Abstract
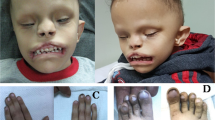
CHARGE syndrome is a multiple anomaly disorder in which patients present with a variety of phenotypes, including ocular coloboma, heart defects, choanal atresia, retarded growth and development, genitourinary hypoplasia and ear abnormalities1. Despite 70–90% of CHARGE syndrome cases resulting from mutations in the gene CHD7, which encodes an ATP-dependent chromatin remodeller, the pathways underlying the diverse phenotypes remain poorly understood2. Surprisingly, our studies of a knock-in mutant mouse strain that expresses a stabilized and transcriptionally dead variant of the tumour-suppressor protein p53 (p5325,26,53,54)3, along with a wild-type allele of p53 (also known as Trp53), revealed late-gestational embryonic lethality associated with a host of phenotypes that are characteristic of CHARGE syndrome, including coloboma, inner and outer ear malformations, heart outflow tract defects and craniofacial defects. We found that the p5325,26,53,54 mutant protein stabilized and hyperactivated wild-type p53, which then inappropriately induced its target genes and triggered cell-cycle arrest or apoptosis during development. Importantly, these phenotypes were only observed with a wild-type p53 allele, as p5325,26,53,54/− embryos were fully viable. Furthermore, we found that CHD7 can bind to the p53 promoter, thereby negatively regulating p53 expression, and that CHD7 loss in mouse neural crest cells or samples from patients with CHARGE syndrome results in p53 activation. Strikingly, we found that p53 heterozygosity partially rescued the phenotypes in Chd7-null mouse embryos, demonstrating that p53 contributes to the phenotypes that result from CHD7 loss. Thus, inappropriate p53 activation during development can promote CHARGE phenotypes, supporting the idea that p53 has a critical role in developmental syndromes and providing important insight into the mechanisms underlying CHARGE syndrome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Davenport, S. L. H., Hefner, M. A. & Mitchell, J. A. The spectrum of clinical features in CHARGE syndrome. Clin. Genet. 29, 298–310 (1986)
Jongmans, M. C. J. et al. CHARGE syndrome: the phenotypic spectrum of mutations in the CHD7 gene. J. Med. Genet. 43, 306–314 (2006)
Brady, C. A. et al. Distinct p53 transcriptional programs dictate acute DNA-damage responses and tumor suppression. Cell 145, 571–583 (2011)
Marine, J. C. et al. Keeping p53 in check: essential and synergistic functions of Mdm2 and Mdm4. Cell Death Differ. 13, 927–934 (2006)
Johnson, T. M., Hammond, E. M., Giaccia, A. & Attardi, L. D. The p53QS transactivation-deficient mutant shows stress-specific apoptotic activity and induces embryonic lethality. Nature Genet. 37, 145–152 (2005)
Brock, K. E., Mathiason, M. A., Rooney, B. L. & Williams, M. S. Quantitative analysis of limb anomalies in CHARGE syndrome: correlation with diagnosis and characteristic CHARGE anomalies. Am. J. Med. Genet. A. 123A, 111–121 (2003)
Zentner, G. E., Layman, W. S., Martin, D. M. & Scacheri, P. C. Molecular and phenotypic aspects of CHD7 mutation in CHARGE syndrome. Am. J. Med. Genet. A. 152A, 674–686 (2010)
Lalani, S. R. et al. Spectrum of CHD7 mutations in 110 individuals with CHARGE syndrome and genotype–phenotype correlation. Am. J. Hum. Genet. 78, 303–314 (2006)
