Abstract
The genetic architecture of autism spectrum disorder involves the interplay of common and rare variants and their impact on hundreds of genes. Using exome sequencing, here we show that analysis of rare coding variation in 3,871 autism cases and 9,937 ancestry-matched or parental controls implicates 22 autosomal genes at a false discovery rate (FDR) < 0.05, plus a set of 107 autosomal genes strongly enriched for those likely to affect risk (FDR < 0.30). These 107 genes, which show unusual evolutionary constraint against mutations, incur de novo loss-of-function mutations in over 5% of autistic subjects. Many of the genes implicated encode proteins for synaptic formation, transcriptional regulation and chromatin-remodelling pathways. These include voltage-gated ion channels regulating the propagation of action potentials, pacemaking and excitability–transcription coupling, as well as histone-modifying enzymes and chromatin remodellers—most prominently those that mediate post-translational lysine methylation/demethylation modifications of histones.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Data deposits
New data included in this manuscript have been deposited at dbGAP merged with our published data under accession number phs000298.v1.p1 and is available for download at (http://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000298.v1.p1).
Change history
12 November 2014
A minor change was made to the author affiliations.
References
Ronald, A. & Hoekstra, R. A. Autism spectrum disorders and autistic traits: a decade of new twin studies. Am. J. Med. Genet. B Neuropsychiatr. Genet. 156, 255–274 (2011)
Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science 316, 445–449 (2007)
Pinto, D. et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 466, 368–372 (2010)
Klei, L. et al. Common genetic variants, acting additively, are a major source of risk for autism. Mol. Autism 3, 9 (2012)
Gaugler, T. et al. Most inherited risk for autism resides with common variation. Nature Genet. 46, 881–885 (2014)
Yu, T. W. et al. Using whole-exome sequencing to identify inherited causes of autism. Neuron 77, 259–273 (2013)
Lim, E. T. et al. Rare complete knockouts in humans: population distribution and significant role in autism spectrum disorders. Neuron 77, 235–242 (2013)
Poultney, C. S. et al. Identification of small exonic CNV from whole-exome sequence data and application to autism spectrum disorder. Am. J. Hum. Genet. 93, 607–619 (2013)
Betancur, C. Etiological heterogeneity in autism spectrum disorders: more than 100 genetic and genomic disorders and still counting. Brain Res. 1380, 42–77 (2011)
Glessner, J. T. et al. Autism genome-wide copy number variation reveals ubiquitin and neuronal genes. Nature 459, 569–573 (2009)
Sanders, S. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012)
O’Roak, B. J. et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 338, 1619–1622 (2012)
O’Roak, B. J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250 (2012)
Neale, B. M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012)
Iossifov, I. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–299 (2012)
Willsey, A. J. et al. Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell 155, 997–1007 (2013)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genet. 43, 491–498 (2011)
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nature Methods 7, 248–249 (2010)
Samocha, K. E. et al. A framework for the interpretation of de novo mutation in human disease. Nature Genet. 46, 944–950 (2014)
He, X. et al. Integrated model of de novo and inherited genetic variants yields greater power to identify risk genes. PLoS Genet. 9, e1003671 (2013)
Girirajan, S. et al. Refinement and discovery of new hotspots of copy-number variation associated with autism spectrum disorder. Am. J. Hum. Genet. 92, 221–237 (2013)
Pinto, D. et al. Convergence of genes and cellular pathways dysregulated in autism spectrum disorders. Am. J. Hum. Genet. 94, 677–694 (2014)
Helsmoortel, C. et al. A SWI/SNF-related autism syndrome caused by de novo mutations in ADNP. Nature Genet. 46, 380–384 (2014)
Long, H. et al. Myo9b and RICS modulate dendritic morphology of cortical neurons. Cereb. Cortex 23, 71–79 (2013)
Yoon, K. J. et al. Mind bomb 1-expressing intermediate progenitors generate Notch signaling to maintain radial glial cells. Neuron 58, 519–531 (2008)
Smrt, R. D. et al. MicroRNA miR-137 regulates neuronal maturation by targeting ubiquitin ligase Mind bomb-1. Stem Cells 28, 1060–1070 (2010)
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nature Genet. 45, 1150–1159 (2013)
Robinson, E. B., Lichtenstein, P., Anckarsater, H., Happe, F. & Ronald, A. Examining and interpreting the female protective effect against autistic behavior. Proc. Natl Acad. Sci. USA 110, 5258–5262 (2013)
