Abstract
The formation of branched lariat RNA is an evolutionarily conserved feature of splicing reactions for both group II and spliceosomal introns. The lariat is important for the fidelity of 5′ splice-site selection and consists of a 2′-5′ phosphodiester bond between a bulged adenosine and the 5′ end of the intron. To gain insight into this ubiquitous intramolecular linkage, we determined the crystal structure of a eukaryotic group IIB intron in the lariat form at 3.7 Å. This revealed that two tandem tetraloop-receptor interactions, η–η′ and π–π′, place domain VI in the core to position the lariat bond in the post-catalytic state. On the basis of structural and biochemical data, we propose that π–π′ is a dynamic interaction that mediates the transition between the two steps of splicing, with η–η′ serving an ancillary role. The structure also reveals a four-magnesium-ion cluster involved in both catalysis and positioning of the 5′ end. Given the evolutionary relationship between group II and nuclear introns, it is likely that this active site configuration exists in the spliceosome as well.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Grabowski, P. J., Padgett, R. A. & Sharp, P. A. Messenger RNA splicing in vitro: an excised intervening sequence and a potential intermediate. Cell 37, 415–427 (1984)
Padgett, R. A., Konarska, M. M., Grabowski, P. J., Hardy, S. F. & Sharp, P. A. Lariat RNA’s as intermediates and products in the splicing of messenger RNA precursors. Science 225, 898–903 (1984)
Konarska, M. M., Grabowski, P. J., Padgett, R. A. & Sharp, P. A. Characterization of the branch site in lariat RNAs produced by splicing of mRNA precursors. Nature 313, 552–557 (1985)
Peebles, C. L. et al. A self-splicing RNA excises an intron lariat. Cell 44, 213–223 (1986)
van der Veen, R. et al. Excised group II introns in yeast mitochondria are lariats and can be formed by self-splicing in vitro. Cell 44, 225–234 (1986)
Di Leo, E. et al. A point mutation in the lariat branch point of intron 6 of NPC1 as the cause of abnormal pre-mRNA splicing in Niemann-Pick type C disease. Hum. Mutat. 24, 440 (2004)
Galej, W. P., Oubridge, C., Newman, A. J. & Nagai, K. Crystal structure of Prp8 reveals active site cavity of the spliceosome. Nature 493, 638–643 (2013)
Fica, S. M. et al. RNA catalyses nuclear pre-mRNA splicing. Nature 503, 229–234 (2013)
Toor, N., Keating, K. S., Taylor, S. D. & Pyle, A. M. Crystal structure of a self-spliced group II intron. Science 320, 77–82 (2008)
Lambowitz, A. M. & Zimmerly, S. Group II introns: mobile ribozymes that invade DNA. Cold Spring Harb. Perspect. Biol. 3, a003616 (2011)
Toor, N., Hausner, G. & Zimmerly, S. Coevolution of group II intron RNA structures with their intron-encoded reverse transcriptases. RNA 7, 1142–1152 (2001)
Michel, F., Umesono, K. & Ozeki, H. Comparative and functional anatomy of group II catalytic introns–a review. Gene 82, 5–30 (1989)
Costa, M., Fontaine, J. M., Loiseaux-de Goër, S. & Michel, F. A group II self-splicing intron from the brown alga Pylaiella littoralis is active at unusually low magnesium concentrations and forms populations of molecules with a uniform conformation. J. Mol. Biol. 274, 353–364 (1997)
Rest, J. S. & Mindell, D. P. Retroids in archaea: phylogeny and lateral origins. Mol. Biol. Evol. 20, 1134–1142 (2003)
Toor, N., Robart, A. R., Christianson, J. & Zimmerly, S. Self-splicing of a group IIC intron: 5′ exon recognition and alternative 5′ splicing events implicate the stem-loop motif of a transcriptional terminator. Nucleic Acids Res. 34, 6461–6471 (2006)
Boudvillain, M. & Pyle, A. M. Defining functional groups, core structural features and inter-domain tertiary contacts essential for group II intron self-splicing: a NAIM analysis. EMBO J. 17, 7091–7104 (1998)
Jacquier, A. & Michel, F. Base-pairing interactions involving the 5′ and 3′-terminal nucleotides of group II self-splicing introns. J. Mol. Biol. 213, 437–447 (1990)
Fedorova, O. & Pyle, A. M. A conserved element that stabilizes the group II intron active site. RNA 14, 1048–1056 (2008)
Fedorova, O., Mitros, T. & Pyle, A. M. Domains 2 and 3 interact to form critical elements of the group II intron active site. J. Mol. Biol. 330, 197–209 (2003)
Chanfreau, G. & Jacquier, A. An RNA conformational change between the two chemical steps of group II self-splicing. EMBO J. 15, 3466–3476 (1996)
Adams, P. L., Stahley, M. R., Kosek, A. B., Wang, J. & Strobel, S. A. Crystal structure of a self-splicing group I intron with both exons. Nature 430, 45–50 (2004)
