Abstract
To broaden our understanding of the evolution of gene regulation mechanisms, we generated occupancy profiles for 34 orthologous transcription factors (TFs) in human–mouse erythroid progenitor, lymphoblast and embryonic stem-cell lines. By combining the genome-wide transcription factor occupancy repertoires, associated epigenetic signals, and co-association patterns, here we deduce several evolutionary principles of gene regulatory features operating since the mouse and human lineages diverged. The genomic distribution profiles, primary binding motifs, chromatin states, and DNA methylation preferences are well conserved for TF-occupied sequences. However, the extent to which orthologous DNA segments are bound by orthologous TFs varies both among TFs and with genomic location: binding at promoters is more highly conserved than binding at distal elements. Notably, occupancy-conserved TF-occupied sequences tend to be pleiotropic; they function in several tissues and also co-associate with many TFs. Single nucleotide variants at sites with potential regulatory functions are enriched in occupancy-conserved TF-occupied sequences.
Similar content being viewed by others
Main
Determining the similarities and differences between mouse and human regulatory networks will not only improve our understanding of the evolution of regulatory mechanisms, but also help to interpret biomedical insights derived from research performed on mouse models. Recent genome-wide binding studies of eight TFs in several species uncovered many regulatory networks that have been highly rewired since the divergence of ancestors to mouse and human1,2,3,4, consistent with early studies in other species5. These results contrast sharply with other data showing that conservation of genomic DNA sequences can be a useful guide to discovery of regulatory regions6, and that the regulatory landscape can be highly conserved among more distant species7. Considering the large numbers of known TFs and their functional diversity, comprehensive studies on a broader range of TFs are needed to resolve these apparent discrepancies. Furthermore, our knowledge of the functional consequences of either divergence or conservation of TF occupancy remains limited.
The mouse–human orthologous occupancy profiles
To examine conservation of TF binding regions both between species and across different cell types, we generated and analysed a large data set of genome-wide binding profiles for 34 TFs in mouse and human. A diverse panel of TFs were chosen including those that bind DNA through specific consensus sequences, comprise part of the general transcriptional machinery such as RNA polymerase 2 (POL2), and modify or remodel chromatin (Extended Data Fig. 1a and Supplementary Information). For simplicity, we refer to the entire collection as TFs, even though some are general factors. We focused on occupancy by 32 TFs in cell line models for erythroid progenitors (mouse erythroleukaemia MEL and human leukaemia K562 cells) and lymphoblasts (mouse lymphoma CH12 and human B lymphoblastoid GM12878 cells) in mouse and human, and we also showed that the results are similar to those obtained in mouse and human embryonic stem cells (Extended Data Fig. 8). Chromatin immunoprecipitation with massively parallel sequencing (ChIP-seq) assays were conducted using replicate experiments and in accordance with ENCODE standards8. A total of 120 data sets were generated and analysed.
Conserved and non-conserved features
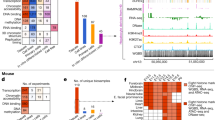
These genome-wide binding data for a large and diverse set of TFs revealed both conserved and non-conserved features of TF occupancy between mouse and human. First, although most TFs can reside at both promoters and distal sites, each shows a pronounced preference (Fig. 1a and Extended Data Fig. 2a, b). The preference is strongly conserved between mouse and human (R = 0.8; Extended Data Fig. 2c). The one exception is ETS1. Even though the primary motif in ETS1 is conserved between mouse and human (Fig. 1b), it preferentially binds proximal to promoters in human but not in mouse. ETS1 is responsible for the mouse-specific expression of the T-cell marker Thy-1 in the thymus9, and we propose that this marked difference in its binding location may contribute to immune system differences between mouse and human10. Second, although the primary motifs of most sequence-specific TFs are conserved between mouse and human, the secondary motifs (for example, motifs of associated factors; see Supplementary Information) tend to be lineage-specific (Fig. 1b and Extended Data Fig. 2d), indicating a change in co-associated partners.
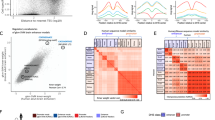
a, Each row represents one TF, and each column represents one genomic region. Heat-map colour shows the proportions of TF OSs (combination of different cell lines in the same species) that are located in each genomic region. b, Motif comparison for sequence-specific TFs examined in lymphoblast cells. In the right panel, each row represents one TF. The level of motif conservation is encoded by colour. Detailed results for the USF2 example are in the left panels. Peaks were divided into different bins according to the occupancy signal (higher signal on the left, lower on the right). The proportions of peaks with the motif in each bin (red lines) and the average distances between motif sites and peak summit in each bin (grey lines) are plotted against ranks of peak bins. Red dots indicate the proportion of control regions (±500 bp flanking the USF2 OS) that have the motif. NA, not available. c, TF OS chromatin state preference comparison between MEL and K562 cells. Heat map shows the percentage of TF OSs (rows) that overlap with eight different chromatin states (columns). d, The average signal distributions for MeDIP-seq and MRE-seq in MEL and K562 cells. Five-kilobase flanking regions centred on the TF OS peak summits were divided into 50-bp bins. Signals were aggregated in each bin.