Ragan, D. C., Casale, A. J., Rink, R. C., Cain, M. P. & Weaver, D. D. Genitourinary anomalies in the CHARGE association. J. Urol. 161, 622–625 (1999)
Inoue, H. et al. Successful cord blood transplantation for a CHARGE syndrome with CHD7 mutation showing DiGeorge sequence including hypoparathyroidism. Eur. J. Pediatr. 169, 839–844 (2010)
Corsten-Janssen, N. et al. The cardiac phenotype in patients with a CHD7 mutation. Circ. Cardiovasc. Genet. 6, 248–254 (2013)
Rinon, A. et al. p53 coordinates cranial neural crest cell growth and epithelial–mesenchymal transition/delamination processes. Development 138, 1827–1838 (2011)
Armstrong, J. F., Kaufman, M. H., Harrison, D. J. & Clarke, A. R. High-frequency developmental abnormalities in p53-deficient mice. Curr. Biol. 5, 931–936 (1995)
Lengner, C. J. et al. Osteoblast differentiation and skeletal development are regulated by Mdm2–p53 signaling. J. Cell Biol. 172, 909–921 (2006)
Sanlaville, D. et al. Phenotypic spectrum of CHARGE syndrome in fetuses with CHD7 truncating mutations correlates with expression during human development. J. Med. Genet. 43, 211–217 (2006)
Legendre, M. et al. Antenatal spectrum of CHARGE syndrome in 40 fetuses with CHD7 mutations. J. Med. Genet. 49, 698–707 (2012)
Veprintsev, D. B. & Fersht, A. R. Algorithm for prediction of tumour suppressor p53 affinity for binding sites in DNA. Nucleic Acids Res. 36, 1589–1598 (2008)
Bajpai, R. et al. CHD7 cooperates with PBAF to control multipotent neural crest formation. Nature 463, 958–962 (2010)
Schnetz, M. P. et al. CHD7 targets active gene enhancer elements to modulate ES cell-specific gene expression. PLoS Genet. 6, e1001023 (2010)
Bosman, E. A. et al. Multiple mutations in mouse Chd7 provide models for CHARGE syndrome. Hum. Mol. Genet. 14, 3463–3476 (2005)
Hurd, E. et al. Loss of Chd7 function in gene-trapped reporter mice is embryonic lethal and associated with severe defects in multiple developing tissues. Mamm. Genome 18, 94–104 (2007)
Adams, M. E. et al. Defects in vestibular sensory epithelia and innervation in mice with loss of Chd7 function: implications for human CHARGE syndrome. J. Comp. Neurol. 504, 519–532 (2007)
Morimoto, A. K. et al. Absent semicircular canals in CHARGE syndrome: radiologic spectrum of findings. AJNR Am. J. Neuroradiol. 27, 1663–1671 (2006)
de Oca Luna, R. M., Wagner, D. S. & Lozano, G. Rescue of early embryonic lethality in mdm2-deficient mice by deletion of p53. Nature 378, 203–206 (1995)
Mendrysa, S. M. et al. mdm2 is critical for inhibition of p53 during lymphopoiesis and the response to ionizing irradiation. Mol. Cell. Biol. 23, 462–472 (2003)
Mendrysa, S. M. et al. Tumor suppression and normal aging in mice with constitutively high p53 activity. Genes Dev. 20, 16–21 (2006)
Tyner, S. D. et al. p53 mutant mice that display early ageing-associated phenotypes. Nature 415, 45–53 (2002)
Balow, S. A. et al. Knockdown of fbxl10/kdm2bb rescues chd7 morphant phenotype in a zebrafish model of CHARGE syndrome. Dev. Biol. 382, 57–69 (2013)
Corsten-Janssen, N. et al. More clinical overlap between 22q11.2 deletion syndrome and CHARGE syndrome than often anticipated. Mol. Syndromol. 4, 235–245 (2013)
