Abstract
Human cancers, including breast cancers, comprise clones differing in mutation content. Clones evolve dynamically in space and time following principles of Darwinian evolution1,2, underpinning important emergent features such as drug resistance and metastasis3,4,5,6,7. Human breast cancer xenoengraftment is used as a means of capturing and studying tumour biology, and breast tumour xenografts are generally assumed to be reasonable models of the originating tumours8,9,10. However, the consequences and reproducibility of engraftment and propagation on the genomic clonal architecture of tumours have not been systematically examined at single-cell resolution. Here we show, using deep-genome and single-cell sequencing methods, the clonal dynamics of initial engraftment and subsequent serial propagation of primary and metastatic human breast cancers in immunodeficient mice. In all 15 cases examined, clonal selection on engraftment was observed in both primary and metastatic breast tumours, varying in degree from extreme selective engraftment of minor (<5% of starting population) clones to moderate, polyclonal engraftment. Furthermore, ongoing clonal dynamics during serial passaging is a feature of tumours experiencing modest initial selection. Through single-cell sequencing, we show that major mutation clusters estimated from tumour population sequencing relate predictably to the most abundant clonal genotypes, even in clonally complex and rapidly evolving cases. Finally, we show that similar clonal expansion patterns can emerge in independent grafts of the same starting tumour population, indicating that genomic aberrations can be reproducible determinants of evolutionary trajectories. Our results show that measurement of genomically defined clonal population dynamics will be highly informative for functional studies using patient-derived breast cancer xenoengraftment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Data deposits
Genome data have been deposited at the European Genome-phenome Archive (http://www.ebi.ac.uk/ega) under accession number EGAS00001000952. Processed data can be viewed at http://www.cbioportal.org.
References
Aparicio, S. & Caldas, C. The implications of clonal genome evolution for cancer medicine. N. Engl. J. Med. 368, 842–851 (2013)
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976)
Diaz, L. A., Jr et al. The molecular evolution of acquired resistance to targeted EGFR blockade in colorectal cancers. Nature 486, 537–540 (2012)
Shah, S. P. et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486, 395–399 (2012)
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012)
Campbell, P. J. et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 467, 1109–1113 (2010)
Bashashati, A. et al. Distinct evolutionary trajectories of primary high-grade serous ovarian cancers revealed through spatial mutational profiling. J. Pathol. 231, 21–34 (2013)
DeRose, Y. S. et al. Tumor grafts derived from women with breast cancer authentically reflect tumor pathology, growth, metastasis and disease outcomes. Nature Med. 17, 1514–1520 (2011)
Zhang, X. et al. A renewable tissue resource of phenotypically stable, biologically and ethnically diverse, patient-derived human breast cancer xenograft models. Cancer Res. 73, 4885–4897 (2013)
Ding, L. et al. Genome remodelling in a basal-like breast cancer metastasis and xenograft. Nature 464, 999–1005 (2010)
Pearson, T. et al. Non-obese diabetic-recombination activating gene-1 (NOD–Rag1null) interleukin (IL)-2 receptor common gamma chain (IL2rγnull) null mice: a radioresistant model for human lymphohaematopoietic engraftment. Clin. Exp. Immunol. 154, 270–284 (2008)
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012)
Ha, G. et al. Integrative analysis of genome-wide loss of heterozygosity and monoallelic expression at nucleotide resolution reveals disrupted pathways in triple-negative breast cancer. Genome Res. 22, 1995–2007 (2012)
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012)
Ellis, M. J. et al. Whole-genome analysis informs breast cancer response to aromatase inhibition. Nature 486, 353–360 (2012)
Banerji, S. et al. Sequence analysis of mutations and translocations across breast cancer subtypes. Nature 486, 405–409 (2012)
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012)
Roth, A. et al. PyClone: statistical inference of clonal population structure in cancer. Nature Methods 11, 396–398 (2014)
Ha, G. et al. Titan: inference of copy number architectures in clonal cell populations from tumor whole genome sequence data. 24, 1881–1893 (2014)
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012)
