Abstract
Mesenchymal stem cells occupy niches in stromal tissues where they provide sources of cells for specialized mesenchymal derivatives during growth and repair1. The origins of mesenchymal stem cells have been the subject of considerable discussion, and current consensus holds that perivascular cells form mesenchymal stem cells in most tissues. The continuously growing mouse incisor tooth offers an excellent model to address the origin of mesenchymal stem cells. These stem cells dwell in a niche at the tooth apex where they produce a variety of differentiated derivatives. Cells constituting the tooth are mostly derived from two embryonic sources: neural crest ectomesenchyme and ectodermal epithelium2. It has been thought for decades that the dental mesenchymal stem cells3 giving rise to pulp cells and odontoblasts derive from neural crest cells after their migration in the early head and formation of ectomesenchymal tissue4,5. Here we show that a significant population of mesenchymal stem cells during development, self-renewal and repair of a tooth are derived from peripheral nerve-associated glia. Glial cells generate multipotent mesenchymal stem cells that produce pulp cells and odontoblasts. By combining a clonal colour-coding technique6 with tracing of peripheral glia, we provide new insights into the dynamics of tooth organogenesis and growth.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Keating, A. Mesenchymal stromal cells: new directions. Cell Stem Cell 10, 709–716 (2012)
Koussoulakou, D. S., Margaritis, L. H. & Koussoulakos, S. L. A curriculum vitae of teeth: evolution, generation, regeneration. Int. J. Biol. Sci. 5, 226–243 (2009)
Zhao, H. et al. Secretion of shh by a neurovascular bundle niche supports mesenchymal stem cell homeostasis in the adult mouse incisor. Cell Stem Cell 14, 160–173 (2014)
Caton, J. & Tucker, A. S. Current knowledge of tooth development: patterning and mineralization of the murine dentition. J. Anat. 214, 502–515 (2009)
Miletich, I. & Sharpe, P. T. Neural crest contribution to mammalian tooth formation. Birth Defect Res C 72, 200–212 (2004)
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010)
Luukko, K. et al. Secondary induction and the development of tooth nerve supply. Ann. Anat. 190, 178–187 (2008)
Laranjeira, C. et al. Glial cells in the mouse enteric nervous system can undergo neurogenesis in response to injury. J. Clin. Invest. 121, 3412–3424 (2011)
Adameyko, I. et al. Schwann cell precursors from nerve innervation are a cellular origin of melanocytes in skin. Cell 139, 366–379 (2009)
Hari, L. et al. Temporal control of neural crest lineage generation by Wnt/beta-catenin signaling. Development 139, 2107–2117 (2012)
Adameyko, I. et al. Sox2 and Mitf cross-regulatory interactions consolidate progenitor and melanocyte lineages in the cranial neural crest. Development 139, 397–410 (2012)
Leone, D. P. et al. Tamoxifen-inducible glia-specific Cre mice for somatic mutagenesis in oligodendrocytes and Schwann cells. Mol. Cell. Neurosci. 22, 430–440 (2003)
Feng, J., Mantesso, A., De Bari, C., Nishiyama, A. & Sharpe, P. T. Dual origin of mesenchymal stem cells contributing to organ growth and repair. Proc. Natl Acad. Sci. USA 108, 6503–6508 (2011)
Crisan, M., Corselli, M., Chen, W. C. & Peault, B. Perivascular cells for regenerative medicine. J. Cell. Mol. Med. 16, 2851–2860 (2012)
Adameyko, I. & Lallemend, F. Glial versus melanocyte cell fate choice: Schwann cell precursors as a cellular origin of melanocytes. Cell. Mol. Life Sci. 67, 3037–3055 (2010)
Chung, I. H. et al. Stem cell property of postmigratory cranial neural crest cells and their utility in alveolar bone regeneration and tooth development. Stem Cells 27, 866–877 (2009)
Jinno, H. et al. Convergent genesis of an adult neural crest-like dermal stem cell from distinct developmental origins. Stem Cells 28, 2027–2040 (2010)
Dewachter, I. et al. Neuronal deficiency of presenilin 1 inhibits amyloid plaque formation and corrects hippocampal long-term potentiation but not a cognitive defect of amyloid precursor protein [V717I] transgenic mice. J Neurosci. 22, 3445–3453 (2002)
Acknowledgements
P. Kovaleva and O. Rogachevskaya helped with illustrations. This study was supported by the Swedish Research Council (I.A., K.F., P.E.), The Bertil Hållsten Research Foundation (I.A.), StratRegen and the Wallenberg Foundation (CLICK, I.A.), the Swiss National Science Foundation (U.S.), Medical Research Council (G0901599; P.S.), a Wallenberg Scholar and European Research Council advanced grant (P.E.), National Graduate School in Odontological Science (M.K.S.), the Swedish Dental Association (N.K., M.K.S.), an EMBO Long-Term Fellowship (M.K.) and Stockholm County Council (N.K.) and Developmental Studies Hybridoma Bank.
