Abstract
In the 1950s, the drug thalidomide, administered as a sedative to pregnant women, led to the birth of thousands of children with multiple defects. Despite the teratogenicity of thalidomide and its derivatives lenalidomide and pomalidomide, these immunomodulatory drugs (IMiDs) recently emerged as effective treatments for multiple myeloma and 5q-deletion-associated dysplasia. IMiDs target the E3 ubiquitin ligase CUL4–RBX1–DDB1–CRBN (known as CRL4CRBN) and promote the ubiquitination of the IKAROS family transcription factors IKZF1 and IKZF3 by CRL4CRBN. Here we present crystal structures of the DDB1–CRBN complex bound to thalidomide, lenalidomide and pomalidomide. The structure establishes that CRBN is a substrate receptor within CRL4CRBN and enantioselectively binds IMiDs. Using an unbiased screen, we identified the homeobox transcription factor MEIS2 as an endogenous substrate of CRL4CRBN. Our studies suggest that IMiDs block endogenous substrates (MEIS2) from binding to CRL4CRBN while the ligase complex is recruiting IKZF1 or IKZF3 for degradation. This dual activity implies that small molecules can modulate an E3 ubiquitin ligase and thereby upregulate or downregulate the ubiquitination of proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Protein Data Bank
Data deposits
Structural coordinates for human DDB1–G. gallus CRBN–thalidomide, human DDB1–G. gallus CRBN–lenalidomide and human DDB1–G. gallus CRBN–pomalidomide have been deposited in the Protein Data Bank under accession numbers 4CI1, 4CI2 and 4CI3. Human protein microarray data sets for this study have been deposited in the Gene Expression Omnibus database under accession number GSE57554.
References
Bartlett, J. B., Dredge, K. & Dalgleish, A. G. The evolution of thalidomide and its IMiD derivatives as anticancer agents. Nature Rev. Cancer 4, 314–322 (2004)
Shortt, J., Hsu, A. K. & Johnstone, R. W. Thalidomide-analogue biology: immunological, molecular and epigenetic targets in cancer therapy. Oncogene 32, 4191–4202 (2013)
Melchert, M. & List, A. The thalidomide saga. Int. J. Biochem. Cell Biol. 39, 1489–1499 (2007)
McBride, W. G. Thalidomide and congenital abnormalities. Lancet 278, 1358 (1961)
Lenz, W., Pfeiffer, R. A., Kosenow, W. & Hayman, D. J. Thalidomide and congenital abnormalities. Lancet 279, 45–46 (1962)
Sheskin, J. Thalidomide in the treatment of lepra reactions. Clin. Pharmacol. Ther. 6, 303–306 (1965)
D'Amato, R. J., Loughnan, M. S., Flynn, E. & Folkman, J. Thalidomide is an inhibitor of angiogenesis. Proc. Natl Acad. Sci. USA 91, 4082–4085 (1994)
Pan, B. & Lentzsch, S. The application and biology of immunomodulatory drugs (IMiDs) in cancer. Pharmacol. Ther. 136, 56–68 (2012)
Singhal, S. et al. Antitumor activity of thalidomide in refractory multiple myeloma. N. Engl. J. Med. 341, 1565–1571 (1999)
Ito, T. et al. Identification of a primary target of thalidomide teratogenicity. Science 327, 1345–1350 (2010)
Higgins, J. J., Pucilowska, J., Lombardi, R. Q. & Rooney, J. P. A mutation in a novel ATP-dependent Lon protease gene in a kindred with mild mental retardation. Neurology 63, 1927–1931 (2004)
Lopez-Girona, A. et al. Cereblon is a direct protein target for immunomodulatory and antiproliferative activities of lenalidomide and pomalidomide. Leukemia 26, 2326–2335 (2012)
Zhu, Y. X. et al. Cereblon expression is required for the antimyeloma activity of lenalidomide and pomalidomide. Blood 118, 4771–4779 (2011)
Lu, G. et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science 343, 305–309 (2014)
Kronke, J. et al. Lenalidomide causes selective degradation of IKZF1 and IKZF3 in multiple myeloma cells. Science 343, 301–305 (2014)
Gandhi, A. K. et al. Immunomodulatory agents lenalidomide and pomalidomide co-stimulate T cells by inducing degradation of T cell repressors Ikaros and Aiolos via modulation of the E3 ubiquitin ligase complex CRL4(CRBN.). Br. J. Haematol. 164, 811–821 (2014)
Li, T., Chen, X., Garbutt, K. C., Zhou, P. & Zheng, N. Structure of DDB1 in complex with a Paramyxovirus V protein: viral hijack of a propeller cluster in ubiquitin ligase. Cell 124, 105–117 (2006)
Scrima, A. et al. Structural basis of UV DNA-damage recognition by the DDB1–DDB2 complex. Cell 135, 1213–1223 (2008)
Li, T., Robert, E. I., van Breugel, P. C., Strubin, M. & Zheng, N. A promiscuous alpha-helical motif anchors viral hijackers and substrate receptors to the CUL4–DDB1 ubiquitin ligase machinery. Nature Struct. Mol. Biol. 17, 105–111 (2010)
Scrima, A. et al. Detecting UV-lesions in the genome: The modular CRL4 ubiquitin ligase does it best!. FEBS Lett. 585, 2818–2825 (2011)