Kreso, A. et al. Variable clonal repopulation dynamics influence chemotherapy response in colorectal cancer. Science 339, 543–548 (2013)
Nolan-Stevaux, O. et al. Measurement of cancer cell growth heterogeneity through lentiviral barcoding identifies clonal dominance as a characteristic of in vivo tumor engraftment. PLoS ONE 8, e67316 (2013)
Li, S. et al. Endocrine-therapy-resistant ESR1 variants revealed by genomic characterization of breast-cancer-derived xenografts. Cell Rep. 4, 1116–1130 (2013)
Notta, F. et al. Evolution of human BCR-ABL1 lymphoblastic leukaemia-initiating cells. Nature 469, 362–367 (2011)
Clappier, E. et al. Clonal selection in xenografted human T cell acute lymphoblastic leukemia recapitulates gain of malignancy at relapse. J. Exp. Med. 208, 653–661 (2011)
Acknowledgements
We are grateful to the staff of the CTAG Molecular Pathology facility, members of the Library Technical Development, Library Construction, Sequencing and Bioinformatics teams at the Michael Smith Genome Sciences Centre for technical assistance with data generation, and S. Kalloger for assistance with sample collection. S.A. and S.P.S. are supported by Canada Research Chairs. P.E. is supported by a Michael Smith Foundation for Health Research (MSFHR) Fellowship. A.S. is supported by an NSERC CREATE scholarship through the graduate program in Genome Science and Technology at UBC. S.P.S. is a MSFHR scholar. We acknowledge long-term funding support provided by the BC Cancer Foundation. The S.A., S.P.S. and C.H. groups receive operating funds from the Canadian Breast Cancer Foundation, Canadian Cancer Society Research Institute, Terry Fox Research Institute, Genome Canada and Canadian Institutes for Health Research (CIHR). We thank S. Mullaly for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
S.A. and S.P.S. designed the study and supervised the research. S.A., S.P.S., P.E. and A.S. wrote the paper. P.E., A.Br., J.S., T.A., W.G., H.C., H.X., L.N., Y.W. and D.L. performed transplants and passaging. K.G., S.C. and C.M. recruited patients and performed tissue biopsies. A.S., P.E., G.H., C.N., H.F., A.J.L.R., C.L., A.Ba., C.S., K.S., J.R., R.A., A.M., C.d.S., S.P.S. and S.A. carried out bioinformatics analyses. J.K., D.Y., E.L., J.Br., A.W., J.Bi., K.L., A.J.M., A.O., R.M., Y.Z., C.H. and M.A.M. assisted with sequence generations and single-cell experiments. T.O., J.L. and D.H. contributed to histological analysis. C.J.E., C.H., M.A.M., C.C., S.P.S. and S.A. provided intellectual contributions to design or interpretation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
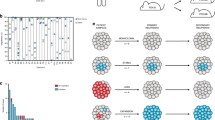
Extended Data Figure 1 Transplant history.
Diagrams show the transplant history of each xenograft line. Line segment colours represent the site used for each transplant (blue, subcutaneous; red, subrenal capsule; green, mammary fat pad). Black points indicate the passage of an engrafted xenograft to the next mouse generation. Grey crosses indicate transplants that did not result in palpable tumours. Samples analysed by whole-genome and/or targeted deep sequencing are indicated (black squares and vertical lines, respectively). The cumulative time in vivo is shown on the x axis. The originating tumour site (Met., pleural effusion; Prim., primary breast) and immunohistochemical expression of biomarkers (ER, oestrogen receptor; PR, progesterone receptor; TN, triple negative for ER, PR and HER2) are shown.
Extended Data Figure 2 Comparison of the prevalence of mutations in six originating tumours and subsequent xenografts in SNV and CNA spaces.
a, Density scatter plots showing the WGSS variant allele prevalence of genome-wide high-confidence SNVs in tumours (x axis) and xenografts (y axis). SNVs in clones undergoing neutral dynamics lie along a diagonal, and SNVs in clones undergoing expansion or contraction lie on/towards the y and x axes, respectively. b, Scatter plots showing the mutation cellular prevalence of selected SNVs in tumours and xenografts, inferred by PyClone from population-level targeted deep sequencing. Circles represent individual SNVs, colours indicate clusters of mutations for which mutation cellular prevalences vary together over all sample time points. c, Scatter plots show co-occurrence of CNA/LOH events inferred by TITAN in tumours and xenografts. The z axis height of each bar shows the number of genes belonging to a unique mutation cluster and present at the indicated mutation cellular prevalence in tumour (x axis) and xenograft (y axis).
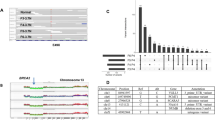
Extended Data Figure 3 Single-cell determination of clonal genotypes recapitulates population-based prediction of minor clone selection.