Jones, N. C. et al. Prevention of the neurocristopathy Treacher Collins syndrome through inhibition of p53 function. Nature Med.14. 125–133 (2008)
Schwenk, F., Baron, U. & Rajewsky, K. A cre-transgenic mouse strain for the ubiquitous deletion of loxP-flanked gene segments including deletion in germ cells. Nucleic Acids Res. 23, 5080–5081 (1995)
Sato, T. & Bartunkova, S. Analysis of embryonic vascular morphogenesis. Methods Mol. Biol. 137, 223–233 (2000)
Ovchinnikov, D. Alcian blue/alizarin red staining of cartilage and bone in mouse. Cold Spring Harb. Protoc. http://dx.doi.org/10.1101/pdb.prot5170 (March 2009)
Hurd, E. A., Poucher, H. K., Cheng, K., Raphael, Y. & Martin, D. M. The ATP-dependent chromatin remodeling enzyme CHD7 regulates pro-neural gene expression and neurogenesis in the inner ear. Development 137, 3139–3150 (2010)
Lane, D. et al. Epitope analysis of the murine p53 tumour suppressor protein. Oncogene 12, 2461–2466 (1996)
Rada-Iglesias, A. et al. Epigenomic annotation of enhancers predicts transcriptional regulators of human neural crest. Cell Stem Cell 11, 633–648 (2012)
Kenzelmann Broz, D. et al. Global genomic profiling reveals an extensive p53-regulated autophagy program contributing to key p53 responses. Genes Dev. 27, 1016–1031 (2013)
Acknowledgements
We thank S. Spano-Mello, K. T. Bieging, N. Raj and M. Monje-Deisseroth for reading the manuscript and S. E. Artandi and T. Williams for discussion. We thank H. Chou for immunohistochemistry assistance; E. L. Van Nostrand, P. Lavori, and A. McMillian for statistical analysis; K. Weinberg and D. Min for thymus analysis assistance; M. Shkreli for kidney analysis assistance; B. Liu and J. A. Helms for craniofacial analysis assistance; and M. Bowen for Chd7 mouse experiment assistance. We thank S. E. Artandi for plasmids; S. E. Artandi and P. Khavari for control human fibroblast cell lines; D. Lane and B. Vojtesek for wild-type p53-specific antibody (pAB242); P. Scacheri for wild-type and Chd7-null mouse embryonic stem cells; and T. Denecker and G. Goudefroye for TP53 sequencing in patients. This work was supported by funding from the NSF and NCI (grant number 1F31CA167917-01) to J.L.V.N.; from the NIH (RO1 GM095555) to J.W.; from the American Heart Association (12EIA8960018), March of Dimes Foundation (#6-FY11-260) and NIH (R01 HL118087 and RO1 HL121197) to C.-P.C.; from the NIH (R01 DC009410) to D.M.M.; and from the ACS, LLS and NIH (RO1 CA140875) to L.D.A.
Author information
Authors and Affiliations
Contributions
J.L.V.N. designed and carried out experiments, interpreted data and wrote the manuscript. C.A.B. generated the p5325,26,53,54 mice, designed and carried out experiments, and interpreted data. H.J. performed p53 ChIP analyses. M.M.K. and T.M.J. performed certain mouse analyses. D.R.F. and J.W. performed NCC differentiation and CHD7 ChIP analyses. C-Y.L., C-J.L. and C-P.C. assisted with heart analyses. D.L.S. and D.M.M. performed inner ear analyses, interpreted data and provided human fibroblasts. H.V. assisted with histological analyses. J.A.B. generated CHARGE and control human fibroblast lines. T.A.-B. performed TP53 sequencing analysis in patients and supplied CHARGE thymus samples. L.D.A. designed experiments, interpreted data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Model for examining p53-associated developmental phenotypes.