The preferred chromatin states, defined by histone modifications, for occupied sequences (OSs) of orthologous TFs are also conserved between mouse and human. Using data on five histone modifications, the mouse and human genomes were segmented into eight chromatin states (Fig. 1c and Extended Data Fig. 3a, b). Most TF OSs are located in states characteristic of promoters and enhancers (states 1–4). By contrast, approximately 50% of OSs for the CTCF–cohesin complex (CTCF, RAD21 and SMC3)11,12 are located in state 5 and 8, which mark quiescent regions with very low signal for all the histone modifications. MAFK also shows preference for quiescent regions. Notably, both the CTCF–cohesin complex and MAFK13 can mediate long-range interactions in the genome. The state preference is conserved between mouse and human (Fig. 1c; R = 0.9; Extended Data Fig. 3b), suggesting that the overall functions of the occupied segments are similar in the two species. Indeed, the proportion of enhancers, predicted by a different approach14,15, is also conserved (R = 0.7) (Extended Data Fig. 4).
We also examined DNA methylation profiles in TF OSs by using both methylated DNA immunoprecipitation (MeDIP) and DNA digestion with methyl-sensitive restriction enzymes followed by sequencing (MRE-seq)16. The TF OSs are highly enriched for MRE-seq signals and depleted of MeDIP-seq signals, showing that TF OSs are generally hypomethylated in both species (Fig. 1d and Extended Data Fig. 3c).
TF- and location-specific occupancy conservation
The TF binding regions are enriched for conservation of DNA sequences, showing a strong signal for evolutionary constraint within ±50 base pairs (bp) of ChIP-seq peak summits (Fig. 2a). This result indicates that purifying selection has acted on DNA sequences in many of the TF OSs, but it does not mean that all TF OSs are uniformly under constraint. Approximately 50% of TF OSs do not align between mouse and human15 because either they are lineage-specific sequences such as transposable elements17, or they have diverged to an extent that they no longer align.
a, Blue and purple lines represent the average phyloP score distribution near (±100 bp) the ChIP-seq peak summit in human and mouse. The grey line represents the distribution for randomly selected background sequences. The x axis is the distance to the peak summit, and the y axis is the average phyloP score. b, The heat map represents the occupancy conservation of TF (rows) OSs in the four cell lines. The colour intensity represents the proportion of TF OSs for which occupancy is conserved between mouse and human in different genomic regions (columns). c, Comparison of the chromatin state change between TF OSs and orthologous sequences. TF OSs that can be aligned between mouse and human are divided into two groups according to the occupancy conservation status (‘occupancy conserved’ versus ‘occupancy not conserved’). Top, the y axis is the proportion of TF OSs and their orthologous sequences in each chromatin state. Bottom, detailed chromatin state change in human orthologues for mouse TF OSs in chromatin states 1 and 3. The pie charts show the distribution of chromatin states in the orthologous sequence in the second species. d, Comparison of the DNA methylation change between TF OSs and orthologous sequences. The y axis gives the normalized DNA methylation signals (MeDIP-seq). TF OSs are divided into two categories according to the occupancy conservation status as in c.
We then focused on the subset of TF OSs in which the sequences aligned between mouse and human to determine whether orthologous DNA sequences are also occupied by orthologous TFs (details in Supplementary Methods). Notably, the proportion of TF OSs at which occupancy was conserved varied markedly both among TFs and with the genomic locations (Fig. 2b). Conservation of occupancy is consistently higher in the promoter regions and lower in distal regions for almost all TFs, suggesting that the promoters may be under stronger selection than distal enhancers. Conserved promoter occupancy is observed both for factors that bind near promoters (NRF1 and MAZ) and for factors with a minority of binding sites in promoter regions (for example, MEF2A and TAL1). A notable exception is the CTCF–cohesin complex, which not only shows high levels of occupancy conservation as described previously18, but also the conservation remains high at proximal, middle and distal regions relative to the transcription start site (TSS) (Fig. 2b). These patterns of variation in conservation of occupancy are robust. One potential confounding factor is the tendency for promoter sequences to be more conserved than other regulatory regions, but adjusting the occupancy conservation by the sequence conservation difference revealed similar trends, that is, the OSs in promoter regions are more conserved than those in other regions (Extended Data Fig. 5a). Similarly, removal of the few TFs for which markedly different numbers of peaks were called between mouse and human did not change the patterns of conservation of occupancy (Extended Data Fig. 5b and Supplementary Information).
