Abstract
Sequencing studies of breast tumour cohorts have identified many prevalent mutations, but provide limited insight into the genomic diversity within tumours. Here we developed a whole-genome and exome single cell sequencing approach called nuc-seq that uses G2/M nuclei to achieve 91% mean coverage breadth. We applied this method to sequence single normal and tumour nuclei from an oestrogen-receptor-positive (ER+) breast cancer and a triple-negative ductal carcinoma. In parallel, we performed single nuclei copy number profiling. Our data show that aneuploid rearrangements occurred early in tumour evolution and remained highly stable as the tumour masses clonally expanded. In contrast, point mutations evolved gradually, generating extensive clonal diversity. Using targeted single-molecule sequencing, many of the diverse mutations were shown to occur at low frequencies (<10%) in the tumour mass. Using mathematical modelling we found that the triple-negative tumour cells had an increased mutation rate (13.3×), whereas the ER+ tumour cells did not. These findings have important implications for the diagnosis, therapeutic treatment and evolution of chemoresistance in breast cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Torres, L. et al. Intratumor genomic heterogeneity in breast cancer with clonal divergence between primary carcinomas and lymph node metastases. Breast Cancer Res. Treat. 102, 143–155 (2007)
Navin, N. et al. Inferring tumor progression from genomic heterogeneity. Genome Res. 20, 68–80 (2010)
Park, S. Y., Gonen, M., Kim, H. J., Michor, F. & Polyak, K. Cellular and genetic diversity in the progression of in situ human breast carcinomas to an invasive phenotype. J. Clin. Invest. 120, 636–644 (2010)
Sørlie, T. et al. Gene expression patterns of carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl Acad. Sci. USA 98, 10869–10874 (2001)
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012)
Shah, S. P. et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486, 395–399 (2012)
The Cancer Genome Atlas Network Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012)
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012)
Ellis, M. J. et al. Whole-genome analysis informs breast cancer response to aromatase inhibition. Nature 486, 353–360 (2012)
Schmitt, M. W. et al. Detection of ultra-rare mutations by next-generation sequencing. Proc. Natl Acad. Sci. USA 109, 14508–14513 (2012)
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011)
Woyke, T. et al. One bacterial cell, one complete genome. PLoS ONE 5, e10314 (2010)
Dichosa, A. E. et al. Artificial polyploidy improves bacterial single cell genome recovery. PLoS ONE 7, e37387 (2012)
Hou, Y. et al. Single-cell exome sequencing and monoclonal evolution of a JAK2-negative myeloproliferative neoplasm. Cell 148, 873–885 (2012)
Klein, C. A. et al. Comparative genomic hybridization, loss of heterozygosity, and DNA sequence analysis of single cells. Proc. Natl Acad. Sci. USA 96, 4494–4499 (1999)
Adey, A. et al. Rapid, low-input, low-bias construction of shotgun fragment libraries by high-density in vitro transposition. Genome Biol. 11, R119 (2010)
Kytola, S. et al. Chromosomal alterations in 15 breast cancer cell lines by comparative genomic hybridization and spectral karyotyping. Genes Chromosomes Cancer 28, 308–317 (2000)
Baslan, T. et al. Genome-wide copy number analysis of single cells. Nature Protocols 7, 1024–1041 (2012)
Zong, C., Lu, S., Chapman, A. R. & Xie, X. S. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338, 1622–1626 (2012)
Lorenz, M. O. Methods of measuring the concentration of wealth. J. Am. Stat. Assoc. 9, 209–219 (1905)
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nature Methods 7, 248–249 (2010)
Ng, P. C. & Henikoff, S. SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res. 31, 3812–3814 (2003)