Author information
Authors and Affiliations
Contributions
N.K., M.K.S., C.K., V.D., M.K., A.F., Z.A., L.W., P.S., I.H., I.A. and K.F. performed experiments, analysed data and wrote the paper. L.A.-R., H.B., H.B., N.A.L., V.P., U.S., H.C., P.S., I.T. and P.E. analysed data and wrote the paper. All authors read and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Expression of SCP markers and genetic tracing in nerves of E11.5–13.5 embryos.
a–c, Whole-mount immunohistochemistry performed on an E11.5 mouse embryo with antibodies against 155 kDa neurofilaments (2H3), MITF and Sox10. Note that Sox10+ cells are localized at the nerves in maxillary and mandibular regions. d–f, Whole-mount immunohistochemistry performed on an E11.5 mouse embryo with antibodies against 155 kDa neurofilaments (2H3) and BFABP. d–e, Arrows point at posterior ventral spinal nerves hosting immature embryonic glial cells that lack BFABP expression. f, Magnified area outlined in d by white rectangle. Roman digits show corresponding numbers of cranial ganglia. Dotted line follows the contour of an embryo. g–l, Sagittal section through E12.5 mandible stained for Sox10 (g–l), Tuj1 and BFABP (g), GFAP (h) and p75 (i). j–l, ErbB3 is expressed in all Sox10+ nerve-adjacent SCPs. k–l, Magnified area outlined by white rectangle in j. Dotted line indicates epithelial organ. m–o2, Sagittal section through E13.5 mandible, sequential single plane optical slices 2 µm in z-axis. m1 –m2 , Magnified area outlined in m; n1 , n2 , magnified area from n; o1 –o2 , respectively, show an area from o. p–p2 , Region on a section shown in m–o, maximum intensity projection image. p1 , p2 , Magnified area that is outlined by white rectangle in p. m–p, White dotted line indicates incisor placode. q–t, Expression of CreERT2, Sox10 and Tuj1 in a mandible traced from E12.5 to E13.5 in PLP-CreERT2 embryo. Note the full co-localization of Sox10 and CreERT2. Sox10+/CreERT2+ cells are adjacent to the innervation. Scale bars, 100 µm (a–c, g–i); 500 µm (d–f); 50 µm (j–l); 50 µm (m–t).
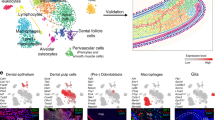
Extended Data Figure 2 Expression of Sox10, YFP and CreERT2 in a genetically traced mouse embryonic head.
a–d, Expression of CreERT2 protein in a 36 h genetically traced PLP-CreERT2/R26YFP embryo at E13.5. c, d, Magnified areas outlined in a and b. Open arrows point at CreERT2+/YFP+ cells attached to the nerve whereas filled arrow points at CreERT2−/YFP+ cell proximal to the nerve. e, f, In situ hybridization with Sox10 riboprobe (e) on a section of a mouse embryonic E12.5 head post-stained with Tuj1 (β-III-tubulin, neuronal marker) antibody (f). g–h, In situ with Sox10 probe on a section of E12.5 developing mandible (g) with incisor tooth bud (outlined by dotted line) post-stained with Tuj1 antibody (h). i–j, Sox10 and PGP9.5 (marker for neurites) are visualized by immunohistochemistry on a section of an embryonic mandible at E12.5. Dotted line outlines incisor bud. k–l, Sections of developing mandible with incisor (k) and molar (l) buds from a Sox10-CreERT2/R26YFP embryo genetically traced from E12.5 to E13.5 and stained with antibody against the neuronal marker PGP9.5. Arrow in l points at developing cartilage. m–p, Expression of CreERT2 protein in a 24 h genetically traced Sox10-CreERT2/R26YFP embryo at E13.5. The developing Meckel’s cartilage is outlined by the red dotted circle in m and o. n, p, Magnified areas outlined in m and o. White dotted line shows the borders of tooth bud. Scale bars, 200 µm (e–h); 100 µm (i, m, o); 50 µm (a, b, j–l, n–p); 10 µm (c–d).