Next, we investigated how epigenetic factors influence TF binding at orthologous sites between mouse and human. As expected, the distribution of chromatin states is highly similar for occupancy-conserved TF OSs. For orthologues of TF OSs that can be aligned between the two species but are bound only in one species, a smaller proportion were in enhancer-associated states (states 3 and 4) and a larger proportion were in either repressed (state 7) or quiescent (states 5 and 8) chromatin OSs (Fig. 2c and Extended Data Fig. 6a, b). Thus species-specific loss of TF occupancy at many sites is accompanied by a shift to repressive or quiescent chromatin. By contrast, the promoter states (states 1 and 2) were largely maintained in the second species even with the loss of TF binding. This result indicates that other TFs may help to maintain conservation of a promoter state in these regions. We also searched for changes in the level of DNA methylation between TF OSs and their orthologous sequences. DNA methylation levels remained low in both species for occupancy-conserved TF OSs (Fig. 2d and Extended Data Fig. 6c), but the DNA methylation levels were significantly increased in the unbound, orthologous sequences. Thus, species-specific loss of TF occupancy is also associated with species-specific increases in DNA methylation.
Occupancy conservation associates with pleiotropy
We proposed that TF OSs with regulatory functions in several tissues would be under increased selective pressure, and thus more likely to be conserved in occupancy. To test this hypothesis, we first examined DNase I hypersensitive sites (DHSs) across 55 mouse tissues and cell lines15 to measure the chromatin accessibility of each TF OS among different tissues. Because DHSs are a proxy for regulatory element activity19, TF OS regions accessible in multiple tissues are more likely to function in those tissues. Chromatin accessibility of TF OSs presents wide variation, ranging from tissue-specific to ubiquitous patterns (Fig. 3a). Notably, the TF OSs with more pervasive chromatin accessibility across different tissues show the highest extent of occupancy conservation between mouse and human. The association between tissue usage and occupancy conservation is general; it was observed for most of the TFs examined (Extended Data Fig. 7b, c). This association is also robust to several potential confounding factors. CTCF–cohesin complexes, which are abundant and conserved across different tissue types and species18,20, might be expected to bias the result; however, we obtained comparable results after removing all the genomic regions occupied by CTCF, RAD21 or SMC3 (Extended Data Fig. 7a). The conservation of promoter regions among several tissues and species14 might also be expected to bias our analysis, but, after removal of occupancy-conserved TF OSs that lie within 2 kilobases (kb) of TSSs, we still found that the association between tissue usage and TF occupancy conservation holds for distal TF OSs (Extended Data Fig. 7d, e). Furthermore, specifically examining distal TF OSs that overlapped with enhancers predicted by chromatin signals14 showed that broad tissue usage of presumptive enhancers tracks strongly with conservation of occupancy between mouse and human (Fig. 3b).
a, Association between occupancy conservation and chromatin accessibility across several tissues. The density plot represents the frequency that TF OSs are in accessible chromatin in varying numbers of cell types. The x axis is the Shannon index density calculated on the basis of the DHS signals in 55 tissues or cell lines in mouse; high values mean the TF OS is in accessible chromatin in many cell types. The red line shows the fraction of TF OSs at which occupancy is conserved within each bin of Shannon index. b, Association between occupancy conservation and enhancer usage across several tissues. The density plot represents the frequency that TF OSs are in chromatin indicative of enhancer activity (calculated using histone H3 acetyl Lys 27 (H3K27ac) ChIP-seq signals) in varying numbers of cell types. The x axis is the Shannon index calculated based on H3K27ac signal across 23 tissues or cell lines. The red line shows the fraction of TF OSs at which occupancy is conserved within each bin of Shannon index. pre-enhancer, presumptive enhancer. c, Results of transgenic mouse enhancer assays of ten occupancy-conserved GATA1 binding sites. The stained embryo images are highlighted by activity in different tissues: light pink for those showing enhancer activity only in heart and vascular tissues, darker pink for those with activities in other tissues. Right panel shows genes, enhancers predicted by histone modifications, chromatin states (using the software ChromHMM, see Methods), factor occupancy, and DHS signals across different tissues for regions containing two GATA1 OSs.
A prediction of our hypothesis is that occupancy-conserved TF OSs will tend to be active in multiple tissues. To test this prediction experimentally, we randomly chose ten occupancy-conserved GATA1 OSs. Even though OSs were chosen on the basis of the occupancy profile of an erythroid-specific regulatory factor, all ten conserved OSs overlapped with DHSs peaks and predicted enhancers in many tissues, such as brain (Fig. 3c). When tested for in vivo enhancer activity in transgenic mouse reporter assays at embryonic day 11.5, nine of the ten showed strong, reproducible in vivo enhancer activity, and four were active in non-erythroid tissues such as midbrain and neural tube (Fig. 3c). We expanded our analysis to examine other mouse GATA1 OSs that overlapped with previously tested enhancers deposited in the VISTA Enhancer Browser (http://enhancer.lbl.gov)21. Six GATA1 OSs that are specific to mouse generated positive enhancer assays; only one (16%) showed expression in tissues other than blood vessels and heart. By contrast, among 12 additional occupancy-conserved GATA1 OSs with in vivo enhancer activity, 6 (50%) were active in non-erythroid tissues such as midbrain (Supplementary Table 5).
