Abstract
Intracellular ISG15 is an interferon (IFN)-α/β-inducible ubiquitin-like modifier which can covalently bind other proteins in a process called ISGylation; it is an effector of IFN-α/β-dependent antiviral immunity in mice1,2,3,4. We previously published a study describing humans with inherited ISG15 deficiency but without unusually severe viral diseases5. We showed that these patients were prone to mycobacterial disease and that human ISG15 was non-redundant as an extracellular IFN-γ-inducing molecule. We show here that ISG15-deficient patients also display unanticipated cellular, immunological and clinical signs of enhanced IFN-α/β immunity, reminiscent of the Mendelian autoinflammatory interferonopathies Aicardi–Goutières syndrome and spondyloenchondrodysplasia6,7,8,9. We further show that an absence of intracellular ISG15 in the patients’ cells prevents the accumulation of USP1810,11, a potent negative regulator of IFN-α/β signalling, resulting in the enhancement and amplification of IFN-α/β responses. Human ISG15, therefore, is not only redundant for antiviral immunity, but is a key negative regulator of IFN-α/β immunity. In humans, intracellular ISG15 is IFN-α/β-inducible not to serve as a substrate for ISGylation-dependent antiviral immunity, but to ensure USP18-dependent regulation of IFN-α/β and prevention of IFN-α/β-dependent autoinflammation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
BioProject
Gene Expression Omnibus
Data deposits
Microarray data have been deposited in the Gene Expression Omnibus under accession number GSE60359; WES data have been deposited in the BioProject database under accession number PRJNA167660.
References
Jeon, Y. J., Yoo, H. M. & Chung, C. H. ISG15 and immune diseases. Biochim. Biophys. Acta 1802, 485–496 (2010)
Morales, D. J. & Lenschow, D. J. The antiviral activities of ISG15. J. Mol. Biol. 425, 4995–5008 (2013)
Skaug, B. & Chen, Z. J. Emerging role of ISG15 in antiviral immunity. Cell 143, 187–190 (2010)
Zhang, D. & Zhang, D. E. Interferon-stimulated gene 15 and the protein ISGylation system. J. Interferon Cytokine Res. 31, 119–130 (2011)
Bogunovic, D. et al. Mycobacterial disease and impaired IFN-γ immunity in humans with inherited ISG15 deficiency. Science 337, 1684–1688 (2012)
Baechler, E. C. et al. Interferon-inducible gene expression signature in peripheral blood cells of patients with severe lupus. Proc. Natl Acad. Sci. USA 100, 2610–2615 (2003)
Crow, Y. J. Aicardi–Goutières syndrome. Handb. Clin. Neurol. 113, 1629–1635 (2013)
Crow, Y. J. & Rehwinkel, J. Aicardi–Goutières syndrome and related phenotypes: linking nucleic acid metabolism with autoimmunity. Hum. Mol. Genet. 18, R130–R136 (2009)
Rice, G. I. et al. Assessment of interferon-related biomarkers in Aicardi–Goutières syndrome associated with mutations in TREX1, RNASEH2A, RNASEH2B, RNASEH2C, SAMHD1, and ADAR: a case–control study. Lancet Neurol. 12, 1159–1169 (2013)
Malakhova, O. A. et al. UBP43 is a novel regulator of interferon signaling independent of its ISG15 isopeptidase activity. EMBO J. 25, 2358–2367 (2006)
Tokarz, S. et al. The ISG15 isopeptidase UBP43 is regulated by proteolysis via the SCFSkp2 ubiquitin ligase. J. Biol. Chem. 279, 46424–46430 (2004)
Kendall, B. & Cavanagh, N. Intracranial calcification in paediatric computed tomography. Neuroradiology 28, 324–330 (1986)
Livingston, J. H., Stivaros, S., van der Knaap, M. S. & Crow, Y. J. Recognizable phenotypes associated with intracranial calcification. Dev. Med. Child Neurol. 55, 46–57 (2013)
Manyam, B. V. What is and what is not ‘Fahr’s disease'. Parkinsonism Relat. Disord. 11, 73–80 (2005)
Wang, C. et al. Mutations in SLC20A2 link familial idiopathic basal ganglia calcification with phosphate homeostasis. Nature Genet. 44, 254–256 (2012)
Keller, A. et al. Mutations in the gene encoding PDGF-B cause brain calcifications in humans and mice. Nature Genet. 45, 1077–1082 (2013)
Nicolas, G. et al. Mutation of the PDGFRB gene as a cause of idiopathic basal ganglia calcification. Neurology 80, 181–187 (2013)