Hur, S., Stroud, R. M. & Finer-Moore, J. Substrate recognition by RNA 5-methyluridine methyltransferases and pseudouridine synthases: a structural perspective. J. Biol. Chem. 281, 38969–38973 (2006)
Ruchelman, A. L. et al. Isosteric analogs of lenalidomide and pomalidomide: Synthesis and biological activity. Bioorg. Med. Chem. Lett. 23, 360–365 (2013)
Jin, J., Arias, E., Chen, J., Harper, J. & Walter, J. A family of diverse Cul4-Ddb1-interacting proteins includes Cdt2, which is required for S phase destruction of the replication factor Cdt1. Mol. Cell 23, 709–721 (2006)
Angers, S. et al. Molecular architecture and assembly of the DDB1–CUL4A ubiquitin ligase machinery. Nature 443, 590–593 (2006)
Fischer, E. S. et al. The molecular basis of CRL4DDB2/CSA ubiquitin ligase architecture, targeting, and activation. Cell 147, 1024–1039 (2011)
Hornbeck, P. V. et al. PhosphoSitePlus: a comprehensive resource for investigating the structure and function of experimentally determined post-translational modifications in man and mouse. Nucleic Acids Res. 40, D261–D270 (2012)
Hofmeister, C. C. et al. Phase I trial of lenalidomide and CCI-779 in patients with relapsed multiple myeloma: evidence for lenalidomide-CCI-779 interaction via P-glycoprotein. J. Clin. Oncol. 29, 3427–3434 (2011)
Zhou, S. et al. Transport of thalidomide by the human intestinal caco-2 monolayers. Eur. J. Drug Metab. Pharmacokinet. 30, 49–61 (2005)
Roche, S. et al. Development, validation and application of a sensitive LC-MS/MS method for the quantification of thalidomide in human serum, cells and cell culture medium. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 902, 16–26 (2012)
Crowley, M. A. et al. Further evidence for the possible role of MEIS2 in the development of cleft palate and cardiac septum. Am. J. Med. Genet. A. 152A, 1326–1327 (2010)
Capdevila, J., Tsukui, T., Rodríquez Esteban, C., Zappavigna, V. & Izpisúa Belmonte, J. C. Control of vertebrate limb outgrowth by the proximal factor Meis2 and distal antagonism of BMPs by Gremlin. Mol. Cell 4, 839–849 (1999)
Paige, S. L. et al. A temporal chromatin signature in human embryonic stem cells identifies regulators of cardiac development. Cell 151, 221–232 (2012)
Skaar, J. R., Pagan, J. K. & Pagano, M. Mechanisms and function of substrate recruitment by F-box proteins. Nature Rev. Mol. Cell Biol. 14, 369–381 (2013)
Bennett, E. J., Rush, J., Gygi, S. P. & Harper, J. W. Dynamics of Cullin-RING Ubiquitin Ligase Network Revealed by Systematic Quantitative Proteomics. Cell 143, 951–965 (2010)
Tan, X. et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645 (2007)
Huai, Q. et al. Crystal structure of calcineurin-cyclophilin-cyclosporin shows common but distinct recognition of immunophilin-drug complexes. Proc. Natl Acad. Sci. USA 99, 12037–12042 (2002)
Kissinger, C. R. et al. Crystal structures of human calcineurin and the human FKBP12–FK506-calcineurin complex. Nature 378, 641–644 (1995)
Landau, M. et al. ConSurf 2005: the projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 33, W299–W302 (2005)
Li, T., Robert, E. I., van Breugel, P. C., Strubin, M. & Zheng, N. A promiscuous α-helical motif anchors viral hijackers and substrate receptors to the CUL4–DDB1 ubiquitin ligase machinery. Nature Struct. Mol. Biol. 17, 105–111 (2010)
Xu, G., Jiang, X. & Jaffrey, S. R. A mental retardation-linked nonsense mutation in cereblon is rescued by proteasome inhibition. J. Biol. Chem. 288, 29573–29585 (2013)
Zhang, L., Lubin, A., Chen, H., Sun, Z. & Gong, F. The deubiquitinating protein USP24 interacts with DDB2 and regulates DDB2 stability. Cell Cycle 11, 4378–4384 (2012)
Zhang, X. et al. Mutations in UVSSA cause UV-sensitive syndrome and destabilize ERCC6 in transcription-coupled DNA repair. Nature Genet. 44, 593–597 (2012)
Forbes, S. A. S. et al. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 39, D945–D950 (2011)
Acknowledgements
This work was supported by the Novartis Research Foundation and grants to N.H.T. from the European Research Council (ERC-2010-StG 260481-MoBa-CS) and to J.W.H. from the National Institutes of Health (AG011085). J.R.L. was supported by a Damon Runyon Postdoctoral Fellowship (DRG 2061-10). We acknowledge J. Reilly for performing immobilized artificial membrane (IAM) experiments. We thank J. Tallarico, J. Porter, W. Sellers, S. Cottens and M. Renatus for help and comments. D. Hess, R. Sack and J. Seebacher for the mass spectrometry analysis and J. Keusch and H. Gut for support. We thank W. Kaelin for kindly providing the IKZF1 reporter plasmid (pCMV–IRES–Renilla luciferase–IRES–Gateway–firefly luciferase). Part of this work was performed at the Swiss Light Source, Paul Scherrer Institute, Villigen, Switzerland.