Kuroishi, T. et al. Tumor growth rate and prognosis of breast cancer mainly detected by mass screening. Jpn. J. Cancer Res. 81, 454–462 (1990)
Peer, P. G., van Dijck, J. A., Hendriks, J. H., Holland, R. & Verbeek, A. L. Age-dependent growth rate of primary breast cancer. Cancer 71, 3547–3551 (1993)
Michaelson, J. et al. Estimates of breast cancer growth rate and sojourn time from screening database information. J. Women’s Imaging 5, 11–19 (2003)
Nachman, M. W. & Crowell, S. L. Estimate of the mutation rate per nucleotide in humans. Genetics 156, 297–304 (2000)
Drake, J. W., Charlesworth, B., Charlesworth, D. & Crow, J. F. Rates of spontaneous mutation. Genetics 148, 1667–1686 (1998)
Preston, B. D., Albertson, T. M. & Herr, A. J. DNA replication fidelity and cancer. Semin. Cancer Biol. 20, 281–293 (2010)
Baca, S. C. et al. Punctuated evolution of prostate cancer genomes. Cell 153, 666–677 (2013)
Hicks, J. et al. Novel patterns of genome rearrangement and their association with survival in breast cancer. Genome Res. 16, 1465–1479 (2006)
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011)
Pellman, D. Cell biology: aneuploidy and cancer. Nature 446, 38–39 (2007)
McClintock, B. The stability of broken ends of chromosomes in Zea mays. Genetics 26, 234–282 (1941)
Loeb, L. A. Human cancers express mutator phenotypes: origin, consequences and targeting. Nature Rev. Cancer 11, 450–457 (2011)
Merlo, L. M. F., Pepper, J. W., Reid, B. J. & Maley, C. C. Cancer as an evolutionary and ecological process. Nature Rev. Cancer 6, 924–935 (2006)
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012)
Luria, S. E. & Delbruck, M. Mutations of bacteria from virus sensitivity to virus resistance. Genetics 28, 491–511 (1943)
Bielas, J. H., Loeb, K. R., Rubin, B. P., True, L. D. & Loeb, L. A. Human cancers express a mutator phenotype. Proc. Natl Acad. Sci. USA 103, 18238–18242 (2006)
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013)
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013)
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
Wang, J. et al. CREST maps somatic structural variation in cancer genomes with base-pair resolution. Nature Methods 8, 652–654 (2011)
Futreal, P. A. et al. A census of human cancer genes. Nature Rev. Cancer 4, 177–183 (2004)
Hsu, F. et al. The UCSC known genes. Bioinformatics 22, 1036–1046 (2006)
Grubor, V. et al. Novel genomic alterations and clonal evolution in chronic lymphocytic leukemia revealed by representational oligonucleotide microarray analysis (ROMA). Blood 113, 1294–1303 (2009)
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010)
Forbes, S. A. et al. COSMIC: mining complete cancer genomes in the catalogue of somatic mutations in cancer. Nucleic Acids Res. 39, D945–D950 (2011)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987)
Acknowledgements
We thank L. Ramagli, H. Tang, E. Thompson, K. Khanna, W. Schober and J. Tyler. We are grateful to S. Kennedy and L. Loeb for help with the duplex protocols. We thank M. Edgerton, J. Hicks, M. Wigler and J. Kendall for discussions. We thank R. Krahe and M. Rui for reviewing the manuscript. N.E.N. is a Nadia’s Gift Foundation Damon Runyon-Rachleff Innovator (DRR-25-13). This research was supported by grants to N.E.N. from NIH (R21CA174397-01) and NCI (1RO1CA169244-01). N.E.N. was supported by T.C. Hsu and the Alice-Reynolds Kleberg Foundation. N.E.N. and P.S. were supported by the Center for Genetics & Genomics. F.M.-B was supported by an NIH UL1 (TR000371) and Susan Komen (SAC10006). K.C. was supported by the NCI (RO1CA172652). H.L. was supported by the NIH (U24CA143883). F.M. was supported by PS-OC (U54CA143798). K.C. and H.L. were supported by the Dell Foundation. M.L.L. is a CPRIT scholar and is supported by ALA. This work was also supported by an NCI center grant (CA016672). A.U. is a Rosalie B. Hite Fellow.
