Abstract
To repair double-strand DNA breaks by homologous recombination, the 5′-terminated DNA strand must first be resected, which generates 3′ single-stranded DNA overhangs. Genetic evidence suggests that this process is initiated by the Mre11–Rad50–Xrs2 (MRX) complex1,2,3. However, its involvement was puzzling, as the complex possesses exonuclease activity with the opposite (3′ to 5′) polarity from that required for homologous recombination4,5. Consequently, a bidirectional model has been proposed6,7,8 whereby dsDNA is first incised endonucleolytically and MRX then proceeds back to the dsDNA end using its 3′ to 5′ exonuclease. The endonuclease creates entry sites for Sgs1–Dna2 and/or Exo1, which then carry out long-range resection in the 5′ to 3′ direction. However, the identity of the endonuclease remained unclear. Using purified Saccharomyces cerevisiae proteins, we show that Sae2 promotes dsDNA-specific endonuclease activity by the Mre11 subunit within the MRX complex. The endonuclease preferentially cleaves the 5′-terminated dsDNA strand, which explains the polarity paradox. The dsDNA end clipping is strongly stimulated by protein blocks at the DNA end, and requires the ATPase activity of Rad50 and physical interactions between MRX and Sae2. Our results suggest that MRX initiates dsDNA break processing by dsDNA endonuclease rather than exonuclease activity, and that Sae2 is the key regulator of this process. These findings demonstrate a probable mechanism for the initiation of dsDNA break processing in both vegetative and meiotic cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Mimitou, E. P. & Symington, L. S. Sae2, Exo1 and Sgs1 collaborate in DNA double-strand break processing. Nature 455, 770–774 (2008)
Zhu, Z., Chung, W. H., Shim, E. Y., Lee, S. E. & Ira, G. Sgs1 helicase and two nucleases Dna2 and Exo1 resect DNA double-strand break ends. Cell 134, 981–994 (2008)
Gravel, S., Chapman, J. R., Magill, C. & Jackson, S. P. DNA helicases Sgs1 and BLM promote DNA double-strand break resection. Genes Dev. 22, 2767–2772 (2008)
Trujillo, K. M. & Sung, P. DNA structure-specific nuclease activities in the Saccharomyces cerevisiae Rad50*Mre11 complex. J. Biol. Chem. 276, 35458–35464 (2001)
Trujillo, K. M. et al. Yeast Xrs2 binds DNA and helps target Rad50 and Mre11 to DNA ends. J. Biol. Chem. 278, 48957–48964 (2003)
Shibata, A. et al. DNA double-strand break repair pathway choice is directed by distinct MRE11 nuclease activities. Mol. Cell 53, 7–18 (2014)
Garcia, V., Phelps, S. E., Gray, S. & Neale, M. J. Bidirectional resection of DNA double-strand breaks by Mre11 and Exo1. Nature 479, 241–244 (2011)
Zakharyevich, K. et al. Temporally and biochemically distinct activities of Exo1 during meiosis: double-strand break resection and resolution of double Holliday junctions. Mol. Cell 40, 1001–1015 (2010)
Bressan, D. A., Olivares, H. A., Nelms, B. E. & Petrini, J. H. Alteration of N-terminal phosphoesterase signature motifs inactivates Saccharomyces cerevisiae Mre11. Genetics 150, 591–600 (1998)
Lengsfeld, B. M., Rattray, A. J., Bhaskara, V., Ghirlando, R. & Paull, T. T. Sae2 is an endonuclease that processes hairpin DNA cooperatively with the Mre11/Rad50/Xrs2 complex. Mol. Cell 28, 638–651 (2007)
Deshpande, R. A. et al. ATP-driven Rad50 conformations regulate DNA tethering, end resection, and ATM checkpoint signaling. EMBO J. 33, 482–500 (2014)
Lammens, K. et al. The Mre11:Rad50 structure shows an ATP-dependent molecular clamp in DNA double-strand break repair. Cell 145, 54–66 (2011)
Chen, L. et al. Effect of amino acid substitutions in the Rad50 ATP binding domain on DNA double strand break repair in yeast. J. Biol. Chem. 280, 2620–2627 (2005)
Hopfner, K. P. et al. Structural biochemistry and interaction architecture of the DNA double-strand break repair Mre11 nuclease and Rad50-ATPase. Cell 105, 473–485 (2001)