Jacquemont, S. et al. A higher mutational burden in females supports a “female protective model” in neurodevelopmental disorders. Am. J. Hum. Genet. 94, 415–425 (2014)
Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014)
Darnell, J. C. et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146, 247–261 (2011)
Ascano, M., Jr. et al. FMRP targets distinct mRNA sequence elements to regulate protein expression. Nature 492, 382–386 (2012)
Weyn-Vanhentenryck, S. M. et al. HITS-CLIP and integrative modeling define the Rbfox splicing-regulatory network linked to brain development and autism. Cell Rep. 6, 1139–1152 (2014)
Voineagu, I. et al. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature 474, 380–384 (2011)
Collins, M. O. et al. Molecular characterization and comparison of the components and multiprotein complexes in the postsynaptic proteome. J. Neurochem. 97 (suppl. 1). 16–23 (2006)
Liu, L. et al. DAWN: a framework to identify autism genes and subnetworks using gene expression and genetics. Mol. Autism 5, 22 (2014)
Tan, C. M., Chen, E. Y., Dannenfelser, R., Clark, N. R. & Ma’ayan, A. Network2Canvas: network visualization on a canvas with enrichment analysis. Bioinformatics 29, 1872–1878 (2013)
Vatta, M. et al. Genetic and biophysical basis of sudden unexplained nocturnal death syndrome (SUNDS), a disease allelic to Brugada syndrome. Hum. Mol. Genet. 11, 337–345 (2002)
Volkers, L. et al. Nav 1.1 dysfunction in genetic epilepsy with febrile seizures-plus or Dravet syndrome. Eur. J. Neurosci. 34, 1268–1275 (2011)
Scholl, U. I. et al. Somatic and germline CACNA1D calcium channel mutations in aldosterone-producing adenomas and primary aldosteronism. Nature Genet. 45, 1050–1054 (2013)
Khare, S. P. et al. HIstome–a relational knowledgebase of human histone proteins and histone modifying enzymes. Nucleic Acids Res. 40, D337–D342 (2012)
Feng, J. et al. Chronic cocaine-regulated epigenomic changes in mouse nucleus accumbens. Genome Biol. 15, R65 (2014)
Lachmann, A. et al. ChEA: transcription factor regulation inferred from integrating genome-wide ChIP-X experiments. Bioinformatics 26, 2438–2444 (2010)
Ronan, J. L., Wu, W. & Crabtree, G. R. From neural development to cognition: unexpected roles for chromatin. Nature Rev. Genet. 14, 347–359 (2013)
Rauch, A. et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet 380, 1674–1682 (2012)
Penzes, P., Cahill, M. E., Jones, K. A., VanLeeuwen, J. E. & Woolfrey, K. M. Dendritic spine pathology in neuropsychiatric disorders. Nature Neurosci. 14, 285–293 (2011)
Zoghbi, H. Y. Postnatal neurodevelopmental disorders: meeting at the synapse? Science 302, 826–830 (2003)
Acknowledgements
This work was supported by National Institutes of Health (NIH) grants U01MH100233, U01MH100209, U01MH100229 and U01MH100239 to the Autism Sequencing Consortium. Sequencing at Broad Institute was supported by NIH grants R01MH089208 (M.J.D.) and new sequencing by U54 HG003067 (S. Gabriel, E. Lander). Other funding includes NIH R01 MH089482, R37 MH057881 (B.D. and K.R.), R01 MH061009 (J.S.S.), UL1TR000445 (NCAT to VUMC); P50 HD055751 (E.H.C.); MH089482 (J.S.S.), NIH RO1 MH083565 and RC2MH089952 (C.A.W.), NIMH MH095034 (P.S), MH077139 (P.F. Sullivan); 5UL1 RR024975 and P30 HD15052. The DDD Study is funded by HICF-1009-003 and WT098051. UK10K is funded by WT091310. We also acknowledge The National Children’s Research Foundation, Our Lady’s Children’s Hospital, Crumlin; The Meath Foundation; AMNCH, Tallaght; The Health Research Board, Ireland and Autism Speaks, U.S.A. C.A.W. is an Investigator of the Howard Hughes Medical Institute. S.D.R., A.P.G., C.S.P., Y.K. and S.-C.F. are Seaver fellows, supported by the Seaver foundation. A.P.G. is also supported by the Charles and Ann Schlaifer Memorial Fund. P.F.B. is supported by a UK National Institute for Health Research (NIHR) Senior Investigator award and the NIHR Biomedical Research Centre in Mental Health at the South London & Maudsley Hospital. A.C. is supported by María José Jove Foundation and the grant FIS PI13/01136 of the Strategic Action from Health Carlos III Institute (FEDER). This work was supported in part through the computational resources and staff expertise provided by the Department of Scientific Computing at the Icahn School of Medicine at Mount Sinai. We acknowledge the assistance of D. Hall and his team at National Database for Autism Research. We thank Jian Feng for providing a list of targets of both RBFOX1 and H3K4me3. We thank M. Potter for data coordination; K. Moore and J. Reichert for technical assistance; and, S. Lindsay for helping with molecular validation. We acknowledge the clinicians and organizations that contributed to samples used in this study. Finally, we are grateful to the many families whose participation made this study possible.