Extended Data Figure 3 Expression of Sox10 and CreERT2 at intermediate stages of incisor development.
a–g, Incisor traced from E12.5 to E15.5 in PLP-CreERT2/R26YFP animals. Enamel organ is outlined by dotted line. a, b, YFP+ cells in the nerve and in the pulp are shown together with Ki67 staining. c–g, Note that the expression of CreERT2 is confined to nerve sites only, as shown in low (c, d) and high (e–g) magnification images of developing incisor. e–g, Area from c, d shown at high magnification. YFP+/CreERT2+ cells are indicated by arrows in e–g. Note the presence of CreERT2−/YFP+ proximally to the nerve and in the apical mesenchyme. h–k, Incisor traced from E15.5 to E16.5 in Sox10-CreERT2/R26YFP animals. Panel i represents a magnified area from h. j, k, Magnified area outlined by white rectangle in i. Arrows in j and k point at CreERT2−/YFP+ cells in the apical mesenchyme; note that the expression of CreERT2 is confined to the nerve sites only. l–m, Developing incisor at E15.5 stained with antibodies against Sox10 to show SCPs, Tuj1 to visualize nerves and Col IV to outline the position of vessels. Note that all Sox10+ cells are nerve adjacent. n, Schematic drawing showing the position of Sox10+ cells during early bell stage of tooth development. Scale bars, 50 µm (a–d, h, i, l, m); 25 µm (e–g, j, k). CL1 and CL2 are the labial and lingual aspects of the cervical loop respectively.
Extended Data Figure 4 Sub-optimal genetic recombination highlights clonal relationships between pulp cells and odontoblasts during tooth organogenesis.
a–c, Consecutive sections through the incisor from PLP-CreERT2/R26Confetti mouse embryo traced from E12.5 to E17.5 (sub-optimal recombination). Note the pulp and odontoblasts progenies of CFP+ SCPs (white arrows in a and b). c, Section throughout the same tooth where cyan arrows point at columnar CFP+ odontoblasts. d, Section through another clonally traced (sub-optimal recombination) incisor. Note the presence of YFP+ clone in the pulp. Arrow points at a single YFP+ cell positioned at the innervation site. e, Reconstructed lineage tree of the neural-crest-derived compartment in the tooth. a–d, Scale bars, 100 µm. CL1 and CL2 are the labial and lingual aspects of the cervical loop respectively.
Extended Data Figure 5 Distribution of Sox10+ cells in adult incisor.
a, Localization of Sox10+ cells and Tuj1+ nerve fibres in the pulp of adult incisor. Note that apical mesenchyme between the aspects of a cervical loop contains a population of nerve-adjacent Sox10+ cells while the distal pulp harbours an additional population of Sox10+ cells (outlined by a yellow dotted line) that are not adjacent to the nerves. b–e, Magnified areas from a where they are outlined by numbered white rectangles. b, Magnified image of apical mesenchyme and cervical loop area with nerve-adjacent Sox10+ cells pointed out by arrows. c, Region in the distal pulp showing scattered Sox10+ cells. Note that these cells are not in contact with Tuj1+ fibres. d, Cervical loop area including a region of proximal pulp and apical mesenchyme. e, Neural bundle in the apical mesenchyme with nerve-adjacent Sox10+ cells pointed out by arrows (projection of a stack). f–k, Localization of Sox10+ cells in relation to the proliferative growth zone outlined by the expression of Ki67. g–h, Magnified cervical loop area outlined in f by white rectangle 1. Arrows point at detected Sox10+ cells. i–j, Magnified area shown in f by white rectangle 2. Arrows point at Ki67+ cells in the distal pulp. Note that Ki67 does not label the distal population of Sox10+ cells outlined by yellow dotted line in f. k, Schematic drawing showing two separated populations of Sox10+ cells in the pulp: one population is represented by nerve-adjacent cells and located in proliferative apical mesenchyme while another population is scattered in distal pulp at a significant distance from a cervical loop. Scale bars, 100 µm (a, b, f); 50 µm (c–e, g–j). CL1 and CL2 are the labial and lingual aspects of the cervical loop respectively. White dotted line shows enamel organ epithelium.