Keeney, S., Giroux, C. N. & Kleckner, N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88, 375–384 (1997)
Alani, E., Padmore, R. & Kleckner, N. Analysis of wild-type and rad50 mutants of yeast suggests an intimate relationship between meiotic chromosome synapsis and recombination. Cell 61, 419–436 (1990)
Neale, M. J., Pan, J. & Keeney, S. Endonucleolytic processing of covalent protein-linked DNA double-strand breaks. Nature 436, 1053–1057 (2005)
Rattray, A. J., McGill, C. B., Shafer, B. K. & Strathern, J. N. Fidelity of mitotic double-strand-break repair in Saccharomyces cerevisiae: a role for SAE2/COM1. Genetics 158, 109–122 (2001)
Langerak, P., Mejia-Ramirez, E., Limbo, O. & Russell, P. Release of Ku and MRN from DNA ends by Mre11 nuclease activity and Ctp1 is required for homologous recombination repair of double-strand breaks. PLoS Genet. 7, e1002271 (2011)
Mimitou, E. P. & Symington, L. S. Ku prevents Exo1 and Sgs1-dependent resection of DNA ends in the absence of a functional MRX complex or Sae2. EMBO J. 29, 3358–3369 (2010)
Shim, E. Y. et al. Saccharomyces cerevisiae Mre11/Rad50/Xrs2 and Ku proteins regulate association of Exo1 and Dna2 with DNA breaks. EMBO J. 29, 3370–3380 (2010)
Liu, C., Pouliot, J. J. & Nash, H. A. Repair of topoisomerase I covalent complexes in the absence of the tyrosyl-DNA phosphodiesterase Tdp1. Proc. Natl Acad. Sci. USA 99, 14970–14975 (2002)
Hartsuiker, E., Neale, M. J. & Carr, A. M. Distinct requirements for the Rad32(Mre11) nuclease and Ctp1(CtIP) in the removal of covalently bound topoisomerase I and II from DNA. Mol. Cell 33, 117–123 (2009)
Hopkins, B. B. & Paull, T. T. The P. furiosus Mre11/Rad50 complex promotes 5′ strand resection at a DNA double-strand break. Cell 135, 250–260 (2008)
Uetz, P. et al. A comprehensive analysis of protein–protein interactions in Saccharomyces cerevisiae. Nature 403, 623–627 (2000)
Kim, H. S. et al. Functional interactions between Sae2 and the Mre11 complex. Genetics 178, 711–723 (2008)
Huertas, P., Cortes-Ledesma, F., Sartori, A. A., Aguilera, A. & Jackson, S. P. CDK targets Sae2 to control DNA-end resection and homologous recombination. Nature 455, 689–692 (2008)
Akamatsu, Y. et al. Molecular characterization of the role of the Schizosaccharomyces pombe nip1+/ctp1+ gene in DNA double-strand break repair in association with the Mre11-Rad50-Nbs1 complex. Mol. Cell. Biol. 28, 3639–3651 (2008)
Penkner, A. et al. A conserved function for a Caenorhabditis elegans Com1/Sae2/CtIP protein homolog in meiotic recombination. EMBO J. 26, 5071–5082 (2007)
Cejka, P. & Kowalczykowski, S. C. The full-length Saccharomyces cerevisiae Sgs1 protein is a vigorous DNA helicase that preferentially unwinds Holliday junctions. J. Biol. Chem. 285, 8290–8301 (2010)
Cannavo, E., Cejka, P. & Kowalczykowski, S. C. Relationship of DNA degradation by Saccharomyces cerevisiae exonuclease 1 and its stimulation by RPA and Mre11-Rad50-Xrs2 to DNA end resection. Proc. Natl Acad. Sci. USA 110, E1661–E1668 (2013)
Levikova, M., Klaue, D., Seidel, R. & Cejka, P. Nuclease activity of Saccharomyces cerevisiae Dna2 inhibits its potent DNA helicase activity. Proc. Natl Acad. Sci. USA 110, E1992–E2001 (2013)
Nicolette, M. L. et al. Mre11-Rad50-Xrs2 and Sae2 promote 5′ strand resection of DNA double-strand breaks. Nature Struct. Mol. Biol. 17, 1478–1485 (2010)
Lisby, M., Barlow, J. H., Burgess, R. C. & Rothstein, R. Choreography of the DNA damage response: spatiotemporal relationships among checkpoint and repair proteins. Cell 118, 699–713 (2004)
Acknowledgements
We thank L. Ranjha, R. Anand, L. Mlejnkova and M. Levikova for assistance and comments on the manuscript. This work was supported by Swiss National Science Foundation Grant PP00P3 133636 to P.C.