a, Schematic of p5325,26, p5353,54 and p5325,26,53,54 mutant p53 proteins. Basic, basic-residue-rich domain; PRD, proline-rich domain; TAD, transcriptional activation domain 1 or 2; Tet, tetramerization domain. b, p53LSL-mut/+ mice (where mut can denote any of the p53 TAD mutants) were crossed to p53+/+; CMV-Cre mice, which express Cre in the germline, to assess the viability and developmental phenotypes of the p53-mutant-expressing progeny. c, Table summarizing the actual genotypes and ultimate functional genotypes of progeny from crosses of p53LSL-25,26,53,54/+ and p53+/−; CMV-Cre mice, as used throughout the manuscript. While p53LSL-25,26,53,54/+; CMV-Cre is the actual initial genotype, when Cre acts to delete the lox–Stop–lox (LSL) cassette, the genotype is written as p5325,26,53,54/+ to reflect this recombination. In the text and figure labels, the Cre nomenclature for both control and p5325,26,53,54/+ embryos is excluded for simplicity. Controls for analyses comprised embryos both with and without the CMV-Cre transgene, as summarized in Extended Data Fig. 3.
Extended Data Figure 2 p5325,26,53,54/− mice, but not p5325,26/+ or p5325,26/− mice, are viable.
a, Crosses of p53LSL-25,26/+ with p53+/+; CMV-Cre or p53+/−; CMV-Cre mice revealed a decrease in viable pups expressing p5325,26 at E9.5–E10.5. The observed numbers of live and dead pups compared with the expected numbers of live pups are indicated: [Observed (Expected)]. The genotypes of the p5325,26/+ and p5325,26/− mice carrying a CMV-Cre transgene lack the LSL designation because the lox–Stop–lox element has been deleted from the genome. Significance was assessed by binomial distribution statistical tests on live pups: P = 0.18 for the p53+/+ cross and 0.09 for the p53+/− cross. b, Crosses of p53LSL-25,26/+ or p53LSL-25,26,53,54/+ with p53+/−; CMV-Cre mice revealed that p5325,26,53,54/− mice, but not p5325,26/− mice, are viable, as assessed at postnatal day 21 (P21). mut denotes either mutant allele. The observed numbers of pups are indicated and compared with the expected numbers of pups: [Observed (Expected)]. The genotypes of p53mut/+ and p53mut/−mice carrying a CMV-Cre transgene lack the LSL designation because the lox–Stop–lox element has been deleted from the genome. The lack of pups is significant at P21 as assessed by binomial distribution statistical tests on live pups compared with expected: p5325,26/+ and p5325,26/−, * P = 3.42 × 10−5; p5325,26,53,54/+, ** P = 4.77 × 10−7. c, Whole-mount images of a p5325,26/+ embryo (right) at E9.5, displaying developmental delay (top) and neural tube defects, including exencephaly and kinked spine (bottom), compared with a littermate control (left). Original magnification, ×50. d, p5325,26,53,54/− mice displayed a shorter lifespan (median lifespan, 128 days; n = 8) than wild-type mice (median lifespan, 774 days) and a similar lifespan to p53−/− mice (median lifespan, 143 days), further indicating that the p5325,26,53,54 allele itself behaves like a p53-null allele; P < 0.0001 by Mantel–Cox statistical analysis comparing wild-type and p5325,26,53,54/− mice.
Extended Data Figure 3 Genotypes of the control embryos in the figures and the genders associated with phenotypes.
a, Table identifying the genotypes of the control embryos shown for each analysis. b, Table showing the number of male and female p5325,26,53,54/+ embryos observed for the indicated phenotypes, as assessed by Zfy PCR. The phenotypes are well represented in both sexes.
Extended Data Figure 4 p5325,26,53,54/+ embryos exhibit additional features of CHARGE syndrome.