Author information
Authors and Affiliations
Consortia
Contributions
Lists of participants appear in the Supplementary Information.
Study conception and design: J.D.B., D.J.C., M.J.D., S.D.R., B.D., M.F., A.P.G., X.H., T.L., C.S.P., K.Ro., M.W.S. and M.E.Z. Data analysis: J.C.B., P.F.B., J.D.B., J.C., A.E.C, D.J.C., M.J.D., S.D.R., B.D., M.F., S.-C.F., A.P.G., X.H., L.K., J.K., Y.K., L.L., A.M., C.S.P., S.P., K.Ro., K.S., C.S., T.S., C.St., S.W., L.W. and M.E.Z. Contribution of samples, WES data or analytical tools: B.A., J.C.B., M.B., P.F.B., J.D.B., J.C., N.G.C., A.C., M.H.C., A.G.C., A.E.C, H.C., E.L.C., L.C., S.R.C., D.J.C., M.J.D., G.D., S.D.R., B.D., E.D., B.A.F., C.M.F., M.F., L.G., E.G., M.G., A.P.G., S.J.G., X.H., R.H., C.M.H., I.I.-L., P.J.G., H.K., S.M.K., L.K., A.K., J.K., Y.K., I.L., J.L., T.Le., C.L., L.L., A.M., C.R.M., A.L.M., B.N., M.J.O., N.O., A.P., M.P., J.R.P., C.S.P., S.P., K.P., D.R., K.R., A.R., K.Ro., A.S., M.S., K.S., S.J.S., C.S., G.D.S., S.W.S., M.S.-R., T.S., P.S., D.S., M.W.S., C.St., J.S.S., P.Sz., K.T., O.V., A.V., S.W., C.A.W., L.W., L.A.W., J.A.W., T.W.Y., R.K.C.Y., M.E.Z. Writing of the paper: J.C.B., J.D.B., E.H.C., D.J.C., M.J.D., S.D.R., B.D., M.G., A.P.G., X.H., C.S.P., K.Ro., S.W.S., M.E.Z. Leads of ASC committees: J.D.B., E.H.C., M.J.D., B.D., M.G., K.Ro., M.W.S., J.S.S., M.E.Z. Administration of ASC: J.M.B.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
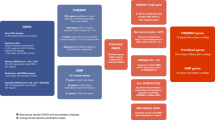
Extended Data Figure 1 Workflow of the study.
The workflow began with 16 sample sets, as listed in Supplementary Table 1. DNA was obtained, and exomes were captured and sequenced. After variant calling, quality control was performed: duplicate subjects and incomplete families were removed and subjects with extreme genotyping, de novo, or variant rates were removed. Following cleaning, 3,871 subjects with ASD remained. Analysis proceeded separately for SNVs and indels, and CNVs. De novo and transmission/non-transmission variants were obtained for trio data (published de novo variants from 825 trios11,13,14,15 were incorporated). This led to the TADA analysis, which found 33 ASD risk genes with an FDR < 0.1; and 107 with an FDR < 0.3. CNVs were called in 2,305 ASD subjects. BAM, binary alignment/map; MAF, minor allele frequency.
Extended Data Figure 2 Expected number of ASD genes discovered as a function of sample size.
