Casutt, M. S., Wendelspiess, S., Steuber, J. & Fritz, G. Crystallization of the Na+-translocating NADH:quinone oxidoreductase from Vibrio cholerae. Acta Crystallogr. F 66, 1677–1679 (2010)
Barquera, B. et al. Purification and characterization of the recombinant Na+-translocating NADH:quinone oxidoreductase from Vibrio cholerae. Biochemistry 41, 3781–3789 (2002)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D 59, 2023–2030 (2003)
Abrahams, J. P. & Leslie, A. G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D 52, 30–42 (1996)
Cowtan, K. Error estimation and bias correction in phase-improvement calculations. Acta Crystallogr. D 55, 1555–1567 (1999)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Söding, J., Biegert, A. & Lupas, A. N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, W244–W248 (2005)
Tao, M. et al. Crystallization of the NADH-oxidizing domain of the Na+-translocating NADH:ubiquinone oxidoreductase from Vibrio cholerae. Acta Crystallogr. F 62, 110–112 (2006)
Vohl, G. et al. Crystallization and preliminary analysis of the NqrA and NqrC subunits of the Na+-translocating NADH:ubiquinone oxidoreductase from Vibrio cholerae. Acta Crystallogr. F 70, 987–992 (2014)
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D 66, 22–25 (2010)
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011)
Headd, J. J. et al. Use of knowledge-based restraints in phenix.refine to improve macromolecular refinement at low resolution. Acta Crystallogr. D 68, 381–390 (2012)
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Chovancova, E. et al. CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLOS Comput. Biol. 8, e1002708 (2012)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
The PyMOL Molecular Graphics System, v. 1.3r1 (Schrödinger, LLC, 2010)
Sääf, A., Johansson, M., Wallin, E. & von Heijne, G. Divergent evolution of membrane protein topology: the Escherichia coli RnfA and RnfE homologues. Proc. Natl Acad. Sci. USA 96, 8540–8544 (1999)
Bogachev, A. V., Bertsova, Y. V., Bloch, D. A. & Verkhovsky, M. I. Thermodynamic properties of the redox centres of Na+-translocating NADH:quinone oxidoreductase. Biochemistry 45, 3421–3428 (2006)
Bogachev, A. V. et al. Redox properties of the prosthetic groups of Na+-translocating NADH:quinone oxidoreductase. 1. Electron paramagnetic resonance study of the enzyme. Biochemistry 48, 6291–6298 (2009)
Acknowledgements
We thank the staff at beamlines X06SA and X06DA at Swiss Light Source for excellent support. This work was supported by contract research ‘Methoden in den Lebenswissenschaften’ of the Baden-Württemberg Stiftung P-LS-Meth/4 (to J.S. and G.F.), and by the Deutsche Forschungsgemeinschaft grant FR 1321/3-1 (to J.S.) and grant FR 1488/3-2 (to G.F.). We thank Y. Obermaier for expression and preparation of Na+-NQR.
Author information
Authors and Affiliations
Contributions
J.S., G.F. and T.V. developed expression constructs; J.S. and M.S.C. developed purification procedures; T.V., M.S.C. and G.V. expressed the protein; M.S.C., G.V. and G.F. purified the entire complex or single subunits; M.S.C., G.V. and G.F. performed crystallization, crystal harvesting and data collection. G.F. and K.D. performed data processing and determination of phases. G.F. performed model building and refinement. G.F. prepared the figures. G.F. and J.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Sequence alignment of the integral membrane subunit NqrB from different organism with the corresponding subunits of the RNF complex.
The localization of transmembrane helices is indicated by cylinders. Connecting loops located in the cytoplasm are shown in red, connecting loops located in the periplasm in blue. Thr 236 covalently binding the FMN, and Asp 346 located in the proposed Na+ channel are indicated by arrows.
