Abstract
Sponges are simple animals with few cell types, but their genomes paradoxically contain a wide variety of developmental transcription factors1,2,3,4, including homeobox genes belonging to the Antennapedia (ANTP) class5,6, which in bilaterians encompass Hox, ParaHox and NK genes. In the genome of the demosponge Amphimedon queenslandica, no Hox or ParaHox genes are present, but NK genes are linked in a tight cluster similar to the NK clusters of bilaterians5. It has been proposed that Hox and ParaHox genes originated from NK cluster genes after divergence of sponges from the lineage leading to cnidarians and bilaterians5,7. On the other hand, synteny analysis lends support to the notion that the absence of Hox and ParaHox genes in Amphimedon is a result of secondary loss (the ghost locus hypothesis)8. Here we analysed complete suites of ANTP-class homeoboxes in two calcareous sponges, Sycon ciliatum and Leucosolenia complicata. Our phylogenetic analyses demonstrate that these calcisponges possess orthologues of bilaterian NK genes (Hex, Hmx and Msx), a varying number of additional NK genes and one ParaHox gene, Cdx. Despite the generation of scaffolds spanning multiple genes, we find no evidence of clustering of Sycon NK genes. All Sycon ANTP-class genes are developmentally expressed, with patterns suggesting their involvement in cell type specification in embryos and adults, metamorphosis and body plan patterning. These results demonstrate that ParaHox genes predate the origin of sponges, thus confirming the ghost locus hypothesis8, and highlight the need to analyse the genomes of multiple sponge lineages to obtain a complete picture of the ancestral composition of the first animal genome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
European Nucleotide Archive
Data deposits
Genome assembly of S. ciliatum and the coding sequences and their translations from transcriptome assemblies of S. ciliatum and L. complicata used in this study can be accessed through http://compagen.zoologie.uni-kiel.de/ and are also deposited at http://dx.doi.org/10.5061/dryad.tn0f3. RNA-seq data have been deposited in ArrayExpress under accession numbers E-MTAB-2430, E-MTAB-2431 and E-MTAB-2890, and the cloned coding sequences of S. ciliatum ANTP-class genes have been deposited in the European Nucleotide Archive under accession numbers LN609546 to LN609553.
References
Srivastava, M. et al. The Amphimedon queenslandica genome and the evolution of animal complexity. Nature 466, 720–726 (2010)
Fortunato, S. et al. Genome-wide analysis of the sox family in the calcareous sponge Sycon ciliatum: multiple genes with unique expression patterns. EvoDevo 3, 14 (2012)
Larroux, C. et al. Genesis and expansion of metazoan transcription factor gene classes. Mol. Biol. Evol. 25, 980–996 (2008)
Fortunato, S. A., Leininger, S. & Adamska, M. Evolution of the Pax-Six-Eya-Dach network: the calcisponge case study. EvoDevo 5, 23 (2014)
Larroux, C. et al. The NK homeobox gene cluster predates the origin of Hox genes. Curr. Biol. 17, 706–710 (2007)
Fahey, B., Larroux, C., Woodcroft, B. J. & Degnan, B. M. Does the high gene density in the sponge NK homeobox gene cluster reflect limited regulatory capacity? Biol. Bull. 214, 205–217 (2008)
Ryan, J. F. et al. The homeodomain complement of the ctenophore Mnemiopsis leidyi suggests that Ctenophora and Porifera diverged prior to the ParaHoxozoa. EvoDevo 1, 9 (2010)
Mendivil Ramos, O., Barker, D. & Ferrier, D. E. K. Ghost loci imply Hox and ParaHox existence in the last common ancestor of animals. Curr. Biol. 22, 1951–1956 (2012)
Wörheide, G. et al. in Advances in Marine Biology Vol. 61 (eds Uriz, M. J., Maldonado, M., Becerro, M. A. & Xavier, T. ) 1–78 (Academic, 2012)
Peterson, K. J. & Sperling, E. A. Poriferan ANTP genes: primitively simple or secondarily reduced? Evol. Dev. 9, 405–408 (2007)
Ferrier, D. E. K. in Hox Genes: Studies from the 20th to the 21st Century Vol. 689 (ed. Deutsch, J. S. ) 91–100 (Springer, 2010)
Garcia-Fernàndez, J. The genesis and evolution of homeobox gene clusters. Nature Rev. Genet. 6, 881–892 (2005)
Chourrout, D. et al. Minimal ProtoHox cluster inferred from bilaterian and cnidarian Hox complements. Nature 442, 684–687 (2006)
Leininger, S. et al. Developmental gene expression provides clues to relationships between sponge and eumetazoan body plans. Nature Commun. 5, 3905 (2014)
Luke, G. N. et al. Dispersal of NK homeobox gene clusters in amphioxus and humans. Proc. Natl Acad. Sci. USA 100, 5292–5295 (2003)
Hui, J. H. L. et al. Extensive chordate and annelid macrosynteny reveals ancestral homeobox gene organization. Mol. Biol. Evol. 29, 157–165 (2012)
