Abstract
Pancreatic ductal adenocarcinoma (PDAC) is one of the deadliest cancers in western countries, with a median survival of 6 months and an extremely low percentage of long-term surviving patients. KRAS mutations are known to be a driver event of PDAC1, but targeting mutant KRAS has proved challenging2. Targeting oncogene-driven signalling pathways is a clinically validated approach for several devastating diseases3,4. Still, despite marked tumour shrinkage, the frequency of relapse indicates that a fraction of tumour cells survives shut down of oncogenic signalling5,6. Here we explore the role of mutant KRAS in PDAC maintenance using a recently developed inducible mouse model of mutated Kras1 (KrasG12D, herein KRas) in a p53LoxP/WT background. We demonstrate that a subpopulation of dormant tumour cells surviving oncogene ablation (surviving cells) and responsible for tumour relapse has features of cancer stem cells and relies on oxidative phosphorylation for survival. Transcriptomic and metabolic analyses of surviving cells reveal prominent expression of genes governing mitochondrial function, autophagy and lysosome activity, as well as a strong reliance on mitochondrial respiration and a decreased dependence on glycolysis for cellular energetics. Accordingly, surviving cells show high sensitivity to oxidative phosphorylation inhibitors, which can inhibit tumour recurrence. Our integrated analyses illuminate a therapeutic strategy of combined targeting of the KRAS pathway and mitochondrial respiration to manage pancreatic cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ying, H. et al. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 149, 656–670 (2012)
Karnoub, A. E. & Weinberg, R. A. Ras oncogenes: split personalities. Nature Rev. Mol. Cell Biol. 9, 517–531 (2008)
Kantarjian, H. et al. Hematologic and cytogenetic responses to imatinib mesylate in chronic myelogenous leukemia. N. Engl. J. Med. 346, 645–652 (2002)
Flaherty, K. T. et al. Inhibition of mutated, activated BRAF in metastatic melanoma. N. Engl. J. Med. 363, 809–819 (2010)
Quintas-Cardama, A., Kantarjian, H. & Cortes, J. Imatinib and beyond—exploring the full potential of targeted therapy for CML. Nature Rev. Clin. Oncol. 6, 535–543 (2009)
Jang, S. & Atkins, M. B. Which drug, and when, for patients with BRAF-mutant melanoma? Lancet Oncol. 14, e60–e69 (2013)
Bardeesy, N. et al. Both p16Ink4a and the p19Arf-p53 pathway constrain progression of pancreatic adenocarcinoma in the mouse. Proc. Natl Acad. Sci. USA 103, 5947–5952 (2006)
Hermann, P. C. et al. Distinct populations of cancer stem cells determine tumor growth and metastatic activity in human pancreatic cancer. Cell Stem Cell 1, 313–323 (2007)
Li, C. et al. Identification of pancreatic cancer stem cells. Cancer Res. 67, 1030–1037 (2007)
Kim, M. P. et al. ALDH activity selectively defines an enhanced tumor-initiating cell population relative to CD133 expression in human pancreatic adenocarcinoma. PLoS ONE 6, e20636 (2011)
Fernandez-Marcos, P. J. & Auwerx, J. Regulation of PGC-1α, a nodal regulator of mitochondrial biogenesis. Am. J. Clin. Nutr. 93, 884S–890S (2011)
Yuan, M., Breitkopf, S. B., Yang, X. & Asara, J. M. A positive/negative ion-switching, targeted mass spectrometry-based metabolomics platform for bodily fluids, cells, and fresh and fixed tissue. Nature Protocols 7, 872–881 (2012)
Gaglio, D. et al. Oncogenic K-Ras decouples glucose and glutamine metabolism to support cancer cell growth. Mol. Syst. Biol. 7, 523 (2011)
Son, J. et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496, 101–105 (2013)
Vizan, P. et al. K-ras codon-specific mutations produce distinctive metabolic phenotypes in NIH3T3 mice fibroblasts. Cancer Res. 65, 5512–5515 (2005)