Extended Data Figure 6 Expression of Schwann cell markers and CreERT2 in adult incisor.
a, Schematic drawing showing the position of a focus area (red frame) selected for demonstration of Schwann cell markers in b–g. Schwann cell markers are expressed in Sox10+ nerve-adjacent cells located in apical mesenchyme: S100β and P0 in b, P0 and p-c-Jun in c, p75 and Krox20 in d, MBP in e and GFAP and P0 in f–g. h–q, Expression of CreERT2 under the control of Sox10- (o, p) and PLP1- (q–u) promoters in adult incisor. h, i, Note that CreERT2 protein is found in cells adjacent to the Tuj1+ nerve fibres in the apical mesenchyme of 30-day traced Sox10-CreERT2/R26YFP animals. j–q, CreERT2 protein is detected in the nuclei of Sox10+ cells (58 ± 7.9% of nerve-adjacent Sox10+ cells are CreERT2+, n = 5) in the apical mesenchyme of 20-h traced PLP-CreERT2/R26YFP animals. l–n, Magnified area outlined in j, k. Arrows point at CreERT2+ nuclei. o, p, Expression of CreERT2 is not detected in a population of Sox10+ non-glial cells in distal pulp. q, Expression of CreERT2 is not found in a cervical loop area. Scale bars, 50 µm. CL1 and CL2 are the labial and lingual aspects of the cervical loop respectively. Dotted line shows enamel organ epithelium.
Extended Data Figure 7 Short and long tracing intervals reveal YFP+ cells at initial positions and uncover the later contribution of Schwann-cell-derived cells to the growing incisor.
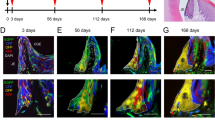
a–e, Section through adult incisor traced for 2 days from a Sox10-CreERT2/R26YFP mouse stained for collagen IV and Tuj1 to visualize blood vessels and nerves, respectively. b, c, Magnified area outlined by rectangle 1 in a. Arrows point at YFP+ cells adjacent to the innervation of a vascular bundle between cervical loops. d, e, YFP+ cells (arrows) also appear adjacent to the nerve fibres of vascular bundle distally from cervical loop (position is outlined by rectangle 2 in a). Images represent maximum intensity projections of confocal stacks. e, f, Incisor after 5 days of genetic tracing in a PLP-CreERT2/R26YFP animal, sequential sections. Note that YFP+ progeny is located at the apex as indicated by the arrows; YFP+ cells are not detected in the distal pulp. Dotted line outlines enamel epithelium and mineralized matrix. i–k, Ki67 labelling of a section of an incisor traced for 6 months from an adult PLP-CreERT2/R26YFP mouse, single tamoxifen injection at sub-optimal concentration. i, Arrows point at a cluster of YFP+ odontoblasts (magnified in inset). Note the presence of YFP+ streams of cells in the pulp after 6 months of genetic tracing. j, k, YFP+ genetically traced pulp cells are positive for Ki67 only in proximity to the cervical loops. k, Magnified area outlined by white rectangle in j. Note the absence of Ki67+ cells in the odontoblast layer and in pulp cells at a distance from the cervical loop (CL). l, Scheme of genetic tracing experiments involving single and serial tamoxifen injections. m, n, Segmentation of odontoblast and pulp cell nuclei in the adult genetically traced incisor (m) injected eight times with subsequent identification of YFP+ nuclei (n) for semi-automated counting in IMARIS. Magnified areas outlined by rectangles are shown in the insets; m.p.i., months post-injection. i–n, CL1 is a labial cervical loop; dotted line outlines enamel epithelium with adjacent hard matrix. o, Contribution of Schwann-cell-derived cells to the incisors from single and multiple injected PLP-CreERT2/R26YFP animals (n = 3 for each type of experiment). For the quantification of contribution to the odontoblast lineage, only labial odontoblasts were analysed. a–k, Scale bars, 100 µm (a, f–j); 50 µm (b–e, k); 100 µm (i–j); 25 µm (k). CL1 and CL2 are the labial and lingual aspects of the cervical loop respectively.
