Abstract
Saturation mutagenesis1,2—coupled to an appropriate biological assay—represents a fundamental means of achieving a high-resolution understanding of regulatory3 and protein-coding4 nucleic acid sequences of interest. However, mutagenized sequences introduced in trans on episomes or via random or “safe-harbour” integration fail to capture the native context of the endogenous chromosomal locus5. This shortcoming markedly limits the interpretability of the resulting measurements of mutational impact. Here, we couple CRISPR/Cas9 RNA-guided cleavage6 with multiplex homology-directed repair using a complex library of donor templates to demonstrate saturation editing of genomic regions. In exon 18 of BRCA1, we replace a six-base-pair (bp) genomic region with all possible hexamers, or the full exon with all possible single nucleotide variants (SNVs), and measure strong effects on transcript abundance attributable to nonsense-mediated decay and exonic splicing elements. We similarly perform saturation genome editing of a well-conserved coding region of an essential gene, DBR1, and measure relative effects on growth that correlate with functional impact. Measurement of the functional consequences of large numbers of mutations with saturation genome editing will potentially facilitate high-resolution functional dissection of both cis-regulatory elements and trans-acting factors, as well as the interpretation of variants of uncertain significance observed in clinical sequencing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Myers, R. M., Tilly, K. & Maniatis, T. Fine structure genetic analysis of a beta-globin promoter. Science 232, 613–618 (1986)
Cunningham, B. C. & Wells, J. A. High-resolution epitope mapping of hGH-receptor interactions by alanine-scanning mutagenesis. Science 244, 1081–1085 (1989)
Patwardhan, R. P. et al. High-resolution analysis of DNA regulatory elements by synthetic saturation mutagenesis. Nature Biotechnol. 27, 1173–1175 (2009)
Fowler, D. M. et al. High-resolution mapping of protein sequence-function relationships. Nature Methods 7, 741–746 (2010)
Botstein, D. & Shortle, D. Strategies and applications of in vitro mutagenesis. Science 229, 1193–1201 (1985)
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012)
Gibson, T. J., Seiler, M. & Veitia, R. A. The transience of transient overexpression. Nature Methods 10, 715–721 (2013)
Gaj, T., Gersbach, C. A. & Barbas, C. F., III ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 31, 397–405 (2013)
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013)
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013)
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013)
Mazoyer, S. et al. A BRCA1 nonsense mutation causes exon skipping. Am. J. Hum. Genet. 62, 713–715 (1998)
Sander, J. D. & Joung, J. K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nature Biotechnol. 32, 347–355 (2014)
Ke, S. et al. Quantitative evaluation of all hexamers as exonic splicing elements. Genome Res. 21, 1360–1374 (2011)
Zhang, J., Kuo, C. C. & Chen, L. GC content around splice sites affects splicing through pre-mRNA secondary structures. BMC Genomics 12, 90 (2011)
Patwardhan, R. P. et al. Massively parallel functional dissection of mammalian enhancers in vivo. Nature Biotechnol. 30, 265–270 (2012)
Mort, M. et al. MutPred Splice: machine learning-based prediction of exonic variants that disrupt splicing. Genome Biol. 15, R19 (2014)
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014)
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nature Biotechnol. 31, 827–832 (2013)
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nature Protocols 8, 2281–2308 (2013)
Carette, J. E. et al. Haploid genetic screens in human cells identify host factors used by pathogens. Science 326, 1231–1235 (2009)
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nature Genet. 46, 310–315 (2014)
Khalid, M. F., Damha, M. J., Shuman, S. & Schwer, B. Structure-function analysis of yeast RNA debranching enzyme (Dbr1), a manganese-dependent phosphodiesterase. Nucleic Acids Res. 33, 6349–6360 (2005)
