Abstract
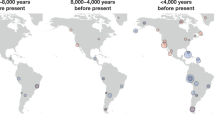
We sequenced the genomes of a ∼7,000-year-old farmer from Germany and eight ∼8,000-year-old hunter-gatherers from Luxembourg and Sweden. We analysed these and other ancient genomes1,2,3,4 with 2,345 contemporary humans to show that most present-day Europeans derive from at least three highly differentiated populations: west European hunter-gatherers, who contributed ancestry to all Europeans but not to Near Easterners; ancient north Eurasians related to Upper Palaeolithic Siberians3, who contributed to both Europeans and Near Easterners; and early European farmers, who were mainly of Near Eastern origin but also harboured west European hunter-gatherer related ancestry. We model these populations’ deep relationships and show that early European farmers had ∼44% ancestry from a ‘basal Eurasian’ population that split before the diversification of other non-African lineages.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
European Nucleotide Archive
Data deposits
The aligned sequences are available through the European Nucleotide Archive under accession number PRJEB6272. The fully public version of the Human Origins dataset can be found at (http://genetics.med.harvard.edu/reichlab/Reich_Lab/Datasets.html). The full version of the dataset (including additional samples) is available to researchers who send a signed letter to D.R. indicating that they will abide by specified usage conditions (Supplementary Information section 9).
References
Keller, A. et al. New insights into the Tyrolean Iceman’s origin and phenotype as inferred by whole-genome sequencing. Nature Commun. 3, 698 (2012)
Olalde, I. et al. Derived immune and ancestral pigmentation alleles in a 7,000-year-old Mesolithic European. Nature 507, 225–228 (2014)
Raghavan, M. et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505, 87–91 (2014)
Skoglund, P. et al. Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science 336, 466–469 (2012)
Bramanti, B. et al. Genetic discontinuity between local hunter-gatherers and Central Europe’s first farmers. Science 326, 137–140 (2009)
Haak, W. et al. Ancient DNA from European early Neolithic farmers reveals their Near Eastern affinities. PLoS Biol. 8, e1000536 (2010)
Lipson, M. et al. Efficient moment-based inference of admixture parameters and sources of gene flow. Mol. Biol. Evol. 30, 1788–1802 (2013)
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012)
Krause, J. et al. A complete mtDNA genome of an early modern human from Kostenki, Russia. Curr. Biol. 20, 231–236 (2010)
Sawyer, S., Krause, J., Guschanski, K., Savolainen, V. & Pääbo, S. Temporal patterns of nucleotide misincorporations and DNA fragmentation in ancient DNA. PLoS ONE 7, e34131 (2012)
Haak, W. et al. Ancient DNA from the first European farmers in 7500-year-old Neolithic sites. Science 310, 1016–1018 (2005)
Perry, G. H. et al. Diet and the evolution of human amylase gene copy number variation. Nature Genet. 39, 1256–1260 (2007)
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009)
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006)
Reich, D., Thangaraj, K., Patterson, N., Price, A. L. & Singh, L. Reconstructing Indian population history. Nature 461, 489–494 (2009)
Moorjani, P. et al. Genetic evidence for recent population mixture in India. Am. J. Hum. Genet. 93, 422–438 (2013)
Reich, D. et al. Reconstructing Native American population history. Nature 488, 370–374 (2012)
Botigué, L. R. et al. Gene flow from North Africa contributes to differential human genetic diversity in southern Europe. Proc. Natl Acad. Sci. USA. 110, 11791–11796 (2013)
Cerezo, M. et al. Reconstructing ancient mitochondrial DNA links between Africa and Europe. Genome Res. 22, 821–826 (2012)