Briggs, T. A. et al. Tartrate-resistant acid phosphatase deficiency causes a bone dysplasia with autoimmunity and a type I interferon expression signature. Nature Genet. 43, 127–131 (2011)
Casanova, J. L. & Abel, L. Genetic dissection of immunity to mycobacteria: the human model. Annu. Rev. Immunol. 20, 581–620 (2002)
Casanova, J. L. & Abel, L. The genetic theory of infectious diseases: a brief history and selected illustrations. Annu. Rev. Genomics Hum. Genet. 14, 215–243 (2013)
Bogunovic, D., Boisson-Dupuis, S. & Casanova, J. L. ISG15: leading a double life as a secreted molecule. Exp. Mol. Med. 45, e18 (2013)
Bennett, L. et al. Interferon and granulopoiesis signatures in systemic lupus erythematosus blood. J. Exp. Med. 197, 711–723 (2003)
Raymond, A. A., Zariah, A. A., Samad, S. A., Chin, C. N. & Kong, N. C. Brain calcification in patients with cerebral lupus. Lupus 5, 123–128 (1996)
Durfee, L. A., Lyon, N., Seo, K. & Huibregtse, J. M. The ISG15 conjugation system broadly targets newly synthesized proteins: implications for the antiviral function of ISG15. Mol. Cell 38, 722–732 (2010)
Broering, R. et al. The interferon stimulated gene 15 functions as a proviral factor for the hepatitis C virus and as a regulator of the IFN response. Gut 59, 1111–1119 (2010)
Chua, P. K. et al. Modulation of alpha interferon anti-hepatitis C virus activity by ISG15. J. Gen. Virol. 90, 2929–2939 (2009)
François-Newton, V. et al. USP18-based negative feedback control is induced by type I and type III interferons and specifically inactivates interferon α response. PLoS ONE 6, e22200 (2011)
Francois-Newton, V., Livingstone, M., Payelle-Brogard, B., Uze, G. & Pellegrini, S. USP18 establishes the transcriptional and anti-proliferative interferon α/β differential. Biochem. J. 446, 509–516 (2012)
Frescas, D. & Pagano, M. Deregulated proteolysis by the F-box proteins SKP2 and β-TrCP: tipping the scales of cancer. Nature Rev. Cancer 8, 438–449 (2008)
Hertzog, P. J. & Williams, B. R. Fine tuning type I interferon responses. Cytokine Growth Factor Rev. 24, 217–225 (2013)
Li, Q. Z. et al. Protein array autoantibody profiles for insights into systemic lupus erythematosus and incomplete lupus syndromes. Clin. Exp. Immunol. 147, 60–70 (2007)
Acknowledgements
The Laboratory of Human Genetics of Infectious Diseases is supported by grants from the French National Agency for Research (ANR), the EU grant HOMITB (HEALTH-F32008-200732), the St Giles Foundation, the National Center for Research Resources and the National Center for Advancing Sciences (NCATS), National Institutes of Health grant number 8UL1TR000043, the Rockefeller University, the National Institute of Allergy and Infectious Diseases grant number R37AI095983, Institut Merieux research grant and the Empire State Stem Cell fund through NYSDOH Contract #C023046 to Flow Cytometry Research Core at the Rockefeller University. The Cytokine Signaling Unit is supported by the Institut Pasteur, CNRS and INSERM. S.P. and G.U. received funding from the EU Seventh Framework Programme under grant agreement 223608. V.F.-N. was supported by the Ligue contre le Cancer. L.R. is a Human Frontier Science Program long-term fellow. L.D.N. was supported by the National Institute of Allergy and Infectious Diseases grant number 1PO1AI076210-01A1. Y.J.C. thanks the Manchester Biomedical Research Centre and the Greater Manchester Comprehensive Local Research Network, the European Union’s Seventh Framework Programme (FP7/2007-2013) under grant agreement 241779, and the European Research Council (GA 309449). A.G.-S. acknowledges NIAID grants U19AI083025 and P01AI090935 for support. We thank C. Daussy for technical assistance, E. Bianchi and F. Michel for discussions. We thank D. Zhang and the members of the Zhang laboratory for assistance, advice and discussions. This work was supported by Chinese National Natural Science Foundation grants (81000079, 81170165) to X.Z. D.B. is supported by the National Institute of Allergy and Infectious Diseases grant number R00AI106942-02.