Author information
Authors and Affiliations
Contributions
Y.W. performed experiments and data analysis. M.L.L., J.W., A.M. and X.S. performed experiments. A.U., W.R., K.C., H.L., P.S. and S.V. performed data and statistical analyses. H.Z. and F.M.-B. obtained clinical samples. R.Z. and F.M. performed modelling. N.E.N. performed experiments, analysed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
The data from this study has been deposited into the Sequence Read Archive (SRA053195).
Extended data figures and tables
Extended Data Figure 1 Nuc-seq method.
a, Nuclear suspensions were prepared and stained with DAPI for flow-sorting, showing distributions of ploidy. The G2/M distribution was gated and single nuclei were deposited into wells. b, Cells were lysed and incubated with the Φ29 polymerase to perform multiple-displacement-amplification for a limited isothermal time-frame. c, d, Sequence libraries were prepared using one of two methods: Tn5 tagmentation (c), or low-input TA ligation cloning (d) (see Methods). e, Exome capture was optionally performed to isolate gDNA in exonic regions. f, Libraries were sequenced on the Illumina HiSeq2000 system. g, Somatic mutations were detected using a custom processing pipeline (Methods).
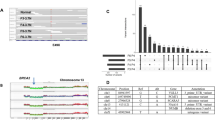
Extended Data Figure 2 Evaluation of WGA efficiency using chromosome-specific primers.
Whole genome amplified DNA from each single cell was used to perform PCR quality control experiments to determine WGA efficiency. For each cell, 22 reactions were performed using primer pairs that target each autosome and the resulting 200 bp PCR product were separated by gel electrophoresis (Methods). a, Two single nuclei were flow-sorted from the G2/M gate and amplified to WGA followed by PCR using 22 primer pairs. b, Two single nuclei were flow-sorted from the G1/0 gate and subject to WGA followed by PCR using 22 primer pairs. PCR products that failed to amplify are marked with an ‘x’ on the gel.
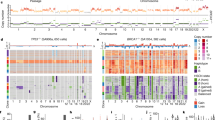
Extended Data Figure 3 Clustered heatmaps of single cell copy number profiles.
Single cell segmented copy number profiles were clustered and used to build heatmaps, showing amplifications in red and deletions in blue. a, Copy number profiles of 50 single cells from the ERBC. b, Copy number profiles of 50 single cells from the TNBC patient.
Extended Data Figure 4 Duplex single-molecule targeted deep-sequencing.
a, Experimental protocol for generating duplex libraries from bulk tumour DNA for custom capture and targeted ultra-deep sequencing. b, Data processing pipeline for duplex data to generate single-molecule data and detect mutation frequencies. c, Distribution of unique molecule tag duplicates for the ER breast cancer patient d, Distribution of unique molecule tag duplicates for the TNBC. e, Single-molecule coverage depth distribution for the ER+ tumour data. f, Single-molecule coverage depth distribution for the TNBC data.
Extended Data Figure 5 TNBC Multi-dimensional scaling and protein prediction plots.
a, Multi-dimensional scaling plot of the nonsynonymous mutations from the single-nuclei exome sequencing data in the TNBC b, Polyphen and SIFT protein impact prediction scores for the subclonal mutations in the TNBC patient.
Extended Data Figure 6 Models of clonal evolution in breast cancer.
a, Clonal evolution in the ERBC inferred from single cell exome and copy number data. b, Clonal evolution in the TNBC inferred from single cell exome and copy number data.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-8. (PDF 854 kb)
Rights and permissions
About this article
Cite this article
Wang, Y., Waters, J., Leung, M. et al. Clonal evolution in breast cancer revealed by single nucleus genome sequencing. Nature 512, 155–160 (2014). https://doi.org/10.1038/nature13600
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13600
This article is cited by
-
scAbsolute: measuring single-cell ploidy and replication status
Genome Biology (2024)
-
Basal–epithelial subpopulations underlie and predict chemotherapy resistance in triple-negative breast cancer
EMBO Molecular Medicine (2024)
-
Selective inhibition of CDK9 in triple negative breast cancer
Oncogene (2024)
-
Single-cell lineage tracing with endogenous markers
Biophysical Reviews (2024)
-
Integrative multi-region molecular profiling of primary prostate cancer in men with synchronous lymph node metastasis
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.