Liu, Y. & Schubert, D. R. The specificity of neuroprotection by antioxidants. J. Biomed. Sci. 16, 98 (2009)
Kim, M. P. et al. Generation of orthotopic and heterotopic human pancreatic cancer xenografts in immunodeficient mice. Nature Protocols 4, 1670–1680 (2009)
Shi, Y. et al. Mammalian transcription factor A is a core component of the mitochondrial transcription machinery. Proc. Natl Acad. Sci. USA 109, 16510–16515 (2012)
Škrtić, M. et al. Inhibition of mitochondrial translation as a therapeutic strategy for human acute myeloid leukemia. Cancer Cell 20, 674–688 (2011)
Singh, R. et al. Autophagy regulates lipid metabolism. Nature 458, 1131–1135 (2009)
Guo, J. Y. et al. Activated Ras requires autophagy to maintain oxidative metabolism and tumorigenesis. Genes Dev. 25, 460–470 (2011)
Yang, S. et al. Pancreatic cancers require autophagy for tumor growth. Genes Dev. 25, 717–729 (2011)
Nakada, D., Saunders, T. L. & Morrison, S. J. Lkb1 regulates cell cycle and energy metabolism in haematopoietic stem cells. Nature 468, 653–658 (2010)
Lagadinou, E. D. et al. BCL-2 inhibition targets oxidative phosphorylation and selectively eradicates quiescent human leukemia stem cells. Cell Stem Cell 12, 329–341 (2013)
Samudio, I. et al. Pharmacologic inhibition of fatty acid oxidation sensitizes human leukemia cells to apoptosis induction. J. Clin. Invest. 120, 142–156 (2010)
Fung, C., Lock, R., Gao, S., Salas, E. & Debnath, J. Induction of autophagy during extracellular matrix detachment promotes cell survival. Mol. Biol. Cell 19, 797–806 (2008)
Zheng, W., Talley Watts, L., Holstein, D. M., Wewer, J. & Lechleiter, J. D. P2Y1R-initiated, IP3R-dependent stimulation of astrocyte mitochondrial metabolism reduces and partially reverses ischemic neuronal damage in mouse. J. Cereb. Blood Flow Metab. 33, 600–611 (2013)
Itoh, Y., Abe, T., Takaoka, R. & Tanahashi, N. Fluorometric determination of glucose utilization in neurons in vitro and in vivo. J. Cereb. Blood Flow Metab. 24, 993–1003 (2004)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Acknowledgements
We thank A. Divakaruni, J. Dunn, C. Smith, K. McGirr and D. Ferrick for their support with the Seahorse Bioscience XF96 Analyser; T. Tieu for vector cloning and J. Kovacs for support with the YSI analyser; J. D. Lechleiter for the protocol to measure mitochondrial potential in vivo; H. Sandoval, C. Tacchetti, M. E. Di Francesco, J. Marszalek and P. Jones for discussions and suggestions; K. Dunner Jr and the High Resolution Electron Microscopy Facility at the MD Anderson Cancer Center (MDACC) for TEM (Cancer Center Core Grant CA16672); W. N. Hittelman and the Center for Targeted Therapy for confocal microscopy; the Dana-Farber Cancer Institute Microarray Core Facility for Affymetrix expression profiling and the MDACC Sequencing and Microarray Facility (SMF) funded by National Cancer Institute (NCI) grant CA016672 (SMF) for exome sequencing; the MDACC Flow Cytometry and Cellular Imaging Core Facility supported by grant NCI#P30CA16672 for flow cytometers and FACS; D. Jayanta for providing GFP–LC3 constructs; B. Perrazzona, U. Varadarajan and R. Dewan for lab management; and S. Jiang for assistance in maintenance of mouse colonies. A.V. is thankful to A. Fantino, S. Rapi, V. Giuliani and P. Viale for their continuous support. This study was supported by grants from the Hirshberg Foundation for Pancreatic Cancer Research to A.V., Harvard Stem Cell Institute to R.A.D. and A.V., Sheikh Ahmed Center for Pancreatic Cancer Research to G.F.D., T.P.H. and A.V., American Italian Cancer Foundation to G.F.D., National Institutes of Health (NIH) P01CA117969 to R.A.D., NIH/NCI P01CA120964 to J.M.A., The Viragh Family Foundation to J.B.F.; C.A.L. is a Pancreatic Cancer Action Network-AACR Pathway to Leadership Fellow.