Hui, J. H. L., Holland, P. W. H. & Ferrier, D. E. K. Do cnidarians have a ParaHox cluster? Analysis of synteny around a Nematostella homeobox gene cluster. Evol. Dev. 10, 725–730 (2008)
Seo, H.-C. et al. Hox cluster disintegration with persistent anteroposterior order of expression in Oikopleura dioica. Nature 431, 67–71 (2004)
Saudemont, A. et al. Complementary striped expression patterns of NK homeobox genes during segment formation in the annelid Platynereis. Dev. Biol. 317, 430–443 (2008)
Amano, S. & Hori, I. Metamorphosis of calcareous sponges. 2. Cell rearrangement and differentiation in metamorphosis. Invertebr. Reprod. Dev. 24, 13–26 (1993)
Voigt, C., Adamski, M., Sluzek, M. & Adamska, M. Calcareous sponge genomes reveal complex evolution of α-carbonic anhydrases and two key biomineralization enzymes. BMC Evol. Biol (in the press)
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010)
Li, B. & Dewey, C. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011)
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience 1, 18 (2012)
Boetzer, M., Henkel, C., Jansen, H., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27, 578–579 (2011)
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nature Biotechnol. 29, 644–652 (2011)
Takatori, N. et al. Comprehensive survey and classification of homeobox genes in the genome of amphioxus, Branchiostoma floridae. Dev. Genes Evol. 218, 579–590 (2008)
Zhong, Y.-F. & Holland, P. HomeoDB2: functional expansion of a comparative homeobox gene database for evolutionary developmental biology. Evol. Dev. 13, 567–568 (2011)
Darriba, D., Taboada, G., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165 (2011)
Keane, T. M., Creevey, C., Pentony, M., Naughton, T. & Mclnerney, J. Assessment of methods for amino acid matrix selection and their use on empirical data shows that ad hoc assumptions for choice of matrix are not justified. BMC Evol. Biol. 6, 29 (2006)
Acknowledgements
This study was funded by the Sars Centre core budget to M. Adamska. Sequencing was performed at the Norwegian High Throughput Sequencing Centre funded by the Norwegian Research Council. O.M.R. and D.E.K.F. acknowledge support from the BBSRC and the School of Biology, University of St Andrews. We thank B. Bergum for collecting samples in the 2011 season and R. Holdhus from the Genomics Core Facility at the University of Bergen for help with DNA shearing.
Author information
Authors and Affiliations
Contributions
S.A.V.F. carried out gene identification and cloning, analysed gene expression by in situ hybridization, and participated in phylogenetic analyses and manuscript writing. M. Adamski performed sequence assembly, annotation, quantification of gene expression and participated in sample collection, phylogenetic analyses and manuscript writing. O.M.R. performed the synteny analyses, participated in phylogenetic analyses, manuscript writing and design of the research approach for synteny and phylogenetic analyses. S.L. isolated samples for sequencing of genomes, generated Mate Pair libraries, and participated in sample collection. J.L. generated samples for sequencing of S. ciliatum metamorphosis stages. D.E.K.F. participated in design of the research approach for synteny and phylogenetic analyses and writing of the manuscript. M. Adamska conceived the study, and participated in data analysis, sample collection and writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
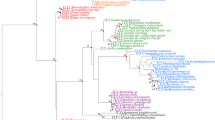
Extended Data Figure 1 Phylogenetic tree of the ANTP-class homeodomains including representative bilaterian and non-bilaterian sequences.
A neighbour-joining tree using the JTT+G (0.5) (1,000 bootstraps) model of protein evolution is displayed. A combination of three support values obtained for three phylogenetic methods is shown: black value is the bootstrap support from the neighbour-joining method, blue value is the bootstrap support from the maximum-likelihood method (LG+G 0.5), and red value is posterior probability from Bayesian analysis (LG+G 0.5). Bootstrap support values below 10% and posterior probability values below 0.5 are not shown except for associations of calcisponge sequences. The root was determined by using selected PRD-class homeodomains as an outgroup. Aqu, A. queenslandica (Porifera/demosponges); Bfl, B. floridae (Chordata); Lco, L. complicata; Mle, M. leidyi (Ctenophora); Nve, N. vectensis (Cnidaria); Sci, S. ciliatum (Porifera/calcisponges); Tad, Trichoplax adhaerens (Placozoa); Tca, T. castaneum (Arthropoda). Scale bar indicates the number of amino acid substitutions per site.