a, Haematoxylin and eosin stained sections of E12.5 control (left) and p5325,26,53,54/+ embryos (right). Examination confirmed neural tube closure defects (arrow). Original magnification, ×10. b, Close-up image of an ultraviolet-radiation-illuminated, ethidium bromide stained E15.5 p5325,26,53,54/+ embryo (right), highlighting the short lower jaw phenotype with protruding tongue (arrow) compared with control littermates (left). Seventy-four per cent of p5325,26,53,54/+ embryos (n = 27) exhibited a short lower jaw. Cleft lip is not shown. Original magnification, ×40. c, Top, Alizarin red (bone) and Alcian blue (cartilage) stained whole mount of an E15.0 p5325,26,53,54/+ embryo (right), showing reduced bone density in the cranium (c), nasal cavity (n), shorter ulna (u), humerus (h), mandible (m) and femur (f), as well as reduced bone formation in the ribs (R), where fewer vertebrae are undergoing ossification than in littermate controls (left). The number of vertebrae with bone formation was 19 in the controls (arrow; V19) and 18 in p5325,26,53,54/+ embryos (arrow; V18). The severity of the bone and cartilage defects was variable, with the most severe defects evident in embryos with exencephaly and severe craniofacial defects (n = 7). Bottom, Quantification of bone lengths shown as percentage of E14.5–15.0 littermate controls. The bone lengths of the mandible, humerus, ulna and femur were measured using the ruler function in Adobe Photoshop on images acquired at 6.3× magnification. Only litters with detectable bone formation in p5325,26,53,54/+ embryos were included in bone length analyses: Student’s t-test; **, P = 0.008 (mandible); **, P = 0.005 (humerus). d, Representative images of haematoxylin and eosin stained sagittal sections of E12.5 control hearts (left) and p5325,26,53,54/+ hearts (right), showing all three cardiac cell types in both genotypes. en, endocardium; ep, epicardium; myo, myocardium (arrows). Original magnification, ×200. e, A haematoxylin and eosin stained E12.5 p5325,26,53,54/+ heart exhibiting persistent truncus arteriosus (PTA) (33%, n = 6). The cardiac outflow tract in the control embryo (left) is septated into the aorta (Ao) and main pulmonary artery (MPA), whereas the cardiac outflow tract (truncus arteriosus (TA)) in the p5325,26,53,54/+ embryo (right) remains unseptated, resulting in PTA. Original magnification, ×100. f, Illustration of a control heart (left) and a p5325,26,53,54/+ embryo heart (right), highlighting double outlet right ventricle (DORV) and atrioventricular cushion defects. Both the aorta (Ao) and the MPA flow out of the right ventricle (RV), resulting in mixed oxygenated and deoxygenated blood in the systemic circulation when combined with concurrent ventricular septal defects (VSDs). The atrioventricular cushions remain bulbous and fail to elongate into mature valve leaflets. Red denotes oxygenated blood; blue denotes deoxygenated blood; and purple (pink) denotes mixed oxygenated and deoxygenated blood. mv, mitral valve; tv, tricuspid valve. Original magnification, ×100. g, Representative haematoxylin and eosin stained transverse section of thymus from a p5325,26,53,54/+ E15.5 embryo (right) reveals a smaller thymus than in littermate controls (left) (63% of controls; n = 4). Original magnification, ×200. h, Representative haematoxylin and eosin analysis of liver sections from E12.5 controls (left) and p5325,26,53,54/+ embryos (right), showing normal liver architecture in both genotypes (top). High magnification image (bottom) of the region of the liver that is outlined by the white box in the top panel shows the presence of nucleated erythrocytes (arrows), indicating proper haematopoiesis. Original magnification top, ×100; bottom, ×400. i, Top, Table summarizing the incidence (%) and sample size (n) of phenotypes assessed qualitatively in p5325,26,53,54/+ embryos. The occurrence of these phenotypes in CHARGE syndrome is also indicated (+, present; −, absent). Bottom, Table summarizing the phenotypes assessed quantitatively in p5325,26,53,54/+ embryos relative to controls, shown as the percentage average size of the control (%), with sample size (n) also indicated. The occurrence of these phenotypes in CHARGE syndrome is also shown (+, present). A detailed description of the bone and cartilage defects is provided in c.