Author information
Authors and Affiliations
Contributions
A.V., P.G., R.A.D. and G.F.D. designed the studies, interpreted the data and wrote the manuscript; A.V., P.G., H.Y., N.S., M.M., A.C., T.G. and V.G. performed the experiments; C.A.L. was responsible for metabolomics and carbon-13 tracing experiments; S.S. was responsible for CNV and bioinformatics analysis; M.K.-A, F.M., S.C., L.N., G.G., A.K.D., A.K., W.Y., E.B., Y.K., T.P.H., A.C.K., H.W. and J.B.F. contributed essential reagents and resources; M.Y. and J.M.A. helped with the metabolomics analysis; F.M., Y.A.W. and L.C.C. assisted with data interpretation; A.K.D. edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
L.C.C. owns equity in, receives compensation from, and serves on the Board of Directors and Scientific Advisory Board of Agios Pharmaceuticals. Agios Pharmaceuticals is identifying metabolic pathways of cancer cells and developing drugs to inhibit such enzymes in order to disrupt tumour cell growth and survival.
Extended data figures and tables
Extended Data Figure 1 Oncogene ablation leads to tumour regression in vitro and in vivo.
a, In vivo experimental scheme. Tumour cells isolated from primary tumours or tumour spheres were injected in nude mice fed with doxycycline in drinking water (+Dox). When mice developed tumours, doxycycline was withdrawn (−Dox) and tumours underwent a complete macroscopic regression after 2–3 weeks (arrows indicate regressed tumours). In residual lesions few tumour cells remain quiescent for months and they can quickly reform tumours upon KRas reactivation (+Dox). b, Tumours expressing KRas (+KRas) and tumour remnants after regression (−KRas) are positive for ductal epithelial marker CK19 (×40). c, Tumours expressing KRas (+KRas) and epithelial remnants after tumour regression (−KRas) were stained for phosphorylated-p42/44 (pErk). No signal is detected in surviving cells (×20). d, In vitro experimental scheme. After digestion to a single-cell suspension, tumour cells isolated from primary tumours were plated in stem cell medium in presence of doxycycline (+Dox, +KRas). Spherogenic cells form tumour spheres (+KRas) that can be maintained by serial replating in presence of doxycycline. Upon doxycycline withdrawal (−Dox) tumour spheres undergo involution and only a minority of cells survive the ablation of KRas (SCs, −KRas). SCs readily reform tumour spheres upon re-activation of KRas (+Dox). e, The amount of active Ras in KRas-expressing cells (+KRas) and SCs (−KRas) was evaluated in three independent tumour spheres by detecting the fraction of Ras protein that co-precipitates with Raf kinase. Total lysates were probed with anti-phospho-p42/44 (pErk), total p42/44 (Erk) antibodies. f, Annexin V staining in tumour spheres after 3 days +/−KRas (n = 3). g, Sphere formation is a regulated process and tumour cells enter and exit cell cycle. BrdU incorporation (pulse of 3 h) was evaluated at different time points during sphere formation and regression. KRas-expressing fully formed spheres (day (D)0 and 8) are quiescent. Upon sphere dissociation and replating (D0), spherogenic cells enter cell cycle (D1) and tumour cells continue to grow until day 3–4, when spheres reach their maximal S phase. Then tumour cells gradually exit the cell cycle and become quiescent (D8). After doxycycline withdrawal (−KRas), tumour spheres undergo involution and surviving cells remain quiescent until KRas is re-expressed (−KRas 24 h +KRas) and spheres are reformed. Ruling out the effect of the cell cycle, transcriptomic and metabolomic characterizations were done, matching quiescent surviving tumour cells to quiescent fully formed KRas-expressing spheres at D8 (n = 3). h, Haematoxylin and eosin staining and immunohistochemistry of regressed tumours after three 8 h pulses of BrdU show that epithelial remnants in regressed tumours after KRas ablation (−KRas) are completely quiescent (left panels). Forty-eight hours after KRas reactivation (doxycycline i.p. injection) tumour cells re-enter massively the cell cycle (right panels). Red arrows indicate mitotic cells (×20). i, Representative annexin V staining with respect to CD133 and CD44 after 3 days of KRas ablation, two independent tumours are represented. Data are mean ± s.d.
Extended Data Figure 2 Transplantation in limiting dilution and characterization of epithelial remnants.