Conservation and divergence of TFs co-association
Because precise gene regulation requires complex interactions among different TFs, we speculated that differences in conservation of TF occupancy may be related, at least in part, to different co-association partners. By calculating the occupancy signals for all the TFs in each TF OS, we found that, in general, occupancy-conserved TF OSs tend to be bound by more TFs compared to lineage-specific TF OSs (P < 2.2 × 10−16, two-tailed t-test; Fig. 4a), suggesting that co-association with several TFs increases the level of purifying selection on the occupied sequences. Furthermore, by examining each co-associated TF pair (Fig. 4b), we determined whether the co-associations were more enriched in occupancy-conserved versus species-specific binding sites (Fig. 4c and Extended Data Fig. 9). The relationships fell into three categories. In the first category, co-association of TFs is not linked with occupancy conservation. For example, RAD21 is highly associated with CTCF in MEL cells; however, this co-association occurs with equivalent frequency at occupancy-conserved and species-specific binding sites. In the second category, TF co-association is negatively correlated with occupancy conservation. For example, the co-association of MYC OSs with EP300, an enhancer-associated factor22, is highly enriched in the mouse-specific binding sites. In the last category, TF co-association is positively correlated with occupancy conservation, as exemplified by the co-association of MYC OSs with the co-repressor SIN3A (ref. 23), suggesting that MYC-associated repressors tend to be conserved between mouse and human.
a, Density plot shows the distribution of co-associated TF numbers in each TF-binding region. The x axis represents the total number of occupied TFs per region. b, Pair-wise TF co-association in MEL cells. The colour intensity represents the extent of co-association between the TFs denoted in the rows and columns compared to the random expectation (details in Supplementary Methods). Red represents co-association higher than random expectation, blue represents co-association lower than random expectation. c, Conditional TF OSs occupancy conservation in MEL cells. The colour intensity represents for a given TF (columns), whether the co-association with the other TF (rows) is more enriched in lineage-specific binding sites (green) or occupancy-conserved binding sites (red). The colour scale represents the extent (–log P value) of the enrichment significance.
Occupancy conservation and functional SNVs
In a previous study, we assigned putative regulatory potential to genome variations by combining high-throughput experimental data sets, computational predictions, and manual annotation24. Interestingly, even though conservation was not considered during the previous classifications, we found that single nucleotide variants (SNVs) with high regulatory potential were highly enriched in occupancy-conserved TF OSs (Extended Data Table 1a). Moreover, examination of the distribution of genome-wide association study (GWAS) single nucleotide polymorphisms (SNPs) as a function of TF OS occupancy conservation revealed a significant enrichment of GWAS SNPs in occupancy-conserved TF OSs (P < 2.2 × 10−16, Fisher’s exact test; see Supplementary Information) compared with the background distribution of all genetic variation in the SNP database (dbSNP). When examining individual phenotypes, we found that SNPs associated with several phenotypes such as type I diabetes are significantly enriched in occupancy-conserved TF OSs (P = 0.019, Fisher’s exact test; Extended Data Table 1b). However, SNPs associated with other phenotypes, such as pulmonary function, are highly human-specific (P = 0.027, Fisher’s exact test; Extended Data Table 1b). Thus, although GWAS SNPs are generally enriched in occupancy-conserved TF OSs, this enrichment is phenotype-specific.
Discussion
Here we report that the conservation of TF occupancy associates with pleiotropic functions. This observation was further validated by in vivo enhancer assays in transgenic mice. To our knowledge, this is the first systematic investigation and validation of the relationship between pleiotropic TF OSs and their occupancy conservation. The pleiotropic functions of a regulatory module subject it to several constraints that preserve the underlying motifs and occupancy patterns. However, the roles in different tissues need not be carried out by the same TF. Paralogous proteins that bind to the same DNA motif (for example, GATA5 or GATA6) could be the active proteins in non-erythroid tissues at the GATA1 OSs with conserved occupancy and pleiotropic functions. This prediction can be tested in future studies.
Cell lines were used in this study because they provide an abundant source of almost identical cells, whereas obtaining primary cells in sufficient number for a study of this scale is problematic for many cell types. One concern is that cell lines across different species may not be entirely analogous. Although this possibility cannot be ruled out, when we compared the expression profile of the four cell lines with those of many other mouse tissues, we found that both MEL and K562, and also CH12 and GM12878, were the most similar pairs (Supplementary Fig. 2a). This close similarity was also seen for genome-wide histone modification signatures (Supplementary Fig. 2b). Thus, we conclude that the K562 and MEL pair of cell lines and the GM12878 and CH12 cell-line pair are sufficiently similar for meaningful cross-species comparisons. Another concern is that the trends observed in cell lines may not be representative of primary cells. Examination of binding of five TFs in mouse and human ES cells confirmed the preferential conservation of binding at promoters and the correlation of occupancy conservation with pleiotropy of DHSs (Extended Data Fig. 8). Thus, the principles gleaned from our examination of many TFs in cell lines are likely to hold for TFs in primary cells.
