Abstract
The incidences of chronic inflammatory disorders have increased considerably over the past three decades1. Recent shifts in dietary consumption may have contributed importantly to this surge, but how dietary consumption modulates inflammatory disease is poorly defined. Pstpip2cmo mice, which express a homozygous Leu98Pro missense mutation in the Pombe Cdc15 homology family protein PSTPIP2 (proline-serine-threonine phosphatase interacting protein 2), spontaneously develop osteomyelitis that resembles chronic recurrent multifocal osteomyelitis in humans2,3,4. Recent reports demonstrated a crucial role for interleukin-1β (IL-1β) in osteomyelitis, but deletion of the inflammasome components caspase-1 and NLRP3 failed to rescue Pstpip2cmo mice from inflammatory bone disease5,6. Thus, the upstream mechanisms controlling IL-1β production in Pstpip2cmo mice remain to be identified. In addition, the environmental factors driving IL-1β-dependent inflammatory bone erosion are unknown. Here we show that the intestinal microbiota of diseased Pstpip2cmo mice was characterized by an outgrowth of Prevotella. Notably, Pstpip2cmo mice that were fed a diet rich in fat and cholesterol maintained a normal body weight, but were markedly protected against inflammatory bone disease and bone erosion. Diet-induced protection against osteomyelitis was accompanied by marked reductions in intestinal Prevotella levels and significantly reduced pro-IL-1β expression in distant neutrophils. Furthermore, pro-IL-1β expression was also decreased in Pstpip2cmo mice treated with antibiotics, and in wild-type mice that were kept under germ-free conditions. We further demonstrate that combined deletion of caspases 1 and 8 was required for protection against IL-1β-dependent inflammatory bone disease, whereas the deletion of either caspase alone or of elastase or neutrophil proteinase 3 failed to prevent inflammatory disease. Collectively, this work reveals diet-associated changes in the intestinal microbiome as a crucial factor regulating inflammasome- and caspase-8-mediated maturation of IL-1β and osteomyelitis in Pstpip2cmo mice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lozano, R. et al. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet 380, 2095–2128 (2012)
Ferguson, P. J. et al. A missense mutation in pstpip2 is associated with the murine autoinflammatory disorder chronic multifocal osteomyelitis. Bone 38, 41–47 (2006)
Chitu, V. et al. Primed innate immunity leads to autoinflammatory disease in PSTPIP2-deficient cmo mice. Blood 114, 2497–2505 (2009)
Grosse, J. et al. Mutation of mouse Mayp/Pstpip2 causes a macrophage autoinflammatory disease. Blood 107, 3350–3358 (2006)
Lukens, J. R. et al. Critical role for inflammasome-independent IL-1β production in osteomyelitis. Proc. Natl Acad. Sci. USA 111, 1066–1071 (2014)
Cassel, S. L. et al. Inflammasome-independent IL-1beta mediates autoinflammatory disease in Pstpip2-deficient mice. Proc. Natl Acad. Sci. USA 111, 1072–1077 (2014)
Maslowski, K. M. & Mackay, C. R. Diet, gut microbiota and immune responses. Nature Immunol. 12, 5–9 (2011)
Montgomery, M. K. et al. Mouse strain-dependent variation in obesity and glucose homeostasis in response to high-fat feeding. Diabetologia 56, 1129–1139 (2013)
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006)
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014)
Dinarello, C. A. Interleukin-1 in the pathogenesis and treatment of inflammatory diseases. Blood. 117, 3720–3732 (2011)
Maelfait, J. et al. Stimulation of Toll-like receptor 3 and 4 induces interleukin-1β maturation by caspase-8. J. Exp. Med. 205, 1967–1973 (2008)
Gringhuis, S. I. et al. Dectin-1 is an extracellular pathogen sensor for the induction and processing of IL-1β via a noncanonical caspase-8 inflammasome. Nature Immunol. 13, 246–254 (2012)
Antonopoulos, C., El Sanadi, C., Kaiser, W. J., Mocarski, E. S. & Dubyak, G. R. Proapoptotic chemotherapeutic drugs induce noncanonical processing and release of IL-1β via caspase-8 in dendritic cells. J. Immunol. 191, 4789–4803 (2013)
Gurung, P. et al. FADD and caspase-8 mediate priming and activation of the canonical and noncanonical Nlrp3 inflammasomes. J. Immunol. 192, 1835–1846 (2014)
Man, S. M. et al. Salmonella infection induces recruitment of Caspase-8 to the inflammasome to modulate IL-1β production. J. Immunol. 191, 5239–5246 (2013)
Joosten, L. A. et al. Inflammatory arthritis in caspase 1 gene-deficient mice: contribution of proteinase 3 to caspase 1-independent production of bioactive interleukin-1β. Arthritis Rheum. 60, 3651–3662 (2009)