Author information
Authors and Affiliations
Contributions
P.C. and E.C. designed the study. E.C. and P.C. carried out the experiments. P.C. and E.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Purification of wild-type Mre11–Rad50–Xrs2 (MRX) and Sae2.
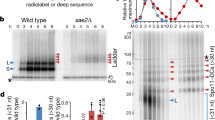
a, Purified MRX used in this study. Gel was stained with Coomassie brilliant blue. b, Nuclease activity of MRX (10 nM) on 5′-labelled dsDNA substrate. Products were separated on a denaturing gel. MRX gradually shortened dsDNA with a 5′ 32P label. c, Nuclease activity of MRX (10 nM) on 3′-labelled dsDNA substrate. MRX directly released the radioactive label from a 3′ 32P-labelled DNA substrate, showing that it is a 3′–5′ exonuclease4. d, Nuclease activity of MRX and its dependence on manganese, ATP and RPA, as indicated. The MRX exonuclease requires manganese (5 mM), is moderately inhibited by ATP (1 mM), and is not affected by RPA (23 nM). e, A scheme of the Sae2 construct. Sae2 contains an N-terminal MBP tag and a C-terminal His-tag (10×His). f, Representative purification of Sae2. Gel was stained with Coomassie brilliant blue. MBP affinity tag was cleaved off during protein purification. MBP, maltose-binding protein; PP, PreScission protease.
Extended Data Figure 2 MRX dsDNA endonuclease is promoted by Sae2 and unaffected by RPA.
a, Quantitation of the data such as from Fig. 1a. Averages shown, n = 2, error, s.e.m. b, Nuclease assay (15 min) was carried out with MRX (20 nM) and a range of Sae2 concentrations, as indicated. The exonuclease of MRX was unaffected by Sae2, but endonuclease cleavage increased with Sae2 concentration. c, Nuclease activity of MRX and Sae2 on dsDNA is not affected by the single-strand DNA binding protein RPA. Nuclease assay was carried out as in b, but with RPA (23 nM). d, Quantitation of the data such as from b and c. Averages shown, n = 2; error bars, s.e.m.
Extended Data Figure 3 Purification of MRX variants.
a, Polyacrylamide gel electrophoresis showing representative purifications. Gel was stained with Coomassie brilliant blue. b, Nuclease assay was performed with 5′-labelled dsDNA substrate and either wild-type or nuclease-deficient M(H125L;D126V)RX variant (MRX-nd). As expected, the mutant MRX possessed no activity, indicating that the nuclease is intrinsic to Mre11. c, Purified MRX mutants used in this study. MR(K40A)X is expected not to bind ATP; MR(K40R)X is expected to bind but not to hydrolyse ATP; MR(K81I)X is a Rad50S MRX variant.