Moorjani, P. et al. The history of African gene flow into southern Europeans, Levantines, and Jews. PLoS Genet. 7, e1001373 (2011)
Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, e1002967 (2012)
Fu, Q. et al. DNA analysis of an early modern human from Tianyuan Cave, China. Proc. Natl Acad. Sci. USA 110, 2223–2227 (2013)
Bar-Yosef, O. The Chronology of the Middle Paleolithic of the Levant 39–56 (Plenum Press, 1998)
Armitage, S. J. et al. The southern route “out of Africa”: evidence for an early expansion of modern humans into Arabia. Science 331, 453–456 (2011)
Rose, J. I. et al. The Nubian Complex of Dhofar, Oman: an African middle stone age industry in Southern Arabia. PLoS ONE 6, e28239 (2011)
Brace, C. L. et al. The questionable contribution of the Neolithic and the Bronze Age to European craniofacial form. Proc. Natl Acad. Sci. USA 103, 242–247 (2006)
Browning, B. L. & Browning, S. R. Improving the accuracy and efficiency of identity-by-descent detection in population data. Genetics 194, 459–471 (2013)
Ralph, P. & Coop, G. The geography of recent genetic ancestry across Europe. PLoS Biol. 11, e1001555 (2013)
Lawson, D. J., Hellenthal, G., Myers, S. & Falush, D. Inference of population structure using dense haplotype data. PLoS Genet. 8, e1002453 (2012)
Brandt, G. et al. Ancient DNA reveals key stages in the formation of central European mitochondrial genetic diversity. Science 342, 257–261 (2013)
Delsate, D., Guinet, J.-M. & Saverwyns, S. De l’ocre sur le crâne mésolithique (haplogroupe U5a) de Reuland-Loschbour (Grand-Duché de Luxembourg)? Bull. Soc. Préhist. Luxembourgeoise 31, 7–30 (2009)
Rohland, N. & Hofreiter, M. Ancient DNA extraction from bones and teeth. Nature Protocols 2, 1756–1762 (2007)
Dabney, J. et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc. Natl Acad. Sci. USA 110, 15758–15763 (2013)
Stäuble, H. Häuser und absolute Datierung der Ältesten Bandkeramik (Habelt, 2005)
Yang, D. Y., Eng, B., Waye, J. S., Dudar, J. C. & Saunders, S. R. Improved DNA extraction from ancient bones using silica-based spin columns. Am. J. Phys. Anthropol. 105, 539–543 (1998)
Meyer, M. & Kircher, M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb. Protoc.. http://dx.doi.org/10.1101/prot5448 (2010)
Meyer, M. et al. A high-coverage genome sequence from an Archaic Denisovan individual. Science 338, 222–226 (2012)
Briggs, A. W. et al. Removal of deaminated cytosines and detection of in vivo methylation in ancient DNA. Nucleic Acids Res. 38, e87 (2010)
Kircher, M. Analysis of high-throughput ancient DNA sequencing data. Methods Mol. Biol. 480, 197–228 (2012)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
Maricic, T., Whitten, M. & Pääbo, S. Multiplexed DNA sequence capture of mitochondrial genomes using PCR products. PLoS ONE 5, e14004 (2010)
Behar, D. M. et al. A Copernican reassessment of the human mitochondrial DNA tree from its root. Am. J. Hum. Genet. 90, 675–684 (2012)
Green, R. E. et al. A complete Neandertal mitochondrial genome sequence determined by high-throughput sequencing. Cell 134, 416–426 (2008)
Fu, Q. et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr. Biol. 23, 553–559 (2013)
Fu, Q. et al. The complete genome sequence of a 45,000-year-old modern human from western Siberia. Nature (in the press)
Rasmussen, M. et al. An Aboriginal Australian genome reveals separate human dispersals into Asia. Science 334, 94–98 (2011)
Vianello, D. et al. HAPLOFIND: a new method for high-throughput mtDNA haplogroup assignment. Hum. Mutat. 34, 1189–1194 (2013)
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011)
Skoglund, P., Storå, J., Götherström, A. & Jakobsson, M. Accurate sex identification of ancient human remains using DNA shotgun sequencing. J. Archaeol. Sci. 40, 4477–4482 (2013)