Karmakar, M., Sun, Y., Hise, A. G., Rietsch, A. & Pearlman, E. Cutting edge: IL-1β processing during Pseudomonas aeruginosa infection is mediated by neutrophil serine proteases and is independent of NLRC4 and caspase-1. J. Immunol. 189, 4231–4235 (2012)
Varfolomeev, E. E. et al. Targeted disruption of the mouse Caspase 8 gene ablates cell death induction by the TNF receptors, Fas/Apo1, and DR3 and is lethal prenatally. Immunity 9, 267–276 (1998)
Yeh, W. C. et al. FADD: essential for embryo development and signaling from some, but not all, inducers of apoptosis. Science 279, 1954–1958 (1998)
Zhang, J., Cado, D., Chen, A., Kabra, N. H. & Winoto, A. Fas-mediated apoptosis and activation-induced T-cell proliferation are defective in mice lacking FADD/Mort1. Nature 392, 296–300 (1998)
Kaiser, W. J. et al. RIP3 mediates the embryonic lethality of caspase-8-deficient mice. Nature 471, 368–372 (2011)
Oberst, A. et al. Catalytic activity of the caspase-8-FLIPL complex inhibits RIPK3-dependent necrosis. Nature 471, 363–367 (2011)
Shornick, L. P. et al. Mice deficient in IL-1β manifest impaired contact hypersensitivity to trinitrochlorobenzone. J. Exp. Med. 183, 1427–1436 (1996)
Kanneganti, T. D. et al. Bacterial RNA and small antiviral compounds activate caspase-1 through cryopyrin/Nalp3. Nature 440, 233–236 (2006)
Salmena, L. et al. Essential role for caspase 8 in T-cell homeostasis and T-cell-mediated immunity. Genes Dev. 17, 883–895 (2003)
Newton, K., Sun, X. & Dixit, V. M. Kinase RIP3 is dispensable for normal NF-κBs, signaling by the B-cell and T-cell receptors, tumor necrosis factor receptor 1, and Toll-like receptors 2 and 4. Mol. Cell. Biol. 24, 1464–1469 (2004)
Belaaouaj, A. et al. Mice lacking neutrophil elastase reveal impaired host defense against gram negative bacterial sepsis. Nature Med. 4, 615–618 (1998)
Kessenbrock, K. et al. Proteinase 3 and neutrophil elastase enhance inflammation in mice by inactivating antiinflammatory progranulin. J. Clin. Invest. 118, 2438–2447 (2008)
Acknowledgements
We thank D. Chaplin, V. Dixit, R. Flavell, D. Green and C. Pham for the supply of mutant mice. We thank J. Kim for helping to acquire and analyse the micro-computed tomography data. We thank S. Olsen, D. Roeber and the Genome Sequencing Facility in the Hartwell Center at St Jude Children’s Research Hospital for performing metagenomics sequencing of 16S rRNA. M.L. is supported by grants from Ghent University (BOF 01N02313 and 01J11113) and the European Research Council (Grant 281600). L.V.W. is a postdoctoral fellow of the Fund for Scientific Research-Flanders. This work was supported by: the National Institute of Arthritis and Musculoskeletal and Skin Diseases, part of the National Institutes of Health, under Award Number AR056296 (T.-D.K.); the National Cancer Institute, part of the National Institutes of Health, under Award Number CA163507 (T.-D.K.); the National Institute of Allergy and Infectious Diseases, part of the National Institutes of Health, under Award Number AI101935 (T.-D.K.); and ALSAC.
Author information
Authors and Affiliations
Contributions
J.R.L., M.L. and T.-D.K. designed the study; J.R.L., P.G., L.V.W., C.R.C. and S.R.B. performed experiments, and G.R.J. provided technical assistance. D.J.M. and R.A.C. analysed the 16S rRNA metagenomics data, and P.V. performed and analysed the histopathology data. J.R.L., M.L. and T.-D.K. analysed data and wrote the manuscript; T.-D.K. oversaw the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
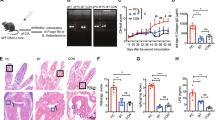
Extended Data Figure 1 Placing Pstpip2cmo mice on a high-fat and cholesterol diet limits the development of inflammatory bone disease.
a–e, Wild-type and Pstpip2cmo mutant mice were fed a LFD or HFD. Representative hind paw images (a) and representative pictures of popliteal lymph nodes (b) from wild-type, LFD Pstpip2cmo and HFD Pstpip2cmo mice at 12–14 weeks of age. c, d, Haematoxylin and eosin staining (original magnification, ×20) (c) and pathology scores (d) of tail samples from 12–14-week-old wild-type, LFD Pstpip2cmo and HFD Pstpip2cmo mice. Pathology scores were assigned in a blinded fashion by a veterinary pathologist based on the extent and severity of inflammation, osteolysis and osteogenesis. e, Representative immunostaining of neutrophils and macrophages in hind paw sections from 14–18-week-old Pstpip2cmo mice that were fed either a LFD or a HFD (original magnification, ×60). ***P < 0.001; Student’s t-test.