DNA prepared from 62 individual lysed SA494 tumour and 58 passage 4 lysed xenograft nuclei was amplified in single reactions using a panel of multiplexed PCR primer pairs targeting amplicons containing 40 SNV and 7 germline variants, and the variant allele ratios were determined by targeted deep sequencing. a, Mutation clusters inferred by the PyClone model from bulk population measurements. b, Bayesian phylogenetic tree derived from multi-locus genotypes of individual nuclei. The tumour and xenograft nuclei group in distinct clades. c, Heat map depicts the multi-locus variant allele prevalences (blue/yellow/red corresponds to wild-type/heterozygous/homozygous loci, respectively) at variant positions (horizontal axis) in individual nuclei (vertical axis, ordered by phylogenetic grouping in b). Upper two blocks show gDNA controls and normal cell nuclei present in tumour samples. The PyClone mutation cluster corresponding to each SNV is indicated by colour in the lowermost horizontal bar. d, Consensus genotypes derived from high-probability splits in the phylogenetic tree confirm a set of high prevalence tumour-specific and xenograft-specific mutations, consistent with the expansion of a minor originating clone to dominance in the xenograft, as well as mutations shared in tumour and xenograft nuclei.
Extended Data Figure 4 Clonal dynamics are reproduced in replicate transplants.
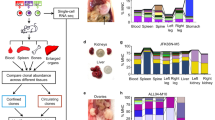
a–c, Upper panels, passaging history of SA532, SA429 and SA496, showing transplants that resulted in successful xenografts. The transplant sites (blue, subcutaneous; red, subrenal; green, mammary fat pad; all subcutaneous for SA532) and host mouse strains (blue, NSG; orange, NRG; all NSG for SA429 and SA496) are shown. Boxed nodes indicate passages analysed by WGSS. Lower panels, change in cellular prevalence of mutation clusters over individual transplants. Plots correspond to passages in upper panels. The clusters are inferred by PyClone using grouped data from all passages and correspond to those displayed in Fig. 1. Arrows in SA429 and SA532 show examples of parallel clonal dynamics of the same mutation cluster in multiple replicate transplants. SA496 exhibits less replicated evolution compared with other cases.
Extended Data Figure 5 Correlation of clonal dynamics in replicate transplants of SA429, SA501, SA532, SA496 and SA535.
a–e, Scatter plots display the inferred mutation cellular prevalence of all SNVs in pairs of same-passage replicates, for cases SA429, SA501, SA532, SA496 and SA535, respectively. For each replicate, prevalences are inferred by a separate PyClone analysis that excludes data from other same-passage transplants. Colours indicate mutation clusters inferred in each individual PyClone analyses; the SNVs clustered and colours assigned may differ in each plot. The Pearson correlation coefficients are shown, indicating closely related evolution in most pairs.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary References, legends for Supplementary Figures 1-15 and Tables 1-15 (see separate files for Supplementary Tables and Figures). (PDF 248 kb)
Supplementary Figures
This file contains Supplementary Figures 1-15 – see Supplementary Information document for legends. (PDF 45314 kb)
Supplementary Tables
This zipped file contains Supplementary Tables 1-10 – see Supplementary Information document for legends. (ZIP 7130 kb)
Supplementary Data
This file contains the script to carry out credible interval significance testing. (ZIP 35 kb)
Rights and permissions
About this article
Cite this article
Eirew, P., Steif, A., Khattra, J. et al. Dynamics of genomic clones in breast cancer patient xenografts at single-cell resolution. Nature 518, 422–426 (2015). https://doi.org/10.1038/nature13952
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13952
This article is cited by
-
Personalizing non-small cell lung cancer treatment through patient-derived xenograft models: preclinical and clinical factors for consideration
Clinical and Translational Oncology (2024)
-
Establishment of a high-fidelity patient-derived xenograft model for cervical cancer enables the evaluation of patient’s response to conventional and novel therapies
Journal of Translational Medicine (2023)
-
Clonal dynamics limits detection of selection in tumour xenograft CRISPR/Cas9 screens
Cancer Gene Therapy (2023)
-
Biology, vulnerabilities and clinical applications of circulating tumour cells
Nature Reviews Cancer (2023)
-
Predictors of success in establishing orthotopic patient-derived xenograft models of triple negative breast cancer
npj Breast Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.