Extended Data Figure 5 p53−/− embryos do not exhibit characteristics of CHARGE syndrome.
a, Whole-mount image of the external ear of an E15.5 p53−/− embryo (right) and a control embryo (left), showing normal ear pinna development. Original magnification, ×100. b, Whole-mount image of an E13.5 p53−/− embryo (right) and a control embryo (left), showing normal retinal development and no evidence of coloboma. Original magnification, ×100. c, Whole-mount image of an E15.5 p53−/−embryo (right) and a control embryo (left), showing normal lower jaw development. Original magnification, ×40. d, Top, Alizarin red (bone) and Alcian blue (cartilage) stained whole-mount E14.5 p53−/−embryo (right), showing normal long bone formation of the ulna (u), humerus (h), mandible (m), and femur (f) relative to littermate controls (left). Bottom, Quantification of the bone lengths shown as a percentage of E14.5 littermate controls (n = 3). Original magnification, ×6.3. e, Representative images of haematoxylin and eosin stained sagittal sections of E13.5 control hearts (left) and p53−/− hearts (right), showing all three cardiac cell types in both genotypes. en, endocardium; ep, epicardium; myo, myocardium (arrows). Original magnification, ×200. f, Analysis of haematoxylin and eosin stained transverse sections of E13.5 p53−/− and control hearts revealing normal outflow tract development. Top, The MPA and Ao are fully septated, and the MPA connects to the right ventricle (RV) in p53−/− hearts. Bottom, The Ao connects to the left ventricle (LV). The symbol Φ denotes that the ventricular outflow tract connects the LV and AO. AV, aortic valve; PV, pulmonary valve. Original magnification, ×100. g, Analysis of transverse sections of haematoxylin and eosin stained E13.5 p53−/− hearts (right) reveals normal atrioventricular cushions, which have undergone remodelling to form mature, elongated mitral valves (mv; arrowhead) and tricuspid valves (tv; arrow) similar to in control hearts (left). LA, left atrium; RA, right atrium. Original magnification, ×100. h, Haematoxylin and eosin stained sagittal section of kidney from p53−/− (right) and control (left) embryos, showing normal renal size and development. Original magnification, ×200. i, Haematoxylin and eosin stained transverse section of thymus in a p53−/− E13.5 embryo (right) reveals a similar thymus size to that in a littermate control (left). Original magnification, ×200.
Extended Data Figure 6 p5325,26,53,54/+ embryo tissues display increased apoptosis and decreased proliferation.
a, Left, Immunofluorescence for phospho-histone H3 (red) in the retina of E13.5 control and p5325,26,53,54/+ embryos. Right, Quantification of phospho-H3-positive cells per retina area relative to littermate controls. **, P = 0.006 by one-tailed Welch’s t-test (n = 4). Original magnification, ×200. b, Left, Immunohistochemistry for cleaved caspase 3 (CC3) in thymi of control (left) and p5325,26,53,54/+ (right) embryos. Inset, Magnified image of CC3-positive region. Right, Quantification of CC3-positive cells per thymic area. *, P = 0.02 by one-tailed Student’s t-test (n = 4). Original magnification, ×400. c, Immunofluorescence for PAX3 (green) in NCCs of E9.5 control and p5325,26,53,54/+ embryos was used to identify NCCs in Fig. 2f. Original magnification, ×200. d, Left, Immunofluorescence for CC3 (red) and PAX3 (green) in NCCs of E9.5 control and p5325,26,53,54/+ embryos. p5325,26,53,54/+ embryos have more apoptotic (red) NCCs, as determined by PAX3-positive staining (green), than littermate controls. Right, Quantification of CC3-positive cells per total NCC number. P = 0.14 by one-tailed Student’s t-test (n = 4). Original magnification, ×200. e, Left, Immunofluorescence for CC3 (red) in the otic vesicles of E9.5 control and p5325,26,53,54/+ embryos. Right, Quantification of CC3-positive cells per total cell number. *, P = 0.03 by one-tailed Student’s t-test (n = 3). Original magnification, ×200. f, CC3 staining in whole-mount E8.5 control and p5325,26,53,54/+ embryos reveals enhanced apoptosis in the neuroepithelium of p5325,26,53,54/+ embryos (right) but not in controls (left). The close-up shows a magnification of the caudal neuroepithelium (bottom). Arrows indicate CC3-positive regions. Original magnification top, ×50.