a–d, Transplantation in limiting dilution. Experiments, number of transplanted mice and percentage of developed tumour are shown. a, Limiting dilution experiments using tumour spheres (+KRas) and surviving cells (−KRas) (genetic model ex vitro). b, Limiting dilution experiments using cells isolated from KRas-expressing (+KRas) and regressed tumours (−KRas) (genetic model ex vivo). c, Top panels, immunoblots of tumour spheres treated with different concentrations of Mek1 (AZD8330) and dual PI3K/mTOR (BEZ235) inhibitors probed with anti-phospho-p42/44 (pErk), phospho-Akt (pAkt), pan-Ras (Ras) and β-actin (Act) antibodies. Bottom panels, effects of AZD8330 (AZD 0.01 μM) and BEZ235 (BEZ 0.1 μM) treatment for 1 week on tumour sphere formation; some cells, as single cells or in small clusters, are able to survive the treatment (×5). d, Limiting dilution experiments using cells surviving pharmacological downregulation of oncogenic pathways (AZD+BEZ, combination of AZD8330 and BEZ235) and control cells (CTRL). e, The plot shows the cumulative distribution of coverage at all the SNVs called by Unified Genotyper (across samples). f, CD44 is expressed during tumorigenesis in mice: no positive cells are detected in normal pancreas (left panel), KRas-expressing tumours express high levels of CD44 (middle panel), epithelial remnants in regressed tumours maintain their positivity for CD44 (right panel) (×10). g, Validation of CD133 (ab16518) in immunohistochemistry: this antibody does not recognize cells and ductal structures in normal pancreas (left panel), a small population of cells is stained by ab16518 in KRas-expressing tumours (middle panel), epithelial remnants in regressed tumours are strongly positive for CD133 (right panel) (×10). At higher magnification (red boxes), it is possible to appreciate the classical polarized pattern of CD133.
Extended Data Figure 3 qPCR validation of pathways enriched in surviving cells.
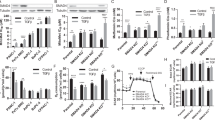
a–e, mRNA fold change in surviving cells normalized to KRas-expressing cells. a, Genes involved in ETC (n = 5). b, Genes involved in the biogenesis and function of mitochondria (Mitochondria) (n = 5). c, d, Genes encoding proteins of the autophagic molecular machinery and its key regulators (n = 5) (c) and β-oxidation (n = 5) (d). e, mRNA fold change in cells surviving AZD8330 plus BEZ235 treatment (AZD+BEZ) versus controls (n = 3). Data are mean ± s.d.
Extended Data Figure 4 Surviving cells have more active mitochondria.
a, mRNA fold change of Ppargc1 genes in −KRas versus +KRas cells (n = 5). b, Quantification of MitoTracker Green staining in +KRas and −KRas cells (n = 3). c, Mitochondrial membrane potential (Δψm) of +KRas and −KRas cells (n = 4); representative flow-cytometry analysis of two tumours. d, Immunoblot of two independent tumour spheres derived from different genetic backgrounds (Ink4a/Arf−/− and p53−/−) treated or not with AZD8330 and BEZ235 (AZD+BEZ) for 7 days and probed with anti-VDAC1 (VDAC1) and β-actin (Actin) antibodies. e, Cells surviving AZD+BEZ treatment have higher mitochondrial transmembrane potential (Δψm) than untreated cells (CTRL) (n = 3); representative flow-cytometry analysis is reported. f, Mice bearing tumours have been treated (AZD+BEZ) or not (CTRL) with a combination of AZD6244 and BEZ235 for 1 week. Upon tail vein injection of a bolus of TMRE, tumours were explanted and analysed by flow cytometry for their mitochondrial potential (Δψm) upon gating on CD44+ DAPI− cells (n = 3). A representatie flow-cytometry analysis of two different tumours is reported, AB+CCCP samples represent reacquisition of AZD+BEZ samples after incubation with CCCP for 5 min. g, ROS production in +KRas and −KRas cells (n = 3); representative flow-cytometry analysis of two tumours. h, Live confocal imaging of SCs stained for mitochondria (MitoTracker Green), ROS (CellRox-Red) and DNA (Hoechst). The vast majority of signal generated by ROS colocalizes with mitochondria. i, Immunoblot of Aldefluor/CD133 double-positive and double-negative cells sorted from two independent tumours probed with anti-VDAC1 (VDAC1) and β-actin (Actin) antibodies. j, KRas-expressing cells positive for aldefluor (Ald+) and CD133 (CD133+) have higher mitochondrial transmembrane potential (Δψm) than tumour cells that do not express the same markers (Ald− and CD133−) (n = 3); a representative flow-cytometry analysis is reported. Data are mean ± s.d.