Author information
Authors and Affiliations
Contributions
D.B., X.Z., B.P.-B., V.F.-N., O.S., D.M., P.G., A.G.-S., L.A., P.L., L.D.N., S.B.-D., Y.J.C., J.-L.C. and S.P. wrote the manuscript. D.B., X.Z., B.P.-B., V.F.-N., S.D.S., C.Y., S.V., Z.L., I.T., G.I.R., C.C., N.M., S.A.M., Y.I., B.B., S.O., L.Z., X.W., H.J., W.L., T.H., D.L., T.M., B.W., D.Y., L.R., G.U., P.G., F.R., S.-Y.Z., E.J., J.B., A.G.-S., L.A., P.L., L.D.N., S.B.-D., Y.J.C., J.-L.C. and S.P. designed and/or performed experiments. M.L., J.-Y.L., Q.K.W., O.S., D.M., N.M., I.T. and S.A.M. took clinical care of the patients and provided advice.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Mutations in ISG15-deficient individuals, allele characterization and serum IFN-α concentrations.
a, Sanger sequencing of ISG15 exon 2 from genomic DNA in kindred C, with the variants highlighted. b, The wild type (WT) and three mutant alleles (G55X, L114fs, E127X- ISG15) were inserted into an expression vector and used to transfect HEK293T cells. Other HEK293T cells were mock-transfected (mock) or left untransfected (not transf.). The cell lysates isolated were subjected to western blotting, with recombinant human (Rh)-ISG15 used as a control. c, d, Plasma samples from P1, P2, P3, P5, P6 and the mother and brother of P5/6 were used in cytopathic protection assays, to measure antiviral activity (c) and blocking antibodies against IFN-α were used to assess specificity (d) (experiment was performed one time).
Extended Data Figure 2 A form of ISG15 that cannot be conjugated rescues the phenotype of ISG15-deficient cells.
a, Lentiviral particles containing luciferase, wild-type (WT) ISG15-RFP or ISG15(ΔGG)-RFP genes were used to transduce hTert-immortalized fibroblasts from C1, a STAT1−/− subject, P1, P2 and P3. RFP-positive cells were obtained by sorting and were cultured for a few weeks. The cells were then treated with 1,000 IU of IFN-α2b for 12 h, washed with PBS and left to rest for 36 h, after which relative mRNA levels for IFIT1 were determined. b, The experimental setting described in a was used in the presence or absence of vehicle control, anti-ISG15 antibodies or control IgG for luciferase and wild-type ISG15–RFP-transduced C1 and P1 hTert-immortalized fibroblasts (showing representative experiments with technical replicates and s.e.m., out of 3 performed).
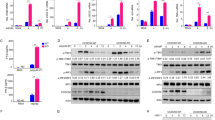
Extended Data Figure 3 Prolonged IFN signalling, low USP18, and high interferon-stimulated-gene-encoded protein levels in patient-derived cells and in ISG15-silenced human fibrosarcoma HLLR1-1.4 cells.
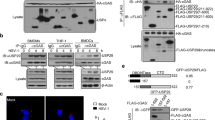
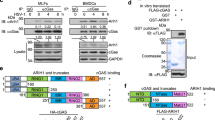
a, Left panels, SV40-immortalized fibroblasts from two controls (C10 and C12) and two ISG15-deficient patients (P1 and P2) were stimulated with IFN-β (500 pM) for 4 to 36 h. Cell lysates (30 µg) were analysed with the indicated antibodies. Right, EBV-transformed B cells from control (C3) and patient P1 were stimulated with IFN-β for 8 to 24 h. Cell lysates (30 µg) were analysed with the indicated antibodies. b, HLLR1-1.4 cells were transfected with control siRNA or ISG15 siRNA. One day post-transfection, IFN-β (500 pM) was added for various amounts of time. Cell lysates (30 µg) were analysed with the indicated antibodies (MxA and MX1 are used synonymously). c, WISH cells were stimulated and lysates analysed as described in b. d, HLLR1-1.4 cells were transfected with control siRNA, USP18 siRNA and UbcH8 (also known as UBE2E2) siRNA (left) or control siRNA, USP18 siRNA and HERC5 siRNA (right). One day post-transfection, cells were left untreated (naive) or were primed for 8 h with IFN-β (500 pM). Cells were washed and left to rest for 16 h before being pulsed for 30 min with 100 pM IFN-α2 or IFN-β. Cell lysates (30 µg) were analysed with the indicated antibodies.