Elliott, B., Richardson, C., Winderbaum, J., Nickoloff, J. A. & Jasin, M. Gene conversion tracts from double-strand break repair in mammalian cells. Mol. Cell. Biol. 18, 93–101 (1998)
Doyon, Y. et al. Transient cold shock enhances zinc-finger nuclease-mediated gene disruption. Nature Methods 7, 459–460 (2010)
Chen, F. et al. High-frequency genome editing using ssDNA oligonucleotides with zinc-finger nucleases. Nature Methods 8, 753–755 (2011)
Reyon, D. et al. FLASH assembly of TALENs for high-throughput genome editing. Nature Biotechnol. 30, 460–465 (2012)
Carroll, D. Genome engineering with targetable nucleases. Annu. Rev. Biochem. 83, 409–439 (2014)
Smurnyy, Y. et al. DNA sequencing and CRISPR-Cas9 gene editing for target validation in mammalian cells. Nature Chem. Biol. 10, 623–625 (2014)
Kinney, J. B., Murugan, A., Callan, C. G., Jr & Cox, E. C. Using deep sequencing to characterize the biophysical mechanism of a transcriptional regulatory sequence. Proc. Natl Acad. Sci. USA 107, 9158–9163 (2010)
Goina, E., Skoko, N. & Pagani, F. Binding of DAZAP1 and hnRNPA1/A2 to an exonic splicing silencer in a natural BRCA1 exon 18 mutant. Mol. Cell. Biol. 28, 3850–3860 (2008)
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Henikoff, S. & Henikoff, J. G. Amino acid substitution matrices from protein blocks. Proc. Natl Acad. Sci. USA 89, 10915–10919 (1992)
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nature Methods 7, 248–249 (2010)
Acknowledgements
We thank F. Zhang and his laboratory for the CRISPR/Cas9 backbone constructs used in this study and G. Church and his laboratory for providing reagents used to establish CRISPR/Cas9 editing techniques in our lab. We also thank members of the Shendure laboratory for helpful discussions and D. Prunkard for assistance with FACS. This work was supported by the National Institutes of Health (DP1HG007811 to J.S.) and the UW Medical Scientist Training Program (G.M.F. and J.K.).
Author information
Authors and Affiliations
Contributions
The project was conceived and designed by G.M.F. and J.S. G.M.F. and E.A.B. performed experiments. E.A.B. and R.J.H. performed data analysis and generated data figures. G.M.F. generated schematic figures. G.M.F., E.A.B., R.J.H. and J.S. wrote the manuscript. J.C.K. assisted G.M.F to establish genome editing techniques in the laboratory.
Corresponding authors
Ethics declarations
Competing interests
We have filed a provisional patent application on the method.
Extended data figures and tables
Extended Data Figure 1 Distributions and pair-wise correlations of hexamer abundances.
a, The relative abundance of hexamers within the HDR library (red), gDNA (blue), cDNA data (green) are shown for a single experiment. The vertical black line represents our threshold of 10 gDNA reads. b–d, Scatterplots from a single replicate show pair-wise correlations between sequencing counts for the HDR library, gDNA, and cDNA for hexamers with at least 10 observations in the gDNA library, excluding wild type and control hexamers (n = 3,633). The HDR library and the gDNA data are most highly correlated (R 95% confidence interval (CI): 0.596–0.636), followed by the gDNA and cDNA (R 95% CI: 0.419–0.471) and the HDR library and cDNA (R 95% CI: 0.341–0.394).
Extended Data Figure 2 Correlations for hexamer genome editing efficiency and enrichment scores between replicates.
a, gDNA counts for all hexamers with at least ten reads in each of two gDNA preps from separate transfections with the same HDR library (n = 2,980) exhibited moderate correlation (R 95% CI: 0.355–0.416). b, However, hexamer editing rates, defined as gDNA counts normalized to HDR library counts, were substantially less correlated (R 95% CI: 0.084–0.155), consistent with a hexamer’s HDR library abundance contributing more to its gDNA abundance than systematic differences in HDR efficiency secondary to the hexamer sequence itself. c, Hexamer enrichment scores for two pools of cells from a single transfection split on D3 were well-correlated (R 95% CI: 0.643–0.681). d, Pooling data from cells split on D3 replicates from a single transfection yielded an improved correlation between biological replicates (that is, independent transfections; R 95% CI: 0.690–0.722).