Methods
ChIP-seq
ChIP for TFs was carried out as previously described25. Cultured cells for biological replicates were grown in separate batches and at separate times. In brief, 5 × 107 cells were grown to a density of 0.6–0.8 × 106 per ml, cells were then cross-linked in 1% formaldehyde for 10 min at room temperature. Nuclear lysates were sonicated using a Branson 250 Sonifier (power setting 7, 100% duty cycle for 12 × 20 s intervals), such that the chromatin fragments ranged from 50 to 2,000 bp. Information on control IgG and TF antibodies used for ChIP-seq experiments is listed in Supplementary Table 2. Protein–DNA–TF antibody complexes were captured on Protein A/G agarose beads (Millipore 16-156/16-266) and eluted in 1% SDS TE buffer at 65 °C. After cross-link reversal and DNA purification, the ChIP DNA sequencing libraries were prepared as described8. Libraries were sequenced on an Illumina Genome Analyzer II and HiSeq 2000.
Uniform ChIP-Seq data processing pipeline
We used a uniform processing pipeline to identify high confidence binding peaks in mouse and human. Reads mapping: for human ChIP-Seq, mapped reads in the form of BAM files were downloaded from ENCODE University of California, Santa Cruz (UCSC) Data Coordination Center (DCC) (http://encodeproject.org/ENCODE/downloads.html). For mouse ChIP-seq, reads were mapped by BWA26. To standardize the mapping protocol, we used custom mappability tracks to filter out multi-mapping reads and only retain unique mapping reads (reads that map to exactly one location in the genome). We also filtered all positional and PCR duplicates. Quality control: several quality metrics for all replicate experiments of each data set were computed. In brief, these metrics measure ChIP enrichment, signal-to-noise ratios, sequencing depth, library complexity and reproducibility of peak calling8. ChIP-seq that did not pass the minimum quality control thresholds were discarded and not used in any analyses. Peak calling: all ChIP-seq experiments were scored against an appropriate control designated by the production groups (either input DNA or DNA obtained from a control immunoprecipitation). We used the SPP peak caller27 to identify and score (rank) potential occupancy sites/peaks. For obtaining optimal thresholds, we used the irreproducible discovery rate (IDR) framework to determine high confidence occupancy events by leveraging the reproducibility and rank consistency of peak identifications across replicate experiments of a data set. Code and detailed step-by-step instructions to call peaks using the IDR framework are available at: https://sites.google.com/site/anshulkundaje/projects/idr. Black list: all peak sets were then screened against specially curated empirical blacklists for each species (A.P.B. and A.K., manuscript submitted). In brief, these blacklist regions typically show the following characteristics: unstructured and extreme high signal in sequenced input DNA and control data sets as well as open chromatin data sets irrespective of cell type identity; an extreme ratio of multi-mapping to unique mapping reads from sequencing experiments; overlap with specific types of repeat regions such as centromeric, telomeric and satellite repeats that often have few unique mappable locations interspersed in repeats. The human blacklist can be found from: http://hgdownload.cse.ucsc.edu/goldenPath/hg19/encodeDCC/wgEncodeMapability/wgEncodeDacMapabilityConsensusExcludable.bed.gz. The mouse blacklist can be downloaded from: http://www.broadinstitute.org/∼anshul/projects/mouse/blacklist/mm9-blacklist.bed.gz. In this study, the blacklist filtered IDR binding peaks for the same TF using the same cell line generated by different institutes were merged. All the raw read files, mapped files and peak files in mouse are deposited in http://mouseencode.org. The human data can be accessed in https://www.encodeproject.org. The access ID in each experiment can be found in Supplementary Table 2.
Motif finding
To compare mouse and human regulatory networks, we applied the de novo motif discovery approach that we developed previously28 and obtained a list of high-confidence sequence motifs using the ChIP-seq data sets. For each ChIP-seq data set, our computational pipeline reported up to five significant motifs. Typically, one of the motifs is the canonical motif of the TF, reflecting its DNA-binding specificity, and we call this the primary motif. If the TF does not have a DNA binding domain, we define the strongest motif as its primary motif. We call the remaining motifs secondary motifs. When the primary motifs of a pair of orthologous TFs are compared, they are either ‘conserved’ or ‘not conserved’ on the basis of whether the similarity between them passes the cut off (1.0 × 10−5). Because a TF may have several secondary motifs, the secondary motifs of two orthologous TFs are ‘partly conserved’ if a subset, but not all, of the motifs are conserved. When neither the human TF nor the mouse TF has a secondary motif, we assign the situation as motif ‘not available’.
