Abstract
Site-specific gene addition can allow stable transgene expression for gene therapy. When possible, this is preferred over the use of promiscuously integrating vectors, which are sometimes associated with clonal expansion1 and oncogenesis2. Site-specific endonucleases that can induce high rates of targeted genome editing are finding increasing applications in biological discovery and gene therapy3. However, two safety concerns persist: endonuclease-associated adverse effects, both on-target4 and off-target5,6; and oncogene activation caused by promoter integration, even without nucleases7. Here we perform recombinant adeno-associated virus (rAAV)-mediated promoterless gene targeting without nucleases and demonstrate amelioration of the bleeding diathesis in haemophilia B mice. In particular, we target a promoterless human coagulation factor IX (F9) gene to the liver-expressed mouse albumin (Alb) locus. F9 is targeted, along with a preceding 2A-peptide coding sequence, to be integrated just upstream to the Alb stop codon. While F9 is fused to Alb at the DNA and RNA levels, two separate proteins are synthesized by way of ribosomal skipping. Thus, F9 expression is linked to robust hepatic albumin expression without disrupting it. We injected an AAV8-F9 vector into neonatal and adult mice and achieved on-target integration into ∼0.5% of the albumin alleles in hepatocytes. We established that F9 was produced only from on-target integration, and ribosomal skipping was highly efficient. Stable F9 plasma levels at 7–20% of normal were obtained, and treated F9-deficient mice had normal coagulation times. In conclusion, transgene integration as a 2A-fusion to a highly expressed endogenous gene may obviate the requirement for nucleases and/or vector-borne promoters. This method may allow for safe and efficacious gene targeting in both infants and adults by greatly diminishing off-target effects while still providing therapeutic levels of expression from integration.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
04 November 2014
Fig. 2e was corrected in the PDF owing to a production error.
References
Aiuti, A. et al. Lentiviral hematopoietic stem cell gene therapy in patients with Wiskott–Aldrich syndrome. Science 341, 1233151 (2013)
Hacein-Bey-Abina, S. et al. Insertional oncogenesis in 4 patients after retrovirus-mediated gene therapy of SCID-X1. J. Clin. Invest. 118, 3132–3142 (2008)
Gaj, T., Gersbach, C. A. & Barbas, C. F., III ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 31, 397–405 (2013)
Hendel, A. et al. Quantifying genome-editing outcomes at endogenous loci with SMRT sequencing. Cell Rep. 7, 293–305 (2014)
Cho, S. W. et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 24, 132–141 (2014)
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nature Biotechnol. 31, 822–826 (2013)
Donsante, A. et al. AAV vector integration sites in mouse hepatocellular carcinoma. Science 317, 477 (2007)
Lisowski, L. et al. Ribosomal DNA integrating rAAV-rDNA vectors allow for stable transgene expression. Mol. Ther. 20, 1912–1923 (2012)
Miller, D. G. et al. Gene targeting in vivo by adeno-associated virus vectors. Nature Biotechnol. 24, 1022–1026 (2006)
Li, H. et al. In vivo genome editing restores haemostasis in a mouse model of haemophilia. Nature 475, 217–221 (2011)
Shi, Y., Falahati, R., Zhang, J., Flebbe-Rehwaldt, L. & Gaensler, K. M. Role of antigen-specific regulatory CD4+CD25+ T cells in tolerance induction after neonatal IP administration of AAV-hF.IX. Gene Ther. 20, 987–996 (2013)
Paulk, N. K., Loza, L. M., Finegold, M. J. & Grompe, M. AAV-mediated gene targeting is significantly enhanced by transient inhibition of nonhomologous end joining or the proteasome in vivo. Hum. Gene Ther. 23, 658–665 (2012)
Garcia-Martinez, R. et al. Albumin: pathophysiologic basis of its role in the treatment of cirrhosis and its complications. Hepatology 58, 1836–1846 (2013)