Extended Data Figure 4 Sae2 does not show nuclease activity, and does not promote MRX nuclease on hairpins.
a, Recombinant Sae2 was assayed on a dsDNA substrate in the presence of either magnesium (5 mM) or manganese (5 mM), with or without RPA (23 nM), as indicated. Free label, carryover of [32P]ATP from the labelling reaction, marks the position of the smallest possible product resulting from potential nuclease activity. Samples were analysed on a 10% native polyacrylamide gel. b, Recombinant Sae2 was assayed on a Y-structure DNA substrate either in the presence of magnesium (5 mM) or manganese (5 mM), with or without RPA (23 nM), as indicated. Free label, carryover of [32P]ATP from the labelling reaction, marks the position of the smallest possible product resulting from potential nuclease activity. Samples were analysed on a 10% native polyacrylamide gel. c, Recombinant Sae2 was assayed on a HL-3 hairpin DNA substrate either in the presence of magnesium (5 mM) or manganese (5 mM), with or without RPA (23 nM), as indicated. Samples were analysed on a 15% denaturing polyacrylamide gel. d, Nuclease assay was performed with MRX and Sae2 on HP-2 DNA, as indicated. Samples were analysed on a 15% denaturing polyacrylamide gel. Sae2 does not promote endonuclease of MRX on HP-2 DNA substrate. e, Nuclease assay was performed with MRX and Sae2 on HL-3 hairpin DNA, as indicated. Samples were analysed on a 15% denaturing polyacrylamide gel. Sae2 does not promote endonuclease of MRX on HL-3 DNA substrate.
Extended Data Figure 5 Analysis of Sae2 and MRX endonuclease activity on 50-bp single-blocked dsDNA substrate.
a, Nuclease assay was performed with ADP or non-hydrolysable ATP analogue ATPγS, and MRX and/or Sae2, as indicated. Neither ADP nor ATPγS supported the endonuclease activity, suggesting that ATP hydrolysis is essential. b, Nuclease assay was performed with MRX and/or Sae2 and various concentrations of magnesium and manganese, as indicated. Higher concentration of magnesium than manganese is required for the endonuclease of MRX stimulated by Sae2. Endo cleavage (%), average percentage of endonucleolytic products, on the basis of two independent experiments.
Extended Data Figure 6 Analysis of Sae2 and MRX-dependent endonuclease activity.
a, Nuclease assay was performed with a 5′-labelled 100-bp-long dsDNA substrate in the absence of a protein block. MRX and Sae2 concentrations were used as indicated. Sae2 promoted MRX endonuclease even in the absence of a protein block; however, the reaction was inefficient and contrast of the image had to be enhanced to visualize the degradation products. The cleavage occurred ∼10 nucleotides away from the DNA end, which is different from protein-blocked substrates, which were cleaved typically ∼15–20 nucleotides away from the end. b, Kinetic analysis of MRX and Sae2 endonuclease activity. The preferred position of cleavage is located ∼50 nucleotides from the 3′ end, and ∼20 nucleotides from the 5′ end. c, Nuclease assay (15 min) was performed with indicated concentrations of MRX and Sae2. The extent of endonuclease cleavage is dependent on concentrations of both Sae2 and MRX. d, Quantification of the data from b. Averages shown, n = 2, error bars, s.e.m. e, f, Quantification of the data from c. Averages shown, n = 2, error bars, s.e.m. g, Experiment as in Fig. 2, but with avidin instead of streptavidin. Both avidin and streptavidin promote Sae2 and MRX endonuclease to a similar extent, showing that there is no need for a specific interaction between Sae2–MRX and the protein block. h, i, Nuclease assays were performed with recombinant proteins as indicated. MX, Mre11–Xrs2. Sae2 promotes only the endonuclease of MRX. *, Nucleolytic product resulting from Exo1 activity, independent of Sae2.