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 7, 214 (2007)
Lippold, S. et al. Human paternal and maternal demographic histories: insights from high-resolution Y chromosome and mtDNA sequences. Preprint at bioRxiv, http://dx.doi.org/10.1101/001792 (2014)
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010)
Reich, D. et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468, 1053–1060 (2010)
Prüfer, K. et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505, 43–49 (2014)
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011)
Hach, F. et al. mrsFAST: a cache-oblivious algorithm for short-read mapping. Nature Methods 7, 576–577 (2010)
The 1000 Genomes Project Consortium An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012)
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011)
Li, H. The sequence alignment/map (SAM) format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Keinan, A., Mullikin, J. C., Patterson, N. & Reich, D. Measurement of the human allele frequency spectrum demonstrates greater genetic drift in East Asians than in Europeans. Nature Genet. 39, 1251–1255 (2007)
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nature Genet. 38, 904–909 (2006)
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007)
Alexander, D. H. & Lange, K. Enhancements to the ADMIXTURE algorithm for individual ancestry estimation. BMC Bioinformatics 12, 246 (2011)
Jakobsson, M. & Rosenberg, N. A. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806 (2007)
Price, A. L., Zaitlen, N. A., Reich, D. & Patterson, N. New approaches to population stratification in genome-wide association studies. Nature Rev. Genet. 11, 459–463 (2010)
Busing, F. T. A., Meijer, E. & Leeden, R. Delete-m Jackknife for Unequal m. Stat. Comput. 9, 3–8 (1999)
Loh, P.-R. et al. Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193, 1233–1254 (2013)
Nelson, M. R. et al. The population reference sample, POPRES: a resource for population, disease, and pharmacological genetics research. Am. J. Hum. Genet. 83, 347–358 (2008)
Acknowledgements
We thank the 1,615 volunteers from 147 diverse populations who donated DNA samples and whose genetic data are newly reported in this study. We are grateful to C. Beall, N. Bradman, A. Gebremedhin, D. Labuda, M. Nelis and A. Di Rienzo for sharing DNA samples; to D. Weigel, C. Lanz, V. Schünemann, P. Bauer and O. Riess for support and access to DNA sequencing facilities; to P. Johnson for advice on contamination estimation; to G. Hellenthal for help with the ChromoPainter software; and to P. Skoglund for sharing graphics software. We thank K. Nordtvedt for alerting us to newly discovered Y-chromosome SNPs. We downloaded the POPRES data from dbGaP at (http://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000145.v4.p2) through dbGaP accession number phs000145.v1.p2. We thank all the volunteers who donated DNA. We thank the staff of the Unità Operativa Complessa di Medicina Trasfusionale, Azienda Ospedaliera Umberto I, Siracusa, Italy for assistance in sample collection; and The National Laboratory for the Genetics of Israeli Populations for facilitating access to DNA. We thank colleagues at the Applied Genomics at the Children’s Hospital of Philadelphia, especially H. Hakonarson, C. Kim, K. Thomas, and C. Hou, for genotyping samples on the Human Origins array. J.Kr., A.M. and C.P. are grateful for support from DFG grant number KR 4015/1-1, the Carl-Zeiss Foundation and the Baden Württemberg Foundation. S.P., G.R., Q.F., C.F., K.P., S.C. and J.Ke. acknowledge support from the Presidential Innovation Fund of the Max Planck Society. G.R. was supported by an NSERC fellowship. J.G.S. acknowledges use of the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by