Grimm, D. et al. In vitro and in vivo gene therapy vector evolution via multispecies interbreeding and retargeting of adeno-associated viruses. J. Virol. 82, 5887–5911 (2008)
Lisowski, L. et al. Selection and evaluation of clinically relevant AAV variants in a xenograft liver model. Nature 506, 382–386 (2014)
Li, C. et al. Single amino acid modification of adeno-associated virus capsid changes transduction and humoral immune profiles. J. Virol. 86, 7752–7759 (2012)
Dalkara, D. et al. In vivo-directed evolution of a new adeno-associated virus for therapeutic outer retinal gene delivery from the vitreous. Sci. Transl Med. 5, 189ra176 (2013)
Nathwani, A. C. et al. Adenovirus-associated virus vector-mediated gene transfer in hemophilia B. N. Engl. J. Med. 365, 2357–2365 (2011)
MacLaren, R. E. et al. Retinal gene therapy in patients with choroideremia: initial findings from a phase 1/2 clinical trial. Lancet 383, 1129–1137 (2014)
Paulk, N. K. et al. Adeno-associated virus gene repair corrects a mouse model of hereditary tyrosinemia in vivo. Hepatology 51, 1200–1208 (2010)
Cavazza, A., Moiani, A. & Mavilio, F. Mechanisms of retroviral integration and mutagenesis. Hum. Gene Ther. 24, 119–131 (2013)
Kim, J. H. et al. High cleavage efficiency of a 2A peptide derived from porcine teschovirus-1 in human cell lines, zebrafish and mice. PLoS ONE 6, e18556 (2011)
Johnson, L. A. et al. Gene therapy with human and mouse T-cell receptors mediates cancer regression and targets normal tissues expressing cognate antigen. Blood 114, 535–546 (2009)
Nakai, H. et al. Unrestricted hepatocyte transduction with adeno-associated virus serotype 8 vectors in mice. J. Virol. 79, 214–224 (2005)
Si-Tayeb, K., Lemaigre, F. P. & Duncan, S. A. Organogenesis and development of the liver. Dev. Cell 18, 175–189 (2010)
Nakai, H. et al. Extrachromosomal recombinant adeno-associated virus vector genomes are primarily responsible for stable liver transduction in vivo. J. Virol. 75, 6969–6976 (2001)
Calcedo, R. & Wilson, J. M. Humoral immune response to AAV. Front. Immunol. 4, 341 (2013)
Anguela, X. M. et al. ZFN mediated targeting of albumin “safe harbor” results in therapeutic levels of human factor viii in a mouse model of hemophilia A. Blood 122, 720 (2013)
Yew, N. S. & Cheng, S. H. Gene therapy for lysosomal storage disorders. Pediatr. Endocrinol. Rev. 11 (suppl. 1). 99–109 (2013)
Balazs, A. B. et al. Antibody-based protection against HIV infection by vectored immunoprophylaxis. Nature 481, 81–84 (2011)
Grimm, D., Pandey, K., Nakai, H., Storm, T. A. & Kay, M. A. Liver transduction with recombinant adeno-associated virus is primarily restricted by capsid serotype not vector genotype. J. Virol. 80, 426–439 (2006)
Lu, J., Zhang, F. & Kay, M. A. A mini-intronic plasmid (MIP): a novel robust transgene expression vector in vivo and in vitro. Mol. Ther. 21, 954–963 (2013)
Mitchell, C. & Willenbring, H. A reproducible and well-tolerated method for 2/3 partial hepatectomy in mice. Nature Protocols 3, 1167–1170 (2008)
Rogers, G. L. & Hoffman, B. E. Optimal immunofluorescent staining for human factor ix and infiltrating T cells following gene therapy for hemophilia B. J. Genet. Syndr. Gene Ther. Suppl. 1, 012 (2012)
Grimm, D. et al. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 441, 537–541 (2006)
Acknowledgements
This work was supported by a grant to M.A.K. from the National Heart Lung & Blood Institute (R01-HL064274). A.B. was supported by a fellowship from the Lucile Packard Foundation for Children’s Health, Stanford NIH-NCATS-CTSA UL1 TR001085 and Child Health Research Institute of Stanford University. N.K.P. was supported by a fellowship from the National Heart Lung & Blood Institute (F32-HL119059), the Hans Popper Memorial Fellowship from the American Liver Foundation, and the Stanford Dean’s Fellowship. M.H.P. was supported by the Laurie Krauss Lacob Faculty Scholar Fund in Pediatric Translational Medicine. The funding organizations played no role in experimental design, data analysis or manuscript preparation.