Extended Data Figure 2 Consumption of a HFD limits hyperinflammatory cytokine production in Pstpip2cmo mutant mice.
a, Wild-type and Pstpip2cmo mutant mice were fed a LFD or HFD for 12 weeks. Relative expression of Cxcl1 (wild type n = 8; LFD Pstpip2cmo n = 4; HFD Pstpip2cmo n = 9) and Il6 (wild type n = 11; LFD Pstpip2cmo n = 10; HFD Pstpip2cmo n = 8) in the hind paws was determined by qRT–PCR. The bar graphs depict combined data from two independent experiments. Data are shown as mean ± s.e.m. b, Wild-type and Pstpip2cmo mutant mice were fed a LFD or a HFD for 12 weeks and cytokines levels in the hind paws were determined by ELISA. Combined data are from two independent experiments. Each point represents an individual mouse, and the line represents the mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001; Student’s t-test.
Extended Data Figure 3 Placing Pstpip2cmo mice on a HFD does not cause abnormal weight gain, intestinal inflammation or extraintestinal translocation of commensal bacteria.
a, b, Wild-type BALB/cJ and Pstpip2cmo mice were fed ad libidum a LFD or a HFD. Body weight was measured in age-matched female (a) and male (b) mice at 12–16 weeks of age. Each point represents an individual mouse and the line represents the mean ± s.e.m. Data were combined from three independent experiments. c–e, Colon length (c), colitis score based on rectal bleeding and stool consistency (d) and representative haematoxylin and eosin-stained sections (original magnification, ×20) (e) of the intestinal tract of LFD- and HFD-fed Pstpip2cmo mice aged 14–18 weeks. f, Presence of commensal bacteria in the spleen, liver, mesenteric lymph nodes and bone of wild-type and diseased LFD-fed Pstpip2cmo mice was evaluated by Gram staining and 16S rDNA qPCR analysis of eubacteria.
Extended Data Figure 4 Dietary- and microbiota-associated factors influence the production of pro-IL-1β.
a, Footpad homogenates of 12–16-week-old wild-type, LFD-fed Pstpip2cmo and HFD-fed Pstpip2cmo mice were immunoblotted for IL-1β. Data are representative of three independent experiments. b, Footpads samples were collected from 10–14-week-old specific pathogen-free wild-type, germ-free wild-type and Pstpip2cmo × Il1b−/− mice and pro-IL-1β protein levels were determined by western blotting. c, CD45+ cells were isolated from the colons of germ-free wild-type mice and cells were left untreated or stimulated with LPS for 1 h. Relative Il1b mRNA expression levels were determined by qRT–PCR. Two biological replicates, with two technical replicates each.
Extended Data Figure 5 Co-housing does not alter disease progression in LFD-fed Pstpip2cmo mice.
a, Pstpip2cmo mice were treated with a cocktail of broad-spectrum antibiotics in their drinking water. Faecal samples were collected from wild-type (n = 5) and Pstpip2cmo mice that received either regular drinking water (n = 5) or antibiotics water (n = 11) 5–7 weeks later. Prevotella and Flexispira 16S rDNA copy numbers were quantified and normalized to total bacteria. The bar graphs depict the mean ± s.e.m. b, Faecal microbiota from diseased Pstpip2cmo mice was orally transplanted into wild-type mice (Pstpip2cmo microbiota ≫ wild type) and the incidence of inflammatory bone disease in control Pstpip2cmo and faecal transplantation mice was evaluated. c, d, Pstpip2cmo mice were singly housed or co-housed with wild-type (c) or Il1b-deficient Pstpip2cmo (d) mice. Clinical development of bone deformity and arthritic inflammation in hind paws and tails was monitored over time. **P < 0.01, ***P < 0.001; Student’s t-test.
Extended Data Figure 6 The neutrophil associated proteases elastase and proteinase 3 are not required for Pstpip2cmo-mediated bone disease.
a, Incidence of inflammatory bone disease in Pstpip2cmo, Pstpip2cmo × Elane−/−, Pstpip2cmo × Elane−/− Prtn3−/−and Pstpip2cmo × Il1b−/− mice. b, Representative footpad images from wild-type, Pstpip2cmo, Pstpip2cmo × Elane−/−, Pstpip2cmo × Elane−/− Prtn3−/−and Pstpip2cmo × Il1b−/− mice.