NSF grant number OCI-1053575. E.B. and O.B. were supported by RFBR grants 13-06-00670, 13-04-01711, 13-04-90420 and by the Molecular and Cell Biology Program of the Presidium, Russian Academy of Sciences. B.M. was supported by grants OTKA 73430 and 103983. A.Saj. was supported by a Finnish Professorpool (Paulo Foundation) Grant. The Lithuanian sampling was supported by the LITGEN project (VP1-3.1-ŠMM-07-K-01-013), funded by the European Social Fund under the Global Grant Measure. A.S. was supported by Spanish grants SAF2011-26983 and EM 2012/045. O.U. was supported by Ukrainian SFFS grant F53.4/071. S.A.T. was supported by NIH Pioneer Award 8DP1ES022577-04 and NSF HOMINID award BCS-0827436. K.T. was supported by an Indian CSIR Network Project (GENESIS: BSC0121). L.S. was supported by an Indian CSIR Bhatnagar Fellowship. R.V., M.M., J.P. and E.M. were supported by the European Union Regional Development Fund through the Centre of Excellence in Genomics to the Estonian Biocentre and University of Tartu and by an Estonian Basic Research grant SF0270177As08. M.M. was additionally supported by Estonian Science Foundation grant number 8973. J.G.S. and M.S. were supported by NIH grant GM40282. P.H.S. and E.E.E. were supported by NIH grants HG004120 and HG002385. D.R. and N.P. were supported by NSF HOMINID award BCS-1032255 and NIH grant GM100233. D.R. and E.E.E. are Howard Hughes Medical Institute investigators. This project has been funded in part with federal funds from the National Cancer Institute, National Institutes of Health, under contract HHSN26120080001E. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government. This Research was supported in part by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research.
Author information
Authors and Affiliations
Contributions
B.B., E.E.E., J.Bu., M.S., S.P., J.Ke., D.R. and J.Kr. supervised the study. I.L., N.P., A.M., G.R., S.M., K.K., P.H.S., J.G.S., S.C., M.L., Q.F., H.L., C.dF., K.P., W.H., M.Met., M.Mey. and D.R. analysed genetic data. F.H., E.F., D.D., M.F., J.-M.G., J.W., A.C. and J.Kr. obtained human remains. A.M., C.E., R.Bo., K.I.B., S.S., C.P., N.R. and J.Kr. processed ancient DNA. I.L., N.P., S.N., N.R., G.A., H.A.B., G.Ba., E.B., O.B., R.Ba., G.Be., H.B.-A., J.Be., F.Be., C.M.B., F.Br., G.B.J.B., F.C., M.C., D.E.C.C., D.Cor., L.D., G.vD., S.D., J.-M.D., S.A.F., I.G.R., M.G., M.H., B.M.H., T.H., U.H., A.R.J., S.K.-Y., R.Kh., E.K., R.Ki., T.K., W.K., V.K., A.K., L.L., S.L., T.L., R.W.M., B.M., E.M., J.Mol., J.Mou., K.N., D.N., T.N., L.O., J.P., F.P., O. P., V.R., F.R., I.R., R.R., H.S., A.Saj., A.Sal., E.B.S., A.Tar., D.T., S.T., I.U., O.U., R.Va., M.Vi., M.Vo., C.A.W., L.Y., P.Z., T.Z., C.C., M.G.T., A.R.-L., S.A.T., L.S., K.T., R.Vi., D.Com., R.S., M.Met., S.P. and D.R. assembled the genotyping dataset. I.L., N.P., D.R. and J.Kr. wrote the manuscript with help from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
U.H. is an employee of Illumina, T.L. is an employee of Amgen, and J.M. is an employee of 23andMe.
Extended data figures and tables
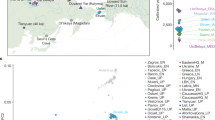
Extended Data Figure 1 Photographs of analysed ancient samples.
a, Loschbour skull. b, Stuttgart skull, missing the lower right M2 we sampled. c, Excavation at Kanaljorden in Motala, Sweden. d, Motala 1 in situ.
Extended Data Figure 2 Pairwise sequential Markovian coalescent (PSMC) analysis.
a, Inference of population size as a function of time, showing a very small recent population size over the most recent period in the ancestry of Loschbour (at least the last 5–10 thousand years). b, Inferred time since the most recent common ancestor from the PSMC for chromosomes 20, 21, 22 (top to bottom); Stuttgart is plotted on top and Loschbour at bottom.