Extended Data Figure 3 Comparison of genome-based hexamer enrichment scores to plasmid-based hexamer scores.
a, There was a modest correlation between ESS and ESE hexamers defined by a previous study14 (x-axis) and the enrichment scores calculated here (y-axis; Spearman ρ = 0.524). The previous study also interrogated hexamers positioned +5 to +10 nucleotides relative to a splice junction, but was plasmid-based rather than genome-based and in the context of different exons. b, To reveal effects of GC content on hexamer abundance, histograms display the distribution of enrichment scores for each possible G+C level (0–6). Hexamers containing two or fewer G+C base pairs exhibited broadly lower enrichment scores than hexamers containing three or more G+C base pairs.
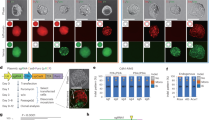
Extended Data Figure 4 Experimental schematic for genome editing and functional analysis of BRCA1 exon 18.
Cultured cells were co-transfected with a single Cas9-sgRNA construct (CRISPR) and an HDR library. Each HDR library was generated from cloning of an oligonucleotide synthesized with 3% nucleotide degeneracy (97WT:1:1:1) for approximately half of the exon and a selective PCR site introduced to the other (fixed) half of the exon (red). CRISPR-induced HDR integrates mutant exons into the genome. Cells were cultured for five days post-transfection, and then harvested for gDNA and total RNA. After reverse transcription, selective PCR was performed before sequencing the edited pools of gDNA and cDNA. Each exon haplotype’s enrichment score was measured by dividing cDNA reads by gDNA reads, and effect sizes for each SNV were calculated via weighted linear regression.
Extended Data Figure 5 Positional SNV editing rates and replication of effect sizes.
a, Editing rates for each SNV in BRCA1 exon 18 were calculated by dividing each SNV’s gDNA sequencing abundance by its HDR library abundance. Editing rates were then plotted across the exon for each library (red = L, blue = R, green = R2) with locations of their selective PCR sites and the CRISPR-targeted PAM illustrated below. For HDR libraries R and R2, there was a subtle decrease in editing rate with increasing distance from the Cas9 cleavage site (rhoR = −0.264, pR = 4.1 × 10−3; rhoR2 = −0.361, pR2 = 4.8 × 10−5). For library L, which allowed re-cutting by not destroying the PAM, there was a sharp peak of editing centred on the Cas9 cleavage site, and a rapid decline in efficiencies in the 5′ direction (further from the 3′ selective PCR handle). b–c, SNV effect sizes were concordant across biological replicates for libraries R2 (b) and L (c) (library R shown in Fig. 2). Notably, variants of high effect size scored similarly across independent transfections.
Extended Data Figure 6 Biological replicate effect size reproducibility for all libraries.
Three separate HDR libraries (R, R2, and L) containing 3% nucleotide degeneracy in either half of BRCA1 exon 18 were introduced to the genome via co-transfection with pCas9-sgBRCA1x18. Enrichment scores were calculated for each haplotype observed at least ten times in the gDNA, and effect sizes of SNVs were determined by weighted linear regression. Effect sizes of individual variants for libraries R2 (left), R (middle), and L (right) were well correlated between biological replicates. Dashed lines represent SNVs that introduce nonsense codons.
Extended Data Figure 7 Correlation between effect sizes and predicted disruption of splicing motifs and indel effects.
a, MutPred Splice17 was used to predict the functional impact of all 234 single nucleotide substitutions on splicing in BRCA1 exon 18 (x-axis), and these scores were compared to absolute values of our empirically measured effect sizes (y-axis; ρ = 0.322). Although nonsense variants contributed to this trend, the sense variants with the largest effect sizes generally had high MutPred Splice scores. b, For indels observed in gDNA from library 2 (virtually all of which occur at the Cas9 cleavage site), size frequencies are plotted. Indel size = 0 includes all haplotypes with wild type length. c, For each indel size, enrichment scores were calculated and normalized to that of the average full length exon. As predicted by nonsense-mediated decay, indels that shift the coding frame were associated with low transcript abundance.
Extended Data Figure 8 Experimental schematic for saturation genome editing and multiplex functional analysis of DBR1 exon 2.