Extended Data Figure 7 Combined deletion of RIPK3 and caspase-8 does not provide protection against Pstpip2cmo-mediated osteomyelitis.
a, Incidence of osteomyelitic bone disease in wild-type, Pstpip2cmo, Pstpip2cmo × Il1b−/− and Pstpip2cmo × Ripk3−/− mice. b, Representative isosurface micro-computed tomography images of hind paw samples from 12–18-week-old Pstpip2cmo, Pstpip2cmo × Ripk3−/− and Pstpip2cmo × Ripk3−/− × Casp8−/− mice. c, Representative haematoxylin and eosin-stained sections of inflammatory caudal vertebrae bone lesions in diseased Pstpip2cmo, Pstpip2cmo × Ripk3−/− and Pstpip2cmo × Ripk3−/− × Casp8−/− mice (original magnification, ×4 (top) and ×10 (bottom)). d, qRT–PCR analysis of Il1b expression in footpads of wild-type (n = 7), Pstpip2cmo (n = 7) and Pstpip2cmo × Ripk3−/− × Casp8−/− × Casp1−/− (n = 7) mice aged 12–16 weeks. Data are expressed as mean ± s.e.m. of combined data from two independent experiments. **P < 0.01, ***P < 0.001; Student’s t-test.
Extended Data Figure 8 Reduced pro-IL-1β expression and IL-1β maturation in neutrophils isolated from HFD-fed Pstpip2cmo mice.
a, Wild-type, Pstpip2cmo and Pstpip2cmo × Il1b−/− bone-marrow-derived macrophages were left untreated or were primed with LPS for 3 h followed by stimulation with ATP (30 min) or silica (12 h), and IL-1β processing was evaluated by western blot. Data are representative of three independent experiments. b, Western blotting for pro-IL-1β in untreated neutrophils that were purified from wild-type, LFD-fed Pstpip2cmo and HFD-fed Pstpip2cmo mice. Data are representative of two independent experiments. c, d, Neutrophils (c) or macrophages (d) from wild-type, LFD-fed Pstpip2cmo and HFD-fed Pstpip2cmo mice were left untreated, or primed with LPS for 3 h and then stimulated with ATP (30 min) or silica (12 h), and IL-1β processing was evaluated by western blotting. Data are representative of two independent experiments.
Extended Data Figure 9 Depletion of neutrophils in anti-Ly6G treated Pstpip2cmo mutant mice.
Wild-type and Pstpip2cmo mice received either PBS or 500 μg per mouse anti-Ly6G antibody (clone IA8) by intraperitoneal injection every 4–5 days starting at 6 weeks of age. a–c, Two weeks after the first anti-Ly6G treatment, FACS analysis was performed on peripheral blood leukocytes (PBLs). a, Representative FACS plots of Gr-1 and CD11b expression on CD45.2+ gated cells. b, Enumeration of CD45.2+Gr-1hiCD11b+ neutrophils in equal volumes of peripheral blood. c, Numbers of T cells (CD45.2+TCRβ+), CD45.2+Gr-1−CD11b+ monocytes/macrophages and CD45.2+Gr-1intCD11b+ cells in equal volumes of peripheral blood. Each point represents an individual mouse and the line represents the mean ± s.e.m. ***P < 0.001; Student’s t-test.
Extended Data Figure 10 Dietary modulation of the intestinal microbiota composition drives autoinflammatory osteomyelitis by setting pro-IL-1β levels available for maturation by caspases 1 and 8.
Proposed model highlighting how dysbiosis and processing of IL-1β by caspases 1 and 8 contribute to the development of inflammatory bone disease.
Rights and permissions
About this article
Cite this article
Lukens, J., Gurung, P., Vogel, P. et al. Dietary modulation of the microbiome affects autoinflammatory disease. Nature 516, 246–249 (2014). https://doi.org/10.1038/nature13788
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13788
This article is cited by
-
NINJ1 mediates inflammatory cell death, PANoptosis, and lethality during infection conditions and heat stress
Nature Communications (2024)
-
Therapeutic effect of adipose-derived mesenchymal stem cells in a porcine model of abdominal sepsis
Stem Cell Research & Therapy (2023)
-
Regulated cell death pathways and their roles in homeostasis, infection, inflammation, and tumorigenesis
Experimental & Molecular Medicine (2023)
-
Triptolide induces PANoptosis in macrophages and causes organ injury in mice
Apoptosis (2023)
-
Results from a pilot study on the oral microbiome in children and adolescents with chronic nonbacterial osteomyelitis
Zeitschrift für Rheumatologie (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.