Extended Data Figure 3 ADMIXTURE analysis (K = 2 to K = 20).
Ancient samples (Loschbour, Stuttgart, Motala_merge, Motala12, MA1, and LaBraña) are on the left.
Extended Data Figure 4 ANE ancestry is present in both Europe and the Near East but WHG ancestry is restricted to Europe, which cannot be due to a single admixture event.
On the x axis we present the statistic f4(Test, Stuttgart; MA1, Chimp), which measures where MA1 shares more alleles with a test population than with Stuttgart. It is positive for most European and Near Eastern populations, consistent with ANE (MA1-related) gene flow into both regions. On the y axis we present the statistic f4(Test, Stuttgart; Loschbour, Chimp), which measures whether Loschbour shares more alleles with a test sample than with Stuttgart. Only European populations show positive values of this statistic, providing evidence of WHG (Loschbour-related) admixture only in Europeans.
Extended Data Figure 5 MA1 is the best surrogate for ANE for which we have data.
Europeans share more alleles with MA1 than with Karitiana, as we see from the fact that in a plot of f4(Test, BedouinB; MA1, Chimp) and f4(Test, BedouinB; Karitiana, Chimp), the European cline deviates in the direction of MA1, rather than Karitiana (the slope is > 1 and European populations are above the line indicating inequality of these two statistics).
Extended Data Figure 6 The differential relatedness of west Eurasians to Stuttgart (EEF), Loschbour (WHG), and MA1 (ANE) cannot be explained by two-way mixture.
We plot on a West Eurasian map the statistic f4(Test, Chimp; A1, A2), where A1 and A2 are a pair of the three ancient samples representing the three ancestral populations of Europe. a, In both Europe and the Near East/Caucasus, populations from the south have more relatedness to Stuttgart than those from the north where ANE influence is also important. b, Northern European populations share more alleles with Loschbour than with Stuttgart, as they have additional WHG ancestry beyond what was already present in EEF. c, We observe a striking contrast between Europe west of the Caucasus and the Near East in degree of relatedness to WHG. In Europe, there is a much higher degree of allele sharing with Loschbour than with MA1, which we ascribe to the 60–80% WHG/(WHG + ANE) ratio in most Europeans that we report in Supplementary Information section 14. In contrast, the Near East has no appreciable WHG ancestry but some ANE ancestry, especially in the northern Caucasus. (Jewish populations are marked with a square in this figure to assist in interpretation as their ancestry is often anomalous for their geographic regions.)
Extended Data Figure 7 Evidence for Siberian gene flow into far north-eastern Europe.
Some north-eastern European populations (Chuvash, Finnish, Russian, Mordovian, Saami) share more alleles with Han Chinese than with other Europeans who are arrayed in a cline from Stuttgart to Lithuanians/Estonians in a plot of f4(Test, BedouinB; Han, Mbuti) against f4(Test, BedouinB; MA1, Mbuti).
Supplementary information
Supplementary Information
This file contains Supplementary Information Parts 1-19 – see Supplementary Contents for details.This file contains Supplementary Information Parts 1-19 – see Supplementary Contents for details. (PDF 9752 kb)
Rights and permissions
About this article
Cite this article
Lazaridis, I., Patterson, N., Mittnik, A. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014). https://doi.org/10.1038/nature13673
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13673
This article is cited by
-
Reconstructing the ancestral gene pool to uncover the origins and genetic links of Hmong–Mien speakers
BMC Biology (2024)
-
Differentiated genomic footprints suggest isolation and long-distance migration of Hmong-Mien populations
BMC Biology (2024)
-
The Allen Ancient DNA Resource (AADR) a curated compendium of ancient human genomes
Scientific Data (2024)
-
Identification of microbial pathogens in Neolithic Scandinavian humans
Scientific Reports (2024)
-
The genetic legacy of the expansion of Bantu-speaking peoples in Africa
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.