Extended Data Figure 5 OCR, ECAR and metabolomics.
a, OCR of KRas-expressing cells (+KRas) and SCs (−KRas) in response to oligomycin, FCCP and rotenone/antimycin (n = 4). b, Same as in a but normalized to basal respiration of +KRas and −Kras cells. c, ECAR response of +KRas and −KRas cells to oligomycin and 2DG. The experiment has been carried out in complete stem cell media to evaluate the glycolytic reserve of tumour cells in a nutrient rich environment. d, Metabolome analysis for +/−KRas cells; unsupervised hierarchical clustering and heat map of significantly (P < 0.05) deregulated metabolites (n = 4). e, Lactate production of +KRas and −KRas cells in response to oligomycin (Oligo) or DMSO (Ctrl) treatment (n = 3). f, Fold change of TCA cycle intermediates in +KRas versus −KRas cells (αKG, α-ketoglutarate) (n = 4). g, Fold change of nucleotide triphosphates and deoxynucleotide triphosphates, glutathione (GSH) and glutathione disulphide (GSSG) in −KRas versus +KRas cells (n = 4). Data are mean ± s.d.
Extended Data Figure 6 Surviving cells in vitro and in vivo have an impaired glucose uptake.
a, KRas-expressing cells (+KRas) and SCs (−KRas) were incubated with 2NBDG (2-(N-(7-nitrobenz-2-oxa-1,3-diazol-4-yl)amino)-2-deoxyglucose) for 6 h then analysed by flow cytometry (n = 3); a representative flow-cytometry analysis of spheres derived from two different tumours is reported. b, Mice bearing KRas-expressing tumours (+KRas) and 3-week regressed tumours (−KRas) were injected with a tail vein bolus of 2NBDG. After 1 h tumours were explanted and analysed by flow cytometry upon gating on CD44+ DAPI− cells (n = 3); a representative flow-cytometry analysis of two different tumours is reported. c, d, Tumour spheres derived from different genetic backgrounds (Ink4a/Arf−/− and p53−/−) were treated (+AB) or not with AZD8330 and BEZ235 for 7 days then plated in fresh stem cell medium. After 24 h medium was collected and analysed by YSI analyser for glucose uptake (c) and lactate production (d) (n = 3). Data are mean ± s.d.
Extended Data Figure 7 Fuel carbon contribution to TCA cycle and TCA branch metabolites.
a–h, Isotopomer distribution for lactate (a), alanine (b), glutamate (c), aspartame (d), fumarate (e), citrate (f), isocitrate (g) and malate (h) in KRas-expressing (+) and SCs (−) following steady-state tracing (36 h labelling) with uniformly carbon-13-labelled substrates: glucose, glutamine, palmitate and pyruvate (n = 3). Data are mean ± s.d.
Extended Data Figure 8 Differential sensitivity of tumour cells to OXPHOS inhibition.
a, Annexin V staining of cells treated with oligomycin 200 nM (Oli) for 24 h shows a significant decrease in viability in surviving cells (−KRas). By contrast, control cells expressing KRas (+KRas) are minimally affected (n = 3); a representative flow-cytometry analysis is reported. b, Effect of oligomycin (Oli), dicyclohexylcarbodiimide (DCCD), veturicidin (Vent), rotenone (Rot), antimycin (Anti) and DMSO (Ctrl) on spherogenic potential of KRas-expressing (+KRas) and SCs (−KRas) (n = 4). c, In vivo treatment experimental scheme: mice were transplanted with tumour cells and fed with doxycycline in drinking water (+KRas, +Dox) until they developed tumours of 1 cm in diameter. Then doxycycline was withdrawn (−KRas, −Dox) and after 2 weeks, when tumours were regressed, mice were treated with oligomycin (0.5 mg kg−1, i.p.) or vehicle for 5 days a week, for 2 weeks. After treatment, KRas was re-induced (+Dox) and mice were monitored for tumour relapse. d, One dose of oligomycin (0.5 mg kg−1, i.p.) is sufficient to increase lactate concentration in plasma of treated mice after 4 h from injection. Oligo, oligomycin; Ctrl, vehicle. n = 4. e, Tumour volume of KRas-expressing tumours treated with either vehicle or oligomycin 0.5 mg kg−1, 5 days a week, for 2 weeks. Treatment was started when tumours reached 5 mm of diameter (5 mice per group). f, SCs after treatment with oligomycin show signs of degeneration and epithelial remnants change their morphology. Red arrows indicate the presence of capillaries (red blood cells) indicating that regressed tumours are vascularized (×40). g, Oligomycin (Oli) induces ROS production in KRas-expressing cells (+KRas) and SCs (−KRas). Its effect is even stronger than that of positive control 4-hydroxynonenal (hne) (n = 3). h, Glutathione levels in KRas-expressing cells (+KRas) and SCs (−KRas) before and after buthionine sulphoximine (BSO) treatment. Glutathione is increased in SCs and BSO treatment is effective in reducing its level (n = 3). i, Effect of glutathione depletion on spherogenic potential of KRas-expressing cells (+KRas) and SCs (−KRas) (n = 3). j, ROS production in SCs after treatment with 4-hydroxynonenal (hne) and oligomycin (oli) in the presence or absence of antioxidants: α-tocopherol (vitE), N-acetylcysteine (nac) and tetrakis (Tet) (n = 2). k, Effect of oligomycin on spherogenic potential of surviving cells pre-treated with antioxidants (n = 4). Data are mean ± s.d.