ChromHMM
ChromHMM29 was applied on the ChIP-seq data of five histone modifications to learn a multivariate HMM model for segmentation of mapped genome in each cell type. Specifically, the ChIP-seq mapped reads were first pooled from replicates for each of the five histone modifications (H3K4me3, H3K4me1, H3K36me3, H3K27ac and H3K27me3). These mapped reads were first processed by ChromHMM into binarized data in every 200-bp window over the entire mapped genome, with ChIP ‘input’ reads as the background control. To learn the model jointly from mouse and human, a pseudo genome table was first constructed by concatenating the mouse mm9 and human hg19 table, then the model was learned from the binarized data in all four cell lines, giving a single model with a common set of emission parameters and transition parameters, which was then used to produce segmentations in all cell types based on the most likely state assignment of the model. We tried models with up to 20 states and selected an eight-state-model as it appeared most parsimonious in the sense that all eight states had clearly distinct emission properties, while the interpretability of distinction between states in models with additional states was less clear.
MeDIP-seq and MRE-seq
MeDIP-seq and MRE-seq experiments were performed as previously described16. The reads were aligned to hg19 and mm9 using BWA. MRE-seq reads were further normalized for difference in enzyme efficiency.
Defining different genomic locations
TSSs were defined by ENCOCDE consortium15. Promoter regions were defined as 2 kb upstream and downstream of the TSS. Distal regions were defined as 10 kb away from TSS. The rest of the genomic regions were defined as middle regions. All the three genomic locations are exclusive to each other, and the priority during the definition is promoter, distal and middle. Each TF OS was assigned to one (and only one) genomic location. If TF OSs overlapped with several regions, the centre of the OS was used to define which region to assign.
TF OSs sequence
phyloP30 wiggle track were downloaded from the UCSC browser. Specifically, hg19 phyloP46way track was used for human and mm9 phyloP30way track was used for mouse. This average phyloP score were calculated at one base pair resolution in 200-bp regions centred on the summit of TF peaks.
Mapping reciprocal orthologous sequences between human and mouse
Orthologous DNA sequences between human and mouse were mapped by bnMapper (O. Denas, R. Sandstrom and J. Taylor, manuscript submitted) using reciprocal chain with default setting (bnMapper.py -f BED12).
RegulomeDB SNV and occupancy conservation
SNPs assigned with pre-calculated regulatory potentials were downloaded from: http://www.regulomedb.org/downloads. dbSNP138 was downloaded from the UCSC genome browser. TF OSs were divided into two exclusive groups: occupancy-conserved and human-specific. The number of SNPs with high regulatory potential and the number of dbSNPs located in each group of TF OSs were calculated. Fisher’s exact test was conducted to examine the enrichment of SNPs with high regulatory potential in each group.
GWAS SNPs and occupancy conservation
GWAS catalogue file was downloaded from: http://www.genome.gov/admin/gwascatalog.txt. Lead SNPs that overlapped with exons were removed. For each lead SNP, if either the SNP itself or the linkage disequilibrium SNPs are located within a given TF OS, it was assigned to that TF OS. Lead SNPs that can be assigned to several TF OSs were also removed. Two-sided Fisher’s exact tests were conduct to calculate the enrichment of conservation in each given phenotype compared with the distribution of all dbSNPs, and P values were further adjusted by Benjamini–Hochberg procedure.
References
Odom, D. T. et al. Tissue-specific transcriptional regulation has diverged significantly between human and mouse. Nature Genet. 39, 730–732 (2007)
Schmidt, D. et al. Five-vertebrate ChIP-seq reveals the evolutionary dynamics of transcription factor binding. Science 328, 1036–1040 (2010)
Stefflova, K. et al. Cooperativity and rapid evolution of cobound transcription factors in closely related mammals. Cell 154, 530–540 (2013)
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nature Genet. 42, 631–634 (2010)
Borneman, A. R. et al. Divergence of transcription factor binding sites across related yeast species. Science 317, 815–819 (2007)
Pennacchio, L. A. & Rubin, E. M. Genomic strategies to identify mammalian regulatory sequences. Nature Rev. Genet. 2, 100 (2001)
He, Q. et al. High conservation of transcription factor binding and evidence for combinatorial regulation across six Drosophila species. Nature Genet. 43, 414–420 (2011)
Landt, S. G. et al. ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813 (2012)
Tokugawa, Y., Koyama, M. & Silver, J. A molecular basis for species differences in Thy-1 expression patterns. Mol. Immunol. 34, 1263 (1997)
Mestas, J. & Hughes, C. C. W. Of mice and not men: differences between mouse and human immunology. J. Immunol. 172, 2731–2738 (2004)
Nitzsche, A. et al. RAD21 cooperates with pluripotency transcription factors in the maintenance of embryonic stem cell identity. PLoS ONE 6, e19470 (2011)
Merkenschlager, M. & Odom, D. T. CTCF and cohesin: linking gene regulatory elements with their targets. Cell 152, 1285–1297 (2013)