Extended Data Figure 7 p5325,26,53,54 is transactivation dead but augments wild-type p53 activity.
a, Western blot analysis of p53 protein levels in untreated or doxorubicin-treated (0.2 μg ml−1 Dox) p53−/−, p53+/−, p5325,26,53,54/− and p5325,26,53,54/+ MEFs. β-Actin served as a loading control. b, Western blot analysis of anti-Flag immunoprecipitation from p53−/− MEFs transiently overexpressing HA–p53 and Flag–p53 or Flag–p5325,26,53,54. HA–MBP and Flag–eGFP were used as negative controls. Immunoprecipitated protein and 10% input were probed with either anti-HA or anti-Flag antibodies. (The microgram ratio of HA–p53 to Flag–p53 or Flag–p5325,26,53,54 plasmid DNA was 1:1 or 1:2.5, as indicated above the blot (see Fig. 3b). c, Heat map examining the transactivation capacity of p5325,26,53,54 on p53-dependent genes identified by microarray analysis through comparison of six HrasV12; WT p53 MEF lines with six HrasV12; p53-null MEF lines, as previously described3. Three independent HrasV12; p5325,26,53,54/25,26,53,54 MEF lines were analysed, and the gene expression profiles were indistinguishable from those of HrasV12; p53-null cells. The numbered columns indicate independent MEF lines. Blue denotes repressed genes, and red denotes induced genes. d, qRT–PCR analysis of p53 target gene expression in untreated MEFs derived from p53+/+ and p5325,26,53,54/+ E13.5 embryos. Graphs indicate the mean ± s.d. from four independent MEF lines after normalization to β-actin gene expression. **, P < 0.01; ***, P < 0.005; Student’s t-test. e, qRT–PCR analysis of p53 target gene expression in p53+/+ and p53−/− MEFs stably transduced with empty vector, Flag–p53 or Flag–p5325,26,53,54. The representative gene expression from one experiment is shown: mean ± s.d. of technical triplicates after normalization to β-actin gene expression. The experiment was performed in duplicate.
Extended Data Figure 8 p53 heterozygosity partially rescues Chd7-null embryos.
a, qRT–PCR analysis of Chd7 expression in untreated MEFs derived from E13.5 p53+/− and p5325,26,53,54/+ embryos. The graphs indicate the mean ± s.d. from four independent MEF lines after normalization to β-actin gene expression. ns, not significant. b, Left, Schematic of NCC differentiation. Right, Representative qRT–PCR analysis of NCC markers in NCC-like cells differentiated from Chd7+/+ and Chd7−/− (whi/whi) mouse embryonic stem cells normalized to β-actin gene expression and relative to matched embryonic stem cells. c, Haematoxylin and eosin stained E10.5 Chd7+/+p53+/− (control), Chd7−/−p53+/+ and Chd7−/−p53+/− embryos. The Chd7−/−p53+/+ embryo shown is necrotic, as evidenced by cellular autolysis. Original magnification, ×32. d, Close-up image of heart region denoted by red box in panel c, in E10.5 Chd7+/+p53+/− (control) and Chd7−/−p53+/− embryos. Original magnification, ×100.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-2. (PDF 115 kb)
Rights and permissions
About this article
Cite this article
Van Nostrand, J., Brady, C., Jung, H. et al. Inappropriate p53 activation during development induces features of CHARGE syndrome. Nature 514, 228–232 (2014). https://doi.org/10.1038/nature13585
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13585
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.