Extended Data Figure 9 Effect of mitochondrial downregulation in human tumour spheres and metabolic stress mediated by inhibition of autophagy.
a, Effects of the combination of AZD8330 and BEZ235 (AZD+BEZ) on human tumour spheres. Some cells, usually doublets, are able to survive the treatment (×5). b, Immunoblots of human tumour spheres treated or not with AZD+BEZ probed with anti-phospho-p42/44 (pErk), total-Erk (Erk), phospho-Akt (pAkt), Akt and β-actin (Actin) antibodies; two independent tumours are reported. c, Annexin V staining of treated (AZD+BEZ) and control (Ctrl) cells after 4 days of treatment (n = 3). d, Mitochondrial transmembrane potential (Δψm) of untreated (Ctrl) and treated (AZD+BEZ) human spheres with AZD8330 and BEZ235 for 7 days (n = 3); representative flow-cytometry analysis of two tumours. e–h, TFAM and TUFM were downregulated using two inducible short hairpin RNAs (shRNAs) each (TFAM: #93, #95; TUFM: #63, #64) in human spheres expressing KRas (untreated) and cells surviving 1 week of treatment with AZD8330 and BEZ235 (AZD+BEZ); after 5 days of shRNA induction cells were replated for evaluating their spherogenic capacity. e, Immunoblots of tumour spheres after 72 h of shRNA induction (+Dox) probed with anti-TFAM, TUFM and HSP90 antibodies. f, Representative calcein staining after sphere replating. g, h, Effects of downregulation of TFAM and TUFM on spherogenic potential of untreated (g) and treated (h) cells; data represent the average of two independent human tumours. i, Immunoblot of KRas-expressing cells treated or not with oligomycin 200 nM (Oligo, +/−) probed with anti-Thr-172-phospho-AMPK and actin antibodies. j, k, Immunoblots of +KRas and −KRas cells treated with etomoxir (Eto, 100 μM for 6 h) (j) and bafilomycin (Baf, 50 nM for 24 h) (k) probed with anti-Thr-172-phospho-AMPK and vinculin antibodies. l, Annexin V staining of cells treated for 48 h with bafilomycin 50 nM (Baf) and etomoxir 100 μM (Eto) clearly shows a significant decrease in viability of SCs (−KRas). Control cells expressing KRas (+KRas) are not affected (n = 3); representative dot-plots are reported. Data are mean ± s.d.
Extended Data Figure 10 Cells surviving oncogene ablation are engorged with autophagosomes and lysosomes and contain lipid droplets.
a, SCs (−KRas) have a cytoplasm full of phagosomes and autophagosomes, a feature absent in KRas-expressing cells (+KRas) (TEM; ×7,500). b, SCs are characterized by the presence of several lipid droplets (arrowheads) in the cytoplasm (TEM; ×7,500). c, Primers used for amplification of mitochondrial and lipid metabolic genes.
Rights and permissions
About this article
Cite this article
Viale, A., Pettazzoni, P., Lyssiotis, C. et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 514, 628–632 (2014). https://doi.org/10.1038/nature13611
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13611
This article is cited by
-
Pancreatic cancer acquires resistance to MAPK pathway inhibition by clonal expansion and adaptive DNA hypermethylation
Clinical Epigenetics (2024)
-
Preneoplastic cells switch to Warburg metabolism from their inception exposing multiple vulnerabilities for targeted elimination
Oncogenesis (2024)
-
Lysosomes as coordinators of cellular catabolism, metabolic signalling and organ physiology
Nature Reviews Molecular Cell Biology (2024)
-
UCP2 and pancreatic cancer: conscious uncoupling for therapeutic effect
Cancer and Metastasis Reviews (2024)
-
Deciphering cellular plasticity in pancreatic cancer for effective treatments
Cancer and Metastasis Reviews (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.