Extended Data Figure 4 ISG15 controls the stability of the USP18 protein, but not of other interferon-stimulated-gene products.
a, HLLR1-1.4 cells were transfected with either control siRNA or ISG15 siRNA. One day post-transfection, cells were stimulated with IFN-β (500 pM) for 6 h. Cycloheximide (CHX, 20 µg ml−1) was then added for various time periods, from 30 min to 5 h. Cell lysates (30 µg) were analysed with the indicated antibodies. b, As in a, with additional controls, cells treated with IFN only. Several interferon-stimulated genes were analysed. c, hTert-immortalized fibroblasts from patient P1 transduced with lentiviral particles expressing RFP and luciferase and wild-type ISG15 (LV ISG15 WT) were stimulated with IFN-β (500 pM) for 6 h. CHX was then added for the indicated times. Cell lysates (15 µg) were analysed by western blotting. d, HEK293T cells were transfected with USP18, HA–ubiquitin and Flag–ISG15 as indicated. Two days later, cells were lysed in modified RIPA buffer, USP18 was immunoprecipitated (IP) and analysed with anti-USP18 antibodies. Left panels, cell lysates (30 µg) were analysed by western blot with the indicated antibodies. Right panels, the immunoprecipitates were gel separated and transferred onto a membrane. The membrane was cut into two parts above the 50 kDa marker, both of which were blotted with anti-USP18 antibodies. The top part was exposed for 2 min, the bottom part for 20 s. Asterisk indicates IgG heavy chain. e, HEK293T cells were transfected with 1 µg of the USP18 construct alone or with 1 µg of HA–ubiquitin, in the presence or absence of Flag-tagged ISG15, either wild type or a mutant form of ISG15 that cannot be conjugated as it lacks the two carboxy-terminal glycine residues (Flag–ISG15(ΔGG)). Two days later, cells were lysed in modified RIPA buffer, USP18 was immunoprecipitated and analysed with anti-HA or anti-USP18 antibodies. Asterisk indicates IgG heavy chain. f, hTert-immortalized fibroblasts from patient P3 were transfected with control siRNA or SKP2 siRNA. We added IFN-β (500 pM) 24 h later and the cells were incubated for the indicated times. Cell lysates were analysed with the indicated antibodies and USP18 levels were determined as a function of actin levels.
Extended Data Figure 5 Autoantibody development in ISG15-deficient individuals.
a, b, Serum samples from ISG15-deficient, SLE and AGS patients were evaluated for the presence of IgG and IgA autoantibodies in a blinded experiment. Values for the negative control samples for each antigen were averaged and ratios of each sample to the mean for the negative controls plus 2 standard deviations were calculated, with values greater than 1 considered positive. A heat map of the ratio values was generated with MultiExperiment Viewer software (MeV, DFCI Boston, MA), with values coded as follows: 0, blue; 1, black; 5, yellow.
Supplementary information
Supplementary Information
This file contains details of the case study. (PDF 117 kb)
Rights and permissions
About this article
Cite this article
Zhang, X., Bogunovic, D., Payelle-Brogard, B. et al. Human intracellular ISG15 prevents interferon-α/β over-amplification and auto-inflammation. Nature 517, 89–93 (2015). https://doi.org/10.1038/nature13801
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13801
This article is cited by
-
Inborn errors of immunity: an expanding universe of disease and genetic architecture
Nature Reviews Genetics (2024)
-
IL4Rα and IL17A Blockade Rescue Autoinflammation in SOCS1 Haploinsufficiency
Journal of Clinical Immunology (2024)
-
ISG15 targets glycosylated PD-L1 and promotes its degradation to enhance antitumor immune effects in lung adenocarcinoma
Journal of Translational Medicine (2023)
-
Mendelian susceptibility to mycobacterial disease: an overview
Egyptian Journal of Medical Human Genetics (2023)
-
The crucial regulatory role of type I interferon in inflammatory diseases
Cell & Bioscience (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.