Hap1 cells were co-transfected with a single Cas9-2A-EGFP-sgRNA construct (CRISPR) and an HDR library cloned from array-synthesized oligonucleotides containing programmed SNVs (orange, blue) and active site codon substitutions (green). The HDR library exon haplotypes also included two synonymous mutations (red) to disrupt PAM and protospacer sequences to prevent Cas9 re-cutting, and a 6 bp selective PCR site (light blue) substituted in the downstream intron. Successfully transfected cells (EGFP+) were selected on D2 by FACS, and cultured. On D5, D8, and D11, samples of cells were taken and selective PCR was performed before targeted sequencing of gDNA. Each haplotype’s enrichment score, a measure of the haplotype’s fitness in cell culture, was calculated by dividing D8 or D11 abundance by D5 abundance.
Extended Data Figure 9 DBR1 editing rates by position and comparison of haplotype abundances between D5 and the HDR library, D8, and D11.
a, Editing rates for programmed SNVs represented in the DBR1 gDNA library above threshold (n = 216) were calculated by normalizing each SNV’s gDNA abundance by its HDR library abundance. Rates are plotted by position, with the locations of the targeted PAM (orange) and selective PCR site (purple) indicated below. The editing rate did not significantly change with position (P > 0.05), consistent with positional effects being negated by eliminating re-cutting and performing selective PCR from a distal site. b, Scatterplots display the frequencies at which each haplotype was observed in the D5 sample vs the HDR library, D8, and D11 samples. To account for bottlenecking from editing of a limited number of cells in this representative experiment, analysis of individual haplotypes was restricted to those present at frequencies above 5 × 10−5 in the D5 sample (n = 377; represented by the vertical line). Selection was evident by the depletion of many haplotypes in D8 and D11 samples.
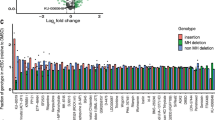
Extended Data Figure 10 Performance of computational predictions of deleterious DBR1 mutations and reproducibility between biological replicates.
a, D11 enrichment scores from a single experiment were used to empirically define deleterious mutations as those with scores fourfold below wild type (vertical line). b, Three in silico metrics of functional impairment were tested for their ability to anticipate the deleteriousness of these mutations as indicated by the area under the receiver operating characteristic curve (AUC): BLOSUM6234 (AUC = 0.672, 214 SNVs), PolyPhen-235 (AUC = 0.671, 155 non-synonymous SNVs), and CADD22 (AUC = 0.701, 214 SNVs). Despite the different approaches of these algorithms, all three exhibited comparably moderate predictive power. c, A biological replicate of the DBR1 experiment was performed and D11 enrichment scores for amino acid substitutions were well correlated (grey lines on scatterplot indicate the ‘deleteriousness’ threshold of fourfold depletion). The distribution of amino acid level enrichment scores for each experiment is displayed along each axis, reflecting bimodality. Notably, unexpected effects (that is, nonsense mutations scoring as tolerated) were among the relatively small percentage of effects not consistent between replicates.
Supplementary information
Supplementary Information
This file contains a Supplementary discussion of the potential sources of noise in the experiments (Supplementary Note 1) and a discussion of potential future applications of the methods presented in the paper (Supplementary Note 2). (PDF 147 kb)
Supplementary Table 1
This table contains a list of oligonucleotide sequences used in this study. (XLSX 14 kb)
Supplementary Table 2
This table contains enrichment scores from the BRCA1 exon 18 hexamer experiment. (XLSX 289 kb)
Supplementary Table 3
This table contains effect sizes from the BRCA1 whole exon 18 SNV experiment. (XLSX 82 kb)
Supplementary Table 4
This table contains enrichment scores from the DBR1 experiment. (XLSX 153 kb)
Rights and permissions
About this article
Cite this article
Findlay, G., Boyle, E., Hause, R. et al. Saturation editing of genomic regions by multiplex homology-directed repair. Nature 513, 120–123 (2014). https://doi.org/10.1038/nature13695
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13695
This article is cited by
-
Building in vitro tools for livestock genomics: chromosomal variation within the PK15 cell line
BMC Genomics (2024)
-
A founder DBR1 variant causes a lethal form of congenital ichthyosis
Human Genetics (2023)
-
Intron turnover is essential to the development and pathogenicity of the plant pathogenic fungus Fusarium graminearum
Communications Biology (2022)
-
Saturation variant interpretation using CRISPR prime editing
Nature Biotechnology (2022)
-
Purification of multiplex oligonucleotide libraries by synthesis and selection
Nature Biotechnology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.