Sawado, T., Igarashi, K. & Groudine, M. Activation of β-major globin gene transcription is associated with recruitment of NF-E2 to the β-globin LCR and gene promoter. Proc. Natl Acad. Sci. USA 98, 10226 (2001)
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012)
Yue, F. et al. A comparative encyclopedia of DNA elements in the mouse genome. Nature http://dx.doi.org/10.1038/nature13992 (this issue)
Xie, M. et al. DNA hypomethylation within specific transposable element families associates with tissue-specific enhancer landscape. Nature Genet. 45, 836–841 (2013)
Sundaram, V., Cheng, Y., Snyder, M. P. & Wang, T. Widespread contribution of transposable elements to the innovation of gene regulatory networks. Genome Res. http://dx.doi.org/10.1101/gr.168872.113 (15 October 2014)
Schmidt, D. et al. Waves of retrotransposon expansion remodel genome organization and CTCF binding in multiple mammalian lineages. Cell 148, 335–348 (2012)
Gross, D. S. & Garrard, W. T. Nuclease hypersensitive sites in chromatin. Annu. Rev. Biochem. 57, 159–197 (1988)
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nature Genet. 39, 311–318 (2007)
Visel, A., Minovitsky, S., Dubchak, I. & Pennacchio, L. A. VISTA Enhancer Browser–a database of tissue-specific human enhancers. Nucleic Acids Res. 35, D88–D92 (2007)
Visel, A. et al. ChIP-seq accurately predicts tissue-specific activity of enhancers. Nature 457, 854–858 (2009)
Kadamb, R., Mittal, S., Bansal, N., Batra, H. & Saluja, D. Sin3: insight into its transcription regulatory functions. Eur. J. Cell Biol. 92, 237–246 (2013)
Boyle, A. P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012)
Kasowski, M. et al. Variation in transcription factor binding among humans. Science 328, 232–235 (2010)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler Transform. Bioinformatics 25, 1754–1760 (2009)
Kharchenko, P. V., Tolstorukov, M. Y. & Park, P. J. Design and analysis of ChIP-seq experiments for DNA-binding proteins. Nature Biotechnol. 26, 1351–1359 (2008)
Wang, J. et al. Sequence features and chromatin structure around the genomic regions bound by 119 human transcription factors. Genome Res. 22, 1798–1812 (2012)
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nature Methods 9, 215–216 (2012)
Cooper, G. M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005)
Acknowledgements
This work is funded by grants 3RC2HG005602, 5U54HG006996 and 1U54HG00699 (M.P.S.), and R01DK065806 and RC2HG005573 (R.C.H.). A.V. and L.A.P. were supported by National Human Genome Research Institute (NHGRI) grant R01HG003988, U54HG006997 and supplementary funds provided by the American Recovery and Reinvestment Act. The in vivo enhancer activity assays were conducted at the E.O. Lawrence Berkeley National Laboratory and performed under Department of Energy Contract DE-AC02-05CH11231, University of California. We acknowledge R. M. Myers for providing access to ChIP-seq data in human embryonic cells. Illumina sequencing services were performed by the Stanford Center for Genomics and Personalized Medicine.
Author information
Authors and Affiliations
Contributions
Y.C., B.-H.K., A.P.B., W.W., J.L. and Z.M. analysed the data. Z.M., Y.C., P.C., X.Y., D.P., G.E., T.K., C.A.K. and B.G. prepared and pre-processed ChIP-seq data. V.S. and X.X. prepared and pre-processed MRE-seq and MEDIP-seq data. A.V. and N.D. conducted the enhancer assay. Y.C., Z.M., R.C.H., M.P.S., K.A., T.W., L.A.P., Z.W., S.L. and Y.L. wrote the paper with input from all authors. M.P.S. and R.C.H. coordinated and supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 TF ChIP-seq data overview and analysis workflow.
a, All TFs in this study are grouped according to species and cell types. TF DNA binding domains are list in the second column. The TFs without binding domains are highlighted in grey. The TFs assayed were cross-marked, whereas TFs not assayed are depicted in white. b, Flowchart for the analysis pipeline for inter- and intra-species comparisons.
Extended Data Figure 2 TF OSs distribution and motifs.
a, An illustration of TF OS distribution relative to TSSs in MEL and K562 cells. Each row represents one TF, each column represents one genomic region. Heat-map colour shows the proportions of TF OSs that are located in different genomic regions. b, Similar TF OS distribution plot as a in CH12 and GM12878 cells. c, Correlation between mouse and human TF OS distribution. Dot plot shows the correlation of orthologous TF OS distribution in each genomic region. Each dot represents proportion of OSs for one TF in one genomic region. The x axis is the proportion in mouse genome, and the y axis is the proportion in human genome. d, Motif comparison for sequence specific TFs examined in erythroid progenitor cells (MEL and K562). Each row represents one TF. The level of motif conservation is encoded by colour.
Extended Data Figure 3 TF OS chromatin states and DNA methylation status preference comparison.
a, Emission matrix of ChromHMM trained by five histone modification markers (H3K4me1, H3K4me3, H3K36me3, H3K27me3 and H3K27ac). b, Heat map shows the proportion of TF OSs (rows) that overlap with each chromatin state (columns) generated by ChromHMM using five different histone markers in CH12 and GM12878 cells. c, The average signal distributions for MeDIP-seq and MRE-seq in CH12 and GM12878 cells. The 5-kb flanking regions centred on the TF OS peak summits were divided into 50-bp bins. Signals were aggregated in each bin.