Extended Data Figure 2 Sequence alignment of the integral membrane subunits NqrD and NqrE from different organism with the corresponding subunits of the RNF complex.
a, b, The localization of transmembrane helices is indicated by cylinders; connecting loops located in the cytoplasm are shown in red, connecting loops located in the periplasm in blue. Cys residues in NqrD (a) and NqrE (b) coordinating the Fe are indicated by arrows.
Extended Data Figure 3 Topology of the transmembrane subunits NqrB, NqrD, and NqrE and arrangement of transmembrane helices.
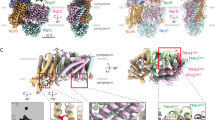
a–c, The schematic topology of the transmembrane helices of NqrB, NqrC and NqrD is shown on the left hand side and the corresponding structure on the right hand side. The membrane plane is indicated in grey and the cytoplasmic aspect is marked by C and the periplasmic aspect by P. a, NqrB contains ten transmembrane helices which can be divided into a N-terminal domain comprising helices I–V and a C-terminal domain comprising helices VI–X, which exhibit an inverted topology. Both domains are connected by a long periplasmic linker. The domains exhibit an inverted topology and align with an r.m.s.d. of 3.3 Å over 113 Cα positions. b, c, NqrD and NqrE each comprise six helices exhibiting an inverted topology. Helix I and helix IV of both subunits are composed of two half helices. Such an inverted topology had been predicted based on the sequence information54. d, Top view from the cytoplasmic side onto the transmembrane helices of subunits NqrB, NqrC, NqrD, NqrE and NqrF. There are a total of 24 transmembrane helices. NqrD and NqrE form a central symmetrical unit. Subunit NqrB resides on one side of the NqrD–E unit whereas the single transmembrane helices from NqrC and NqrF reside on the opposed side. NqrB is closely attached to NqrE via helices V and VI from NqrE and IV, V, IX and X from NqrB, forming an interaction surface of 1,280 Å2, whereas NqrD exhibits a much smaller contact area to NqrB via helices VI from NqrD and IV and V from NqrB, covering 335 Å2. The transmembrane helices of NqrC and NqrF are close to each other but interact with different subunits: the transmembrane helix of NqrC forms contacts with helix III of NqrD, whereas the transmembrane helix of NqrF interacts with helix III of NqrE. e, Top view of the transmembrane part of Na+-NQR and 2Fo − Fc electron density displayed at a contour level of 1.0σ. The map coefficients were sharpened by a B-factor of −80 Å2.
Extended Data Figure 4 Subunit NqrA.
a, Interactions of NqrA with other subunits in the Na+-NQR complex. The subunits of Na+-NQR are shown in different colours: NqrA in blue, NqrB in orange, NqrC in green, NqrD in magenta, NqrE in cyan, and NqrF in red. Subunit B is shown as cartoon and all other subunits as surface representation. The C-terminal domain of NqrA located proximal to the membrane forms minor contacts with the integral membrane subunit NqrB via the NqrA residues 376–379 and 425–428, located in two short loops. A long N-terminal stretch of NqrB encompassing residues 39–53 lies in a groove of NqrA interacting over a total area of 820 Å2 and anchoring NqrA to the membrane subunits. The residues shown as transparent van der Waals spheres fill almost the entire groove of NqrA. At the C terminus of NqrB, transmembrane helix 10 is elongated and protrudes into the cytoplasm, forming contacts with the C-terminal domain and the Rossmann-fold domain of NqrA, covering a total area of 430 Å2. b, c, NqrA is composed of four domains, an N-terminal domain similar to a biotin carboxyl carrier domain (blue, residues 28–100), a Rossmann-fold domain (green, residues 102–254), an ubiquitin-like domain (orange, residues 258–329), and a C-terminal helical domain (red, residues 376–446). The N-terminal residues 1–27 wrap around the Rossmann-fold domain and the ubiquitin-like domain and form two short β-strands that align with β-sheets of both domains, respectively. The C-terminal helical domain of NqrA shows similarity to a 2[4Fe–4S] cluster ferredoxin fold like for example, in fumarate reductase (PDB code 1KF6), but does not contain a FeS centre. Consistently, the Cys residues required for FeS coordination are not present in NqrA. d, Structural alignment of NqrA with Nqo1 from complex I (grey). The proteins align with an r.m.s.d. of 3.9 Å over 234 Cα positions. NqrA comprises a deep solvent-accessible cavity that is formed by residues of the Rossmann-fold domain and the ubiquitin-like domain that is large enough to accommodate ubiquinone. In case of Nqo1 of complex I the corresponding cavity harbours the isoalloxazine moiety of the FMN cofactor.