Extended Data Figure 7 Sae2 does not promote ssDNA endonuclease activity of MRX.
a, Circular M13 ssDNA was used as a substrate in a nuclease assay with MRX and/or Sae2, as indicated. Sae2 did not affect the ssDNA endonuclease of MRX. Reaction products were analysed on 1% agarose gel and stained with Gel red (Invitrogen). b, The ssDNA endonuclease activity of MRX is dependent on manganese and inhibited by saturating concentrations of RPA (1.5 μM). Mg2+ only, 5 mM magnesium acetate, no manganese; Mn2+ only, 5 mM manganese acetate, no magnesium. Reaction products were analysed on 1% agarose gel and stained with Gel red.
Extended Data Figure 8 Sae2 and MRX endonuclease preferentially cleave 5′-terminated DNA strand.
a, Nuclease assay was performed with a 5′ 32P-labelled 2.7-kb-long dsDNA substrate either without streptavidin (lanes 2–5) or with streptavidin (lanes 7–10), and MRX and Sae2, as indicated, for 60 min. The 2.7-kb-long substrate was prepared by reacting pATTP-S plasmid with annealed labelled oligonucleotides and ΦC31 integrase, as described in Methods. Reaction products were separated on a 15% denaturing polyacrylamide gel. Unprocessed DNA substrate did not enter the gel and remained trapped in the wells. MRX alone has the capacity to cleave dsDNA endonucleolytically at various distances from the 5′ end (lane 3), in agreement with previous reports24,33. This endonuclease activity is not affected by the protein block (compare lanes 3 and 8). Sae2 promotes endonucleolytic cleavage specifically near the protein-blocked DNA end (lane 10, indicated by red arrows). b, Assay as in a, but with a 3′-labelled DNA substrate. No endonuclease activity of MRX and Sae2 near the 3′ end was detected. c, Nuclease assay as in Fig. 3a, but with a DNA substrate of 100 bp in length (instead of 70 bp). Concerted action of MRX and Sae2 resulted in DNA cleavage ∼15–20 nucleotides away from the streptavidin-blocked 5′ DNA end. The position of the cleavage was identical for both 100- and 70-bp-long DNA substrates (compare with Fig. 3a), suggesting that the protein-blocked DNA end directs the position of cleavage by MRX and Sae2. d, A cartoon depicting the position of endonucleolytic cleavage by MRX–Sae2. For simplicity, the MRX complex is depicted as a monomer.
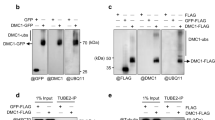
Extended Data Figure 9 Sae2 interacts with Mre11 and Xrs2 subunits of the MRX complex.
a, Amylose pull-down was carried out with MBP–Sae2 and Xrs2 or MBP and Xrs2. Xrs2 bound to MBP–Sae2 (lane 2) but not to MBP (lane 3), showing that Xrs2 binds Sae2 but not the MBP tag or the amylose resin. We point out that the interaction was very weak, and the amount of Xrs2 we pulled down with MBP–Sae2 was near the limit of detection by silver staining. Lane 4, 63 ng of recombinant Xrs2 was loaded as a control. Samples in lanes 2 and 4 were treated with PreScission protease before gel analysis. MBP1, maltose-binding protein expressed in E. coli; MBP2, maltose-binding protein resulting from cleavage with PreScission protease; PP, PreScission protease. b, Amylose pull-down was carried out with MBP–Sae2 and Mre11 or MBP and Mre11. Mre11 bound to MBP–Sae2 (lane 5), but not to MBP (lane 4), showing that Mre11 binds Sae2 but not the MBP tag or the amylose resin. Lane 3, 10 ng of recombinant Mre11 was loaded as a control. Lane 2, amylose pull-down was carried out with MBP–Sae2 alone (without Mre11). The interaction between Sae2 and Mre11 is very weak, as the amount of Mre11 we pulled down with MBP–Sae2 is very small. Samples in lanes 2, 3 and 5 were treated with PreScission protease. The band migrating just below Mre11 (indicated by an asterisk) is likely to be residual uncleaved MBP–Sae2. The image in the upper panel was stretched vertically for visualization purposes. MBP1, maltose-binding protein expressed in E. coli; MBP2, maltose-binding protein resulting from cleavage with PreScission protease; PP, PreScission protease. c, Amylose pull-down was carried out with MBP–Sae2 and variants of MRX (lanes 3–6). Lanes 7–10, 250 ng of the respective MRX preparations was loaded as a control. Control Sae2, 120 ng recombinant Sae2. Sae2 interacts with both MRX variants deficient in ATP binding and/or hydrolysis, as expected. Sae2 also interacts with MR(K81I)X, indicating that the defects in the activation of the endonuclease of the Rad50S MRX variant by Sae2 cannot simply be explained by a lack of interaction, which is in accordance with proper Sae2 recruitment to double strand breaks in rad50s mutants34. However, since Sae2 likely interacts with multiple subunits of the MRX complex, we cannot exclude a defect in a subset of the interaction sites, which may abrogate the functional interplay between MR(K81I)X and Sae2.