Extended Data Figure 4 Proportion of predicted enhancers in the orthologous TF OSs.
Bar graphs show the proportions of TF OSs that overlapped with the predicted enhancers. a, Results in MEL and K562 cells. b, Results in CH12 and GM12878 cells. The x axis represents different TFs, the y axis represents the proportion of TF OSs that overlapped with predicted enhancers.
Extended Data Figure 5 Occupancy conservation adjusted by sequence conservation.
a, The heat map represents the adjusted occupancy conservation of TF (row) OSs in the four cell lines. The colour intensity represents the proportion of TF OSs that are occupancy-conserved between mouse and human in different genomic regions (column). To remove the bias introduced by variation of sequence conservation at different genomic loci, only TF OSs in which the sequence can be aligned between mouse and human were included in this analysis. b, The heat map is similar to Fig. 2b. TFs showing remarkable difference on total binding peaks numbers between the mouse and human were excluded.
Extended Data Figure 6 Comparison of the epigenetic features between TF OSs and orthologous sequences.
a, The y axis represents the proportion of TF OSs in each chromatin state. TF OSs that can be aligned between mouse and human are divided into two categories according to the occupancy conservation status. Each panel represents distribution of TF OSs in one cell line. b, Each panel represents mouse TF OSs in one chromatin state. The pie chart in each panel shows the proportions of chromatin states in the orthologous sequence in human. Panels in the left column represent the occupancy-conserved TF OSs, and panels in the right column represent the TF OSs that can be aligned but without occupancy conservation. c, The y axis represents the normalized DNA methylation signals (MeDIP-seq). TF OSs that can be aligned between mouse and human are divided into two categories according to the occupancy conservation status (both sequence and occupancy are conserved (OCC) and sequence is conserved but occupancy is not conserved (SCNC)). Each panel represents distribution in one cell line.
Extended Data Figure 7 Conservation of occupancy is associated with chromatin accessibility and enhancer activity in several tissues.
a, Association between occupancy conservation and chromatin accessibility across several tissues. The density plot represents the frequency that TF OSs (removed DNA sequences occupied by CTCF, RAD21 and SMC3) are in accessible chromatin in varying numbers of cell types. The x axis is the Shannon index calculated based on the DHS signals in 55 mouse tissues or cell lines; high values mean the TF OS is in accessible chromatin in many cell types. The red line shows the fraction of TF OSs at which occupancy is conserved within each bin of the Shannon index. b, c, The association between occupancy conservation and chromatin accessibility across multiple tissues for each TF (row) in CH12 and MEL cells. TF OSs are divided into different bins according to the value of the Shannon index (columns). The colour intensity represents the proportion of occupancy-conserved TF OSs within each bin. d, e, Similar distribution to b and c but only for TF OSs that are located 2 kb away from TSSs.
Extended Data Figure 8 Consistency of observations between embryonic stem cells and cell lines.
a, Genomic distribution of five TF OSs in embryonic stem cells. b, Occupancy conservation in different genomic locations between human and mouse embryonic stem cells. c, Occupancy conservation of TF OSs in embryonic stem cells is associated with function in many tissues.
Extended Data Figure 9 Relationship between occupancy conservation and pair-wised TFs co-association.
a–d, Occupancy conservation and TF co-association analysis was conducted as described in Fig. 4c for all four cell lines. The TFs were kept in the same order across the four cell lines for easy visualization.
Supplementary information
Supplementary Information
This file contains a full list of authors and affiliations, Supplementary Methods and Analysis, Supplementary Figures 1-4 and Supplementary References. (PDF 5810 kb)
Supplementary Data
This file contains Supplementary Table 1 (XLSX 49 kb)
Supplementary Data
This file contains Supplementary Table 2 (XLSX 35 kb)
Supplementary Data
This file contains Supplementary Table 3 (XLSX 48 kb)
Supplementary Data
This file contains Supplementary Table 4 (XLSX 95 kb)
Supplementary Data
This file contains Supplementary Table 5 (XLSX 66 kb)
Supplementary Data
This file contains Supplementary Table 6 (XLSX 1063 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported licence. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Cheng, Y., Ma, Z., Kim, BH. et al. Principles of regulatory information conservation between mouse and human. Nature 515, 371–375 (2014). https://doi.org/10.1038/nature13985
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13985
This article is cited by
-
Disrupted chromatin architecture in olfactory sensory neurons: looking for the link from COVID-19 infection to anosmia
Scientific Reports (2023)
-
Signaling mechanisms in renal compensatory hypertrophy revealed by multi-omics
Nature Communications (2023)
-
Microbiota in health and diseases
Signal Transduction and Targeted Therapy (2022)
-
LINE retrotransposons characterize mammalian tissue-specific and evolutionarily dynamic regulatory regions
Genome Biology (2021)
-
INFIMA leverages multi-omics model organism data to identify effector genes of human GWAS variants
Genome Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.