Li, C. F., Costa, M. & Michel, F. Linking the branchpoint helix to a newly found receptor allows lariat formation by a group II intron. EMBO J. 30, 3040–3051 (2011)
Klein, D. J., Moore, P. B. & Steitz, T. A. The contribution of metal ions to the structural stability of the large ribosomal subunit. RNA 10, 1366–1379 (2004)
Fica, S. M., Mefford, M. A., Piccirilli, J. A. & Staley, J. P. Evidence for a group II intron-like catalytic triplex in the spliceosome. Nature Struct. Mol. Biol. 21, 464–471 (2014)
Marcia, M. & Pyle, A. M. Visualizing group II intron catalysis through the stages of splicing. Cell 151, 497–507 (2012)
Query, C. C., Moore, M. J. & Sharp, P. A. Branch nucleophile selection in pre-mRNA splicing: evidence for the bulged duplex model. Genes Dev. 8, 587–597 (1994)
Anokhina, M. et al. RNA structure analysis of human spliceosomes reveals a compact 3D arrangement of snRNAs at the catalytic core. EMBO J. 32, 2804–2818 (2013)
Parker, R. & Guthrie, C. A point mutation in the conserved hexanucleotide at a yeast 5′ splice junction uncouples recognition, cleavage, and ligation. Cell 41, 107–118 (1985)
Lesser, C. F. & Guthrie, C. Mutations in U6 snRNA that alter splice site specificity: implications for the active site. Science 262, 1982–1988 (1993)
Roitzsch, M. & Pyle, A. M. The linear form of a group II intron catalyzes efficient autocatalytic reverse splicing, establishing a potential for mobility. RNA 15, 473–482 (2009)
Yang, J., Zimmerly, S., Perlman, P. S. & Lambowitz, A. M. Efficient integration of an intron RNA into double-stranded DNA by reverse splicing. Nature 381, 332–335 (1996)
Lynch, M. & Richardson, A. O. The evolution of spliceosomal introns. Curr. Opin. Genet. Dev. 12, 701–710 (2002)
Tseng, C. K. & Cheng, S. C. Both catalytic steps of nuclear pre-mRNA splicing are reversible. Science 320, 1782–1784 (2008)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Keating, K. S. & Pyle, A. M. RCrane: semi-automated RNA model building. Acta Crystallogr. D 68, 985–995 (2012)
Blanc, E. et al. Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr. D 60, 2210–2221 (2004)
Schroder, G. F., Levitt, M. & Brunger, A. T. Super-resolution biomolecular crystallography with low-resolution data. Nature 464, 1218–1222 (2010)
Chou, F. C., Sripakdeevong, P., Dibrov, S. M., Hermann, T. & Das, R. Correcting pervasive errors in RNA crystallography through enumerative structure prediction. Nature Methods 10, 74–76 (2013)
Morin, A. et al. Collaboration gets the most out of software. eLife 2, e01456 (2013)
Chanfreau, G. & Jacquier, A. Interaction of intronic boundaries is required for the second splicing step efficiency of a group II intron. EMBO J. 12, 5173–5180 (1993)
Parker, R. & Siliciano, P. G. Evidence for an essential non-Watson-Crick interaction between the first and last nucleotides of a nuclear pre-mRNA intron. Nature 361, 660–662 (1993)
Basu, S. et al. A specific monovalent metal ion integral to the AA platform of the RNA tetraloop receptor. Nature Struct. Biol. 5, 986–992 (1998)
Acknowledgements
We thank S. Banerjee and the staff of the NE-CAT beamlines at the Advanced Photon Source (APS) of Argonne National Laboratory. We thank P. Ghosh, S. Joseph, G. Ghosh, R. Doolittle, Y. Tor, D. Donoghue and T. Wiryaman for comments on the manuscript. We thank R. Das and F.-C. Chou for assistance with phenix.erasser for structure refinement and G. Bricogne for advice on Buster refinement. We also thank N. T. Schirle for preliminary biochemical characterization of the P.li.LSUI2 intron. R.T.C. was supported by the Cell, Molecular, and Genetics Training Program funded by NIH predoctoral training grant 5T32GM007240. J.K.P. was supported by the UCSD Molecular Biophysics Training Program funded by NIH predoctoral training grant 5T32GM008326. NE-CAT is supported by NIH grant 8P41GM103403-10 and APS is supported by the US DOE under contract number DE-AC02-06CH11357. This work was supported by a Hellman Foundation Fellowship and NIH grant 5R01GM102216 awarded to N.T.
Author information
Authors and Affiliations
Contributions
A.R.R. and J.K.P. performed the experiments. A.R.R., R.T.C., J.K.P. and N.T. designed the experiments. A.R.R., R.T.C., J.K.P., K.R.R. and N.T. analysed the data. A.R.R., R.T.C. and N.T. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
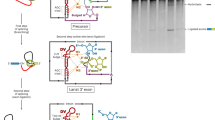
Extended Data Figure 1 Secondary structure of P.li.LSUI2 intron crystallization construct.