The multiple LoF test (red) is a restricted version of TADA that uses only the de novo LoF data. TADA (blue) models de novo LoF, de novo Mis3, LoF variants transmitted/not transmitted and LoF variants observed in case-control samples. The sample size (n) indicates either n trios for which we record de novo and transmitted variation (TADA), or n trios for which we record only de novo events (multiple LoF), plus n cases and n controls.
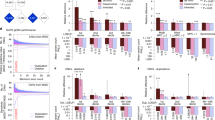
Extended Data Figure 3 Heat map of the numbers of variants used in TADA analysis from each data set in genes with an FDR < 0.3.
Left, variants in affected subjects; right, unaffected subjects. For the counts, we only included de novo LoF and Mis3 variants, transmitted/untransmitted and case-control LoF variants. These variant counts are normalized by the length of coding regions of each gene and sample size of each data set (|trio| + |case| for the left, |trio| + |control| for the right). Description of the samples can be found in Supplementary Table 1.
Extended Data Figure 4 Genome browser view of the CNV deletions identified in ASD-affected subjects.
The deletions are displayed in red if with unknown inheritance, in grey if inherited, and in black in unaffected subjects. Deletions in parents are not shown. For deletions within a single gene, all splicing isoforms are shown.
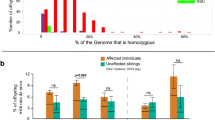
Extended Data Figure 5 Frequency of variants by gender.
Frequency of de novo (dn) and transmitted (Tr) variants per sample in males (black) and females (white) for genes with an FDR < 0.1 (top row), FDR < 0.3 (middle row), or all TADA genes (bottom row). The P values were determined by one-tailed permutation tests (*P < 0.05; **P < 0.01; ***P < 0.01).
Extended Data Figure 6 Enrichment terms for the four clusters identified by protein–protein interaction networks.
P values calculated using mouse-genome-informatics–mammalian-phenotype (MGI_Mammalian phenotype, blue), Kyoto encyclopaedia of genes and genomes (KEGG) pathways (red), and gene ontology biological processes (yellow) are indicated.
Extended Data Figure 7 De novo variants in SET lysine methyltransferases and jumonji lysine demethylases.
Mis3 variants are in black, LoF in red, and variants identified in other disorders in grey (Fig. 5). ARID, AT-rich interacting domain; AWS, associated with SET domain; BAH, bromo adjacent homology; bromo, bromodomain; FYR C, FY-rich C-terminal domain; FYR N, FY-rich N-terminal domain; HiMG, high mobility group box; JmjC, jumonji C domain; JmjN, jumonji N domain; PHD, plant homeodomain; PWWP, Pro-Trp-Trp-Pro domain; SET, Su(var)3-9, enhancer-of-zeste, trithorax domain.
Extended Data Figure 8 Transcription regulation network of TADA genes only.
Edges indicate transcription regulators (source nodes) and their gene targets (target nodes) based on the ChEA network.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Notes and additional references. (PDF 1894 kb)
Supplementary Table 1
This file contains Supplementary Table 1. (XLSX 31 kb)
Supplementary Table 2
This file contains Supplementary Table 2. (XLSX 2333 kb)
Supplementary Table 3
This file contains Supplementary Table 3. (XLSX 895 kb)
Supplementary Table 4
This file contains Supplementary Table 4. (XLSX 32 kb)
Supplementary Table 5
This file contains Supplementary Table 5. (XLSX 35 kb)
Supplementary Table 6
This file contains Supplementary Table 6. (XLSX 34 kb)
Rights and permissions
About this article
Cite this article
De Rubeis, S., He, X., Goldberg, A. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014). https://doi.org/10.1038/nature13772
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13772
This article is cited by
-
Detection of autism spectrum disorder-related pathogenic trio variants by a novel structure-based approach
Molecular Autism (2024)
-
Pathogenic/likely pathogenic mutations identified in Vietnamese children diagnosed with autism spectrum disorder using high-resolution SNP genotyping platform
Scientific Reports (2024)
-
Rare genetic brain disorders with overlapping neurological and psychiatric phenotypes
Nature Reviews Neurology (2024)
-
The complex etiology of autism spectrum disorder due to missense mutations of CHD8
Molecular Psychiatry (2024)
-
Transcriptomic dysregulation and autistic-like behaviors in Kmt2c haploinsufficient mice rescued by an LSD1 inhibitor
Molecular Psychiatry (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.