Extended Data Figure 8 Clonal analysis reveals diversity of progeny originating from different Schwann-cell-derived MSCs that are nerve dependent.
a–f, Consecutive sections of an incisor traced for 1.5 months from a PLP-CreERT2/R26Confetti mouse (sub-optimal recombination). b, d, f, Magnified areas from a, c and e respectively. Note numerous dispersed RFP+ pulp cells in a–d and large amounts of adjacent RFP+ odontoblasts in c–f. g–i, Another example of an incisor traced for 1.5 months (g) from a PLP-CreERT2/R26Confetti mouse also stained for PGP9.5 to visualize nerve fibres. h, i, Magnified areas from g. Note the narrow stream of RFP+ cells in the middle of the dental pulp in g and h. j, k, Contralateral control (j) and denervated (k) incisor teeth 10 days after inferior alveolar nerve transection surgery and 11 days after initiation of genetic tracing by single tamoxifen injection (PLP-CreERT2/R26YFP animals). Arrows in j point at abundant clusters of YFP+ odontoblasts and pulp cells. White dotted line indicates enamel organ and hard matrix. l–m, Wallerian degeneration within the distal stump of the inferior alveolar nerve after the surgery, an area outlined by red rectangle in k. n, Control nerve. o, Quantification of YFP+ progeny in denervated and contralateral control teeth 10 days after surgery (paired t-test P < 0.0001, mean difference 86.6; 95% confidence interval 83.58–89.63; n = 5). p, Quantification of distances between a cervical loop and most distal YFP+ progeny in control (1,479 ± 246.5 µm) and denervated (282.4 ± 71.52 µm) teeth 10 days after the surgery (unpaired t-test P = 0.0016, n = 5). a–i, Scale bars, 100 µm (a, c, e, g); 25 µm (b, d, f, h, i); 100 µm (j, k); 50 µm (l–n). CL1 and CL2 are the labial and lingual aspects of the cervical loop respectively.
Extended Data Figure 9 Schwann-cell-derived MSCs express stem cell marker Thy1.
a–c, Flow cytometry analysis of cell populations from dissociated incisor pulp cells. Incisors were isolated from two PLP-CreERT2/R26YFP animals that were traced for 5 months after a single tamoxifen injection (5 mg). a, Dot plots of isotypic control and immunostaining of non-fixed dissociated pulp cells on the basis of expression of YFP. A region, P1, has been drawn around the YFP+ cells (also shown in black). b, Dot plots of isotypic control and immunostaining of non-fixed dissociated pulp cells on the basis of Thy1 expression. A region, P2, has been drawn around the Thy1+ cells (shown in blue). c, Dot plots of selected sub-populations on the basis of expression of proliferative marker Ki67 and Thy1, staining on fixed cells. YFP+ cells are shown in black. Right panel shows population of YFP+ cells only. A region, P3, has been drawn around Thy1−/Ki67+ cells. P4 has been drawn around Thy1+/Ki67+ cells and P5 around Thy1+/Ki67− cells. d, d1 , Immunostaining for Thy1 and Tuj1 in an adult incisor. d1 Magnified area outlined by white rectangle in d. e, f, Ki67, Sox10 and Thy1 in dental MSC niche between labial and lingual aspects of a cervical loop. Arrows point at Sox10+ nuclei of Schwann cells. Note that some Schwann cells are Ki67+. Inset in e shows only the Thy1 staining channel from area outlined by white rectangle; inset in f shows only the Ki67 staining channel from the same area. g–h, Incisor from a PLP-CreERT2/R26YFP mouse genetically traced for 8 months. Note Thy1+/YFP+ cells located proximally to the cervical loop CL1 in the low-magnification image (g) and indicated by arrows in magnified images (g1 –h). Magnified area is highlighted by white rectangle in g. i, j, Genetically traced incisor from a Thy1-Cre/R26YFP