Extended Data Figure 2 Variability of the YIT/YIS homeodomain motif within the Cdx/Cad, En and Dbx families in bilaterians, cnidarians, a placozoan and sponges.
Ame, Apis mellifera; Aqu, A. queenslandica; Bfl, B. floridae; Cte, Capitella teleta; Dme, Drosophila melanogaster; Dre, Danio rerio; Edi, Eleutheria dichotoma; Gga, Gallus gallus; Hsa, Homo sapiens; Lgi, Lottia gigantea; Mmu, Mus musculus; Nv, Nereis virens; Nve, N. vectensis; Pdu, Platynereis dumerilii; Pfl, Ptychodera flava; Tad, T. adhaerens; Tca, T. castaneum; Xla, Xenopus laevis.
Extended Data Figure 3 Phylogenetic tree including ANTP-class homeodomain families represented in sponges and two additional families characterized by the presence of the YIT motif but excluding divergent A. queenslandica sequences.
Neighbour-joining (JTT, 1,000) bootstrap support values are in black, maximum-likelihood (LG+G 0.4, 1,000 replicates) bootstrap support values are in blue and Bayesian (LG+G 0.4) posterior probability values are in red. Only bootstrap support values equal to or above 500 are shown. All families except Cdx are collapsed for clarity. Ame, A. mellifera; Aqu, A. queenslandica; Bfl, B. floridae; Cte, C. teleta; Dme, D. melanogaster; Dre, D. rerio; Edi, E. dichotoma; Gga, G. gallus; Hsa, H. sapiens; Lgi, L. gigantea; Mmu, M. musculus; Nv, N. virens; Nve, N. vectensis; Pdu, P. dumerilii; Pfl, P. flava; Tad, T. adhaerens; Tca, T. castaneum; Xla, X. laevis. Scale bar indicates the number of amino acid substitutions per site.
Extended Data Figure 4 Phylogenetic tree including ANTP-class homeodomain families represented in sponges and three additional families characterized by the presence of YIT/YIS motifs but excluding some divergent A. queenslandica sequences.
Neighbour-joining (JTT, 1,000 replicates) bootstrap support values are in black, maximum-likelihood (LG+G 0.4, 1,000 replicates) bootstrap support values are in blue and Bayesian (LG+G 0.4) posterior probability values are in red. Only bootstrap support values equal to or above 500 are shown. All subfamilies except Cdx are collapsed for clarity. Ame, A. mellifera; Aqu, A. queenslandica; Bfl, B. floridae; Cte, C. teleta; Dme, D. melanogaster; Dre, D. rerio; Edi, E. dichotoma; Gga, G. gallus; Hsa, H. sapiens; Lgi, L. gigantea; Mmu, M. musculus; Nv, N. virens; Nve, N. vectensis; Pdu, P. dumerilii; Pfl, P. flava; Tad, T. adhaerens; Tca, T. castaneum; Xla, X. laevis. Scale bar indicates the number of amino acid substitutions per site.
Extended Data Figure 5 Phylogenetic tree including ANTP-class homeodomain families represented in sponges and three additional families characterized by the presence of YIT/YIS motifs.
Neighbour-joining (JTT, 1,000 replicates) bootstrap support values are in black, maximum-likelihood (LG+G 0.4, 1,000 replicates) bootstrap support values are in blue and Bayesian (LG+G 0.4) posterior probability values are in red. Only bootstrap support values equal to or above 500 are shown. All families except Cdx are collapsed for clarity. Ame, A. mellifera; Aqu, A. queenslandica; Bfl, B. floridae; Cte, C. teleta; Dme, D. melanogaster; Dre, D. rerio; Edi, E. dichotoma; Gga, G. gallus; Hsa, H. sapiens; Lgi, L. gigantea; Mmu, M. musculus; Nv, N. virens; Nve, N. vectensis; Pdu, P. dumerilii; Pfl, P. flava; Tad, T. adhaerens; Tca, T. castaneum; Xla, X. laevis. Scale bar indicates the number of amino acid substitutions per site.