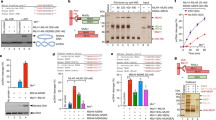
Extended Data Figure 10 C-terminal region of Sae2 is critical for the stimulation of the MRX endonuclease.
a, A scheme depicting Sae2 truncation mutants analysed in this study. b, Nuclease assay (with 50-mer single-blocked DNA) was performed with MRX and N or C-terminal truncation mutants of Sae2, as indicated. N-terminal region of Sae2 is dispensable, while C-terminal region is essential for the stimulation of the MRX endonuclease. c, A scheme depicting Sae2 mutants analysed in this study. d, e, Nuclease assay (with 50-mer single-blocked DNA) was performed with MRX and Sae2 variants, as indicated. Endo cleavage (% of WT), average percentage of endonucleolytic products, normalized to wild-type Sae2, on the basis of two independent experiments. f, Electrophoretic mobility shift assay was used to determine the capacity of Sae2 variants to bind DNA. The results (average) are based on disappearance of the substrate band; n = 3, error bars, s.e.m. g, Amylose pull-down was carried out with MBP-tagged Sae2 variants and MRX. The presence of MRX in the pull-downs was detected by western blotting using anti-Flag antibody against Xrs2. Sae2 was detected by silver staining. h, Nuclease assay (with 50-mer single-blocked DNA) was performed with MRX and Sae2 either mock-treated (incubated with protein phosphatase 1 reaction buffer for 15 min at 30 °C) or protein phosphatase 1-treated Sae2 (New England Biolabs, 1.25 U per 2.5 μg recombinant Sae2, 15 min at 30 °C). Endo cleavage (%), average percentage of endonucleolytic products, on the basis of three independent experiments. Treatment of Sae2 with protein phosphatase 1 leads to a reduction of Sae2 capacity to promote MRX endonucleolytic activity. This suggests that Sae2 purified from Sf9 cells is phosphorylated, and this post-translational modification promotes its capacity to stimulate MRX endonuclease.
Rights and permissions
About this article
Cite this article
Cannavo, E., Cejka, P. Sae2 promotes dsDNA endonuclease activity within Mre11–Rad50–Xrs2 to resect DNA breaks. Nature 514, 122–125 (2014). https://doi.org/10.1038/nature13771
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13771
This article is cited by
-
Molecular mechanisms and regulation of recombination frequency and distribution in plants
Theoretical and Applied Genetics (2024)
-
MUS81 cleaves TOP1-derived lesions and other DNA–protein cross-links
BMC Biology (2023)
-
Double-strand breaks induce inverted duplication chromosome rearrangements by a DNA polymerase δ-dependent mechanism
Nature Communications (2023)
-
Genome-wide analysis of DNA-PK-bound MRN cleavage products supports a sequential model of DSB repair pathway choice
Nature Communications (2023)
-
Mre11-Rad50 oligomerization promotes DNA double-strand break repair
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.