Author information
Authors and Affiliations
Contributions
A.B., N.K.P., M.H.P., K.M.G. and M.A.K. designed the experiments. A.B., N.K.P., Y.S., Y.H., K.C., F.Z., P.N.V. and L.P.S. generated reagents and performed the experiments. A.B., N.K.P. and M.A.K. wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.B. and M.A.K. are founders of LogicBio Therapeutics, a startup biotechnology company with interests in the technology described in the manuscript.
Extended data figures and tables
Extended Data Figure 1 Human F9 liver immunohistochemistry.
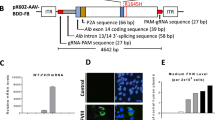
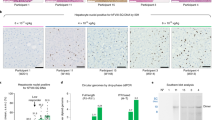
From top to bottom, panels show human F9 staining (red) with 4′,6-diamidino-2-phenylindole (DAPI) nuclear counterstain (blue) in positive control human liver, negative control untreated mouse liver, and two sets of representative stains from mice treated as neonates or adults with AAV8-F9. Original magnification, ×200.
Extended Data Figure 2 Scheme of targeting rate assessment.
Assessment of on-target integration rate begins using linear amplification with biotinylated primer 1 (black), annealing to the genomic locus but not to the vector (step 1). Linear amplicons are then bound to streptavidinylated beads and washed to exclude episomal vectors (step 2). Subsequent second-strand DNA synthesis with random primers (step 3) was followed by CviQI restriction digestion (step 4). A compatible linker is then ligated (step 5) followed by two rounds of nested PCR amplifications (primers 2–3 in blue (step 6), and then primers 4–5 in red (step 7)). CviQI cleaves at the same distance from the homology border in both targeted and wild-type alleles, thus allowing for unbiased amplification. The amplicons of the second nested PCR then serve as a template for qPCR assays with either primers 6–7 (green) or 8–9 (orange) (step 8).
Extended Data Figure 4 Toxicity assessment by ALT measurement.
Alanine transaminase levels (ALT) were evaluated 7 days after injection in mice injected with AAV8 coding for our experimental vector (1 × 1012) or a negative control coding for a known non-toxic cassette (1 × 1012 of H1 promoter-driven shRNA), or a positive control coding for a known toxic cassette (5 × 1011 of U6 promoter-driven shRNA). Data represent mean of two measurements of four independent mice for each groups. The statistical significance is defined here as having P < 0.05 in a one-tailed t-test between samples of different variance.
Extended Data Figure 5 Vector copy number.
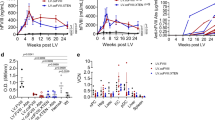
Vector copy number assessed by qPCR using primers 8 and 9 (Fig. 3). n = 7 for mice injected as adults; n = 6 for mice injected as neonates and analysed before or after partial hepatectomy (PH). Error bars represent s.d.
Rights and permissions
About this article
Cite this article
Barzel, A., Paulk, N., Shi, Y. et al. Promoterless gene targeting without nucleases ameliorates haemophilia B in mice. Nature 517, 360–364 (2015). https://doi.org/10.1038/nature13864
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13864
This article is cited by
-
Targeted Gene Insertion: The Cutting Edge of CRISPR Drug Development with Hemophilia as a Highlight
BioDrugs (2024)
-
Novel expression cassettes for increasing apolipoprotein AI transgene expression in vascular endothelial cells
Scientific Reports (2022)
-
Fludarabine increases nuclease-free AAV- and CRISPR/Cas9-mediated homologous recombination in mice
Nature Biotechnology (2022)
-
Therapeutic homology-independent targeted integration in retina and liver
Nature Communications (2022)
-
Frequent aneuploidy in primary human T cells after CRISPR–Cas9 cleavage
Nature Biotechnology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.