Tertiary interactions are indicated with Greek letters and domains are labelled with Roman numerals. Colouring of the individual domains is consistent with the overall view of the tertiary structure shown in Fig. 1.
Extended Data Figure 3 The path of the 5′ exon through the intron structure.
The EBS1–IBS1 and EBS2–IBS2 interactions position the 5′ exon. They do not form a continuous binding interface with the presence of a highly distorted backbone at the junction between these two motifs. As a result, the helical axes of the EBS1–IBS1 and EBS2–IBS2 pairings are positioned ∼90° relative to each other. The EBS3–IBS3 interaction places the 3′ exon in the active site.
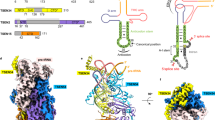
Extended Data Figure 4 Overall tertiary structure of the P.li.LSUI2 intron.
Individual domains and subdomains are depicted in different colours. Domain names are labelled with Roman numerals. a and b show different rotations of the intron structure.
Extended Data Figure 5 Companion to Fig. 2 showing the location of the individual tertiary interactions relative to the overall structure.
a, κ–κ′. b, ε–ε′. c, ρ–ρ′.
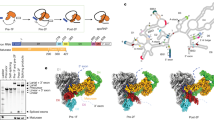
Extended Data Figure 6 Splicing assays for the DVI mutants showing the proportion of branched product.
Blue and orange bars correspond to lariat 3′ exon and lariat, respectively. The Δη′ and Δπ mutants accumulate large amounts of lariat 3′ exon, thus indicating a second-step splicing defect. The Δη′Δπ double mutant is almost completely blocked before the second step. The A620C mutant shows 2.4-fold greater accumulation of lariat 3′ exon compared to the wild-type intron, indicating that the interaction between G1 and A620 is important for the second step. In the yeast aI5γ intron, G1 instead interacts with the penultimate residue44, indicating a certain degree of flexibility for this pairing. There is evidence for a similar interaction between the termini of nuclear introns45 involving nearby (but not exactly equivalent) residues, which also has a significant effect upon the second step of splicing. Therefore, the 5′ and 3′ ends of nuclear introns may have a similar arrangement within the spliceosome.
Extended Data Figure 8 Anomalous maps identifying core metal ions.
a, b, Depiction of the RNA ligands surrounding metals M3 and M4, respectively. Yb3+ anomalous map contoured at 9σ. c, Yb3+ anomalous map for wild type contoured at 9σ. d, Compared with the wild-type intron, the Yb3+ anomalous map for the  G79A mutant (contoured at 4σ) is lacking the peaks corresponding to M3 and M4, even at a lower contour level. e, Tl+ was used as a probe for monovalent ions in the RNA structure46. The Tl+ anomalous map (purple mesh contoured at 5.5σ) revealed a strong peak located 3.8 Å from M1 that coordinates to the nucleobase of J2/3 residue G421 and the backbone of DV nucleotide G550. This sodium ion Na1 (purple sphere) is significantly closer to M1 than the equivalent K+ ion found in O. iheyensis25. Otherwise, this monovalent ion binding site is relatively conserved between these two introns.
G79A mutant (contoured at 4σ) is lacking the peaks corresponding to M3 and M4, even at a lower contour level. e, Tl+ was used as a probe for monovalent ions in the RNA structure46. The Tl+ anomalous map (purple mesh contoured at 5.5σ) revealed a strong peak located 3.8 Å from M1 that coordinates to the nucleobase of J2/3 residue G421 and the backbone of DV nucleotide G550. This sodium ion Na1 (purple sphere) is significantly closer to M1 than the equivalent K+ ion found in O. iheyensis25. Otherwise, this monovalent ion binding site is relatively conserved between these two introns.
Extended Data Figure 9 2Fo − Fc density for DVI in the pre-catalytic structure contoured at 1σ.
The η–η′ interaction persists throughout the splicing reaction and is visible in the pre-catalytic state. The weaker density for the central region of DVI suggests a partially disordered, dynamic region with possible helical remodelling in the conserved internal loop during splicing. The general pattern of side-by-side packing of domains II and VI persists between the two steps. Catalytic triad mutation consisted of an AGC→GAU substitution.
Rights and permissions
About this article
Cite this article
Robart, A., Chan, R., Peters, J. et al. Crystal structure of a eukaryotic group II intron lariat. Nature 514, 193–197 (2014). https://doi.org/10.1038/nature13790
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13790
This article is cited by
-
Identification of polycistronic transcriptional units and non-canonical introns in green algal chloroplasts based on long-read RNA sequencing data
BMC Genomics (2021)
-
Zinc-finger protein CNBP alters the 3-D structure of lncRNA Braveheart in solution
Nature Communications (2020)
-
Metal ions and sugar puckering balance single-molecule kinetic heterogeneity in RNA and DNA tertiary contacts
Nature Communications (2020)
-
Visualizing group II intron dynamics between the first and second steps of splicing
Nature Communications (2020)
-
Structural basis for the second step of group II intron splicing
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.