Extended Data Figure 5 A putative Na+ channel in subunit NqrB.
a, b, Structural alignments of NqrB with urea transporter and ammonium transporter are shown. In NqrB the central helices I, III, VI and VIII form a membrane-spanning channel. Some backbone carbonyls, for example, from Val 161, Ile 164, Leu 168 from helix III deviate notably from the ideal geometry and point inwards the channel. Such a distortion indicates a putative involvement in Na+ coordination. a, The left hand side represents the side view and the right hand side the top view of NqrB (orange) aligned with bovine urea transporter (blue). Helix VIII of NqrB carrying residues forming the constriction is shown in red. The gating helices of urea transporter, which have no corresponding helices in NqrB, are depicted in dark blue. b, Structural alignment of NqrB (orange) with ammonium transporter from Archaeoglobus fulgidus. The outer helix of ammonium transporter that has no homologous helix in NqrB is shown in grey. The high structural similarity of NqrB with urea and ammonium transporter shows that the subunit preserved the basic architecture of a transporter, but has acquired an additional and completely different function as a redox protein. These structural rearrangements in the periplasmic aspect of NqrB required to embed the FMN cofactor might have contributed to the closure of the channel. c, Cross section through NqrB. The surface is coloured according to the electrostatic surface potential. The cytoplasmic half channel exhibits a negative surface charge (red) whereas the periplasmic half channel is positively charged (blue). The localization of residues Phe 338, Phe 342 and Asp 346 is indicated. The constriction is located halfway through the membrane. The borders of the cytoplasmic membrane are indicated by grey lines.
Extended Data Figure 6 Localization of riboflavin.
A large patch of Fo − Fc density was observed between NqrB (orange) and NqrE (cyan) and assigned to the riboflavin. The isoalloxazine moiety of riboflavin fits well into the Fo − Fc density. Several interactions with the protein matrix can stabilize the riboflavin. The flavin is stacked between the side chain of Val 399 and the CB, CG of Glu 402 of NqrB on one side (Si side) and the side chain of Phe 39 of NqrE on the opposed side (Re-side). Moreover, the imidazole of His 398 of NqrB on the Si-side can form a hydrogen bond to N5 of isoalloxazine.
Rights and permissions
About this article
Cite this article
Steuber, J., Vohl, G., Casutt, M. et al. Structure of the V. cholerae Na+-pumping NADH:quinone oxidoreductase. Nature 516, 62–67 (2014). https://doi.org/10.1038/nature14003
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14003
This article is cited by
-
Cryo-EM structures of Na+-pumping NADH-ubiquinone oxidoreductase from Vibrio cholerae
Nature Communications (2022)
-
Molecular dynamics modeling of the Vibrio cholera Na+-translocating NADH:quinone oxidoreductase NqrB–NqrD subunit interface
Molecular and Cellular Biochemistry (2022)
-
Structural characterization of the microbial enzyme urocanate reductase mediating imidazole propionate production
Nature Communications (2021)
-
Architecture of bacterial respiratory chains
Nature Reviews Microbiology (2021)
-
Aerobic nitrogen-fixing bacteria for hydrogen and ammonium production: current state and perspectives
Applied Microbiology and Biotechnology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.