Extended Data Figure 6 S. ciliatum scaffolds containing NK genes and A. queenslandica scaffold containing cluster of NK genes.
S. ciliatum NK genes are indicated in blue and A. queenslandica NK genes are indicated in green (modified with permission from ref. 6). Annotation of the neighbouring genes (genes within 50 kb from the NK gene) in S. ciliatum was performed using BLASTp searches against the RefSeq database.
Extended Data Figure 7 Additional expression patterns of ANTP-class homeobox genes in embryonic development and during metamorphosis.
a–g, All of the investigated genes (except Hex, data not shown) are expressed in oocytes. The expression of SciNKA is detectable in all blastomeres of the cleavage-stage embryos, but the transcripts are concentrated in the corner-most cytoplasm, which becomes gradually partitioned to the cross cells (arrows). l–q, This subcellular localization of cross-cell-enriched transcripts is also observed for SciNanos, expression of which, similarly to SciNKA, becomes ultimately restricted to cross cells and macromeres in pre-inversion-stage embryos. r–v, In metamorphosing post-larvae, SciNKA is expressed in the cells of the outer layer (r), SciNKB and SciNKG in (possibly non-overlapping) fractions of cells in the inner cell mass (s, t); SciHex is weakly expressed throughout the inner cell mass (u), and SciNKC (v) and NKD (data not shown) are not detectable in the juveniles. Scale bars: 10 μm (a–q); 25 μm (r–v).
Extended Data Figure 8 Samples used for quantification of expression.
a–f, Metamorphosis in S. ciliatum, with stages based on those described previously20 with modifications. Stage I, approximately 12 h after settlement: large flat cells derived from larval macromeres envelop the inner cell mass composed of former micromeres (a). Stage II, approximately 24 h after settlement: single-axis spicules (monaxons) are produced by sclerocytes, which have differentiated from the inner cell mass cells (b). Stage III, 2–3 days after settlement: choanocytes that have differentiated from the inner cells mass cells form a single internal chamber (c). Stage IV, approximately 4 days after settlement: osculum (exhalant opening) forms at the apical end of the spherical juvenile; first tri-radial spicules become apparent (d). Stage V, approximately 10 days after settlement: the juvenile is elongated along the apical–basal axis, long straight spicules form a crown around the osculum (e). Young syconoid sponges, approximately 8 weeks after settlement (f). a–e, Photographs of live specimens in culture; photographs a–d are top (apical) views, cartoon representations of sections and photograph e are side views. Scale bars: 100 μm (a–e); 1 mm (f). g, Details of replicates used for the analysis. Several hundred juveniles were used in each sample. h, Plot demonstrating the results of principal component analysis of the metamorphosis series and axial dissection series of non-reproductive adults calculated according to the previously described method22 and using information about the top 500 differentially expressed genes as default parameters. Metamorphosis stages and parts of sponges are colour coded, with the ovals added manually for easier visualization of similarities and differences between the samples. Progress of development, starting from freshly released larvae and until the emergence of adult, but not yet reproductive sponges, is indicated by arrows. Note similarities of samples within replicates and with neighbouring stages of the metamorphosis series, and distinctiveness of the top (apical) samples from the basal and middle samples of the adults. i, Heat-map representation of sample-to-sample distances among all samples used in this study, calculated according to the previously described method22 and based on expression of all coding genes in S. ciliatum (approximately 18,000 sequences). Note that replicates and neighbouring stages group together, as indicated by highlighting.
Extended Data Figure 9 Heat-map representation of expression profiles.
As demonstrated in Fig. 3b but with data from individual libraries presented separately.
Supplementary information
Supplementary information
This file contains Supplementary Notes 1-3. (PDF 339 kb)
Rights and permissions
About this article
Cite this article
Fortunato, S., Adamski, M., Ramos, O. et al. Calcisponges have a ParaHox gene and dynamic expression of dispersed NK homeobox genes. Nature 514, 620–623 (2014). https://doi.org/10.1038/nature13881
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13881
This article is cited by
-
The compact genome of the sponge Oopsacas minuta (Hexactinellida) is lacking key metazoan core genes
BMC Biology (2023)
-
Microbiome diversity and host immune functions influence survivorship of sponge holobionts under future ocean conditions
The ISME Journal (2022)
-
Progression of regeneration in demosponge Cinachyrella cf cavernosa based on wound location
Hydrobiologia (2020)
-
Convergent evolution of a vertebrate-like methylome in a marine sponge
Nature Ecology & Evolution (2019)
-
Wnt signaling and polarity in freshwater sponges
BMC Evolutionary Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.