mouse. Note abundant progeny in the pulp (i, j) and odontoblast layer specifically shown in inset in j. k–m2 , Genetically traced incisor from a Thy1-Cre/R26Confetti mouse. Note streams of YFP+ cells in the pulp (k–l1 ) and a few odontoblasts (m2 ). l1 , m2 , Magnified area outlined by white rectangle in k. Dotted line marks enamel epithelium and mineralized matrix. n–q, BrdU incorporation analysis of fast and slowly cycling cells in the incisor. r, s, EdU+ slowly cycling cells 71 days after the last EdU injection in Thy1-stained and genetically traced incisor (9 months after last tamoxifen injection). t1 , u1 , Magnified area from r and s outlined by white rectangle 1. EdU+/YFP+ cells are marked by arrows. Note numerous EdU+ cells in the Thy1+ zone proximal to the cervical loop (CL1) in s. v2 , w2 , Magnified area from r and s outlined by white rectangle 2. Arrows indicate Thy1+/EdU+ cell. x–z, Flow cytometry analysis of EdU-retaining (21 days after last the injection) and Thy1+ cells from dental pulp. x, Dot plot showing the gating for Thy1-expressing cells. y, Dot plot showing the gating of EdU+ cells among total DAPI+ population. Note that EdU+ cells represent 4.3% of total cell numbers. z, Dot plot of subpopulations on the basis of expression of Thy1 and incorporation of EdU. EdU+/Thy1+ cells are in the upper right square, constituting 1.34% of total population. Scale bars, 100 µm (d, i, k); 50 µm (g, j); 100 µm (n–s); 25 µm (t1 –w2 ). CL1 and CL2 are the labial and lingual aspects of the cervical loop respectively.
Extended Data Figure 10 Populations of pericytes and glia-derived cells do not overlap in the tooth.
a–e, Sox10 and NG2 immunohistochemistry on a section of mandibular incisor from a PLP-CreERT2/R26YFP embryo genetically traced from E8.5 to E17.5. a, Dotted line outlines enamel organ. b, c, and d, e, Magnified areas from a outlined by white rectangles. Arrows in d, e show NG2+/YFP− pericytes. f, g, YFP+/NG2+ pericytes (arrows) in the forebrain of an E8.5 to E17.5 genetically traced PLP-CreERT2/R26YFP embryo. h, Immunohistochemistry with Sox10 and NG2 antibodies on a section of mandibular incisor traced for 30 days in a PLP-CreERT2/R26YFP adult mouse. Note the stream of YFP+/NG2− cells in the pulp of the incisor and numerous YFP−/NG2+ pericytes on the same section. a–h, Scale bars, 50 µm (a); 25 µm (b–h). CL1 and CL2 are the labial and lingual aspects of the cervical loop respectively.
Supplementary information
In situ hybridization reveals Thy1 expression in apical region of incisor
Optical projection tomography video showing expression pattern of Thy1 in adult mouse incisor. Note specific localization of Thy1 at the tooth apex. (MOV 5651 kb)
Rights and permissions
About this article
Cite this article
Kaukua, N., Shahidi, M., Konstantinidou, C. et al. Glial origin of mesenchymal stem cells in a tooth model system. Nature 513, 551–554 (2014). https://doi.org/10.1038/nature13536
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13536
This article is cited by
-
Effect of Octamer-Binding Transcription Factor 4 Overexpression on the Neural Induction of Human Dental Pulp Stem Cells
Stem Cell Reviews and Reports (2024)
-
Hallmarks of peripheral nerve function in bone regeneration
Bone Research (2023)
-
C5L2 modulates BDNF production in human dental pulp stem cells via p38α pathway
Scientific Reports (2023)
-
New Insights on the Role of Satellite Glial Cells
Stem Cell Reviews and Reports (2023)
-
Cav1.2 regulated odontogenic differentiation of NG2+ pericytes during pulp injury
Odontology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.