Author information
Authors and Affiliations
Contributions
E.S.F., N.H.T., J.L.J. and W.C.F. initiated the project. E.S.F. and K.B. conducted the protein purification and crystallization. G.M.L. provided recombinant CSN, and S.C. pre-screened protein complexes by electron microscopy. E.S.F. collected data and processed and refined X-ray data. E.S.F. and N.H.T. analysed the structures. E.S.F. performed in vitro experiments and, with the help of U.H., developed and performed TR-FRET and fluorescence polarization assays. E.S.F. performed protein array experiments. M.B.S. and E.S.F. analysed the data. E.S.F., K.B., J.R.L., H.Y., M.H., J.W.H. and N.H.T. conceived and performed the cell-biological characterization. R.B.T. and R.E.J.B. conceived and conducted the chemical syntheses. J.N. and M. Schirle performed proteomics. V.A. and J.O. carried out the differential scanning fluorimetry experiments. F.S. and M. Schebesta carried out the zebrafish experiments. E.S.F. and N.H.T. wrote the manuscript. All authors assisted in editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Sequence conservation of CRBN.
a, Alignment of G. gallus CRBN and human CRBN; the Zn2+-coordinating domain and residues involved in compound binding are highlighted in yellow and cyan, respectively. The secondary structure of the CRBN protein is shown at the top. b, Surface representation of CRBN orthologues highlighting the evolutionarily conserved residues (sequences used for the alignment include Homo sapiens, Nomascus leucogenys, Macaca mulatta, Gorilla gorilla, Ailuropoda melanoleuca, Callithrix jacchus, Felis catus, Canis lupus, Equus caballus, Mustela putorius furo, Geospiza fortis, Loxodonta africana, Heterocephalus glaber, Pongo abelii, Cavia porcellus, Ovis aries, Cricetulus griseus, Mus musculus, Rattus norvegicus, Desmodus rotundus, Bos taurus, Monodelphis domestica, Anolis carolinensis, G. gallus, Taeniopygia guttata, Meleagris gallopavo, Xenopus tropicalis, Oreochromis niloticus, Takifugu rubripes and Danio rerio). Residues that cluster around the compound-binding pocket are highly conserved38.
Extended Data Figure 2 Structure of the DDB1–CRBN complex bound to thalidomide and its derivatives.
a, Close-up view and topology of the G. gallus CRBN N-terminal domain (NTD). b, Overlay of the G. gallus CRBN-NTD (dark blue) and helical bundle domain (HBD) (light blue) with the Lon protease domain (yellow; PDB 3LJC; r.m.s.d., 2.7 Å over 178 residues aligned). How Lon proteases recognize their substrates is unclear at present. As both the NTD and the HBD are present in the Lon fold, it is conceivable that CRBN originated from a fusion of a Lon protease with a PUA-fold-containing enzyme (see below). In the course of divergent evolution, the helical element of the Lon fold appeared to be used for DDB1 binding. c, The G. gallus CRBN C-terminal domain (CTD; residues 318–445) harbours the thalidomide, lenalidomide or pomalidomide-binding pocket and displays structural similarity to PUA folds. Alignment with the methionine sulphoxide reductase family, the closest structural orthologue of G. gallus CRBN (PDB 3MAO; r.m.s.d., 2.0 Å over 79 residues) is shown. The PUA domain fold family frequently carries a conserved metal coordination site, which in G. gallus CRBN appears to bind Zn2+ and is coordinated by Cys 325, 328, 393 and 396. d, Residues in the active site of the methionine sulphoxide reductase MSRB2 (Trp 103, His 148, Phe 150, Arg 160, Cys 162 and Asn 164). The catalytic cysteine (Cys 162) residue is not conserved in G. gallus CRBN, which makes it unlikely that G. gallus CRBN acts as an MSRB2-like reductase. Interestingly, however, residues equivalent to Trp 402 and Phe 404 involved in G. gallus CRBN thalidomide binding are also involved in substrate binding and catalysis in MSRB2. The conserved binding patch surrounding lenalidomide may thus act as a substrate-binding interface, with thalidomide possibly blocking the binding of a hitherto unknown protein substrate. e, The G. gallus CRBN interaction with DDB1 is mediated through the HBD motif using G. gallus CRBN α-helices H5 (224–233) and H6 (244–249) and the 310 helix H4 (200–204). G. gallus CRBN helices H5 and H6, together with the intervening loop segment, form extensive interactions with the DDB1-BPC propeller, with additional contributions from residues 190–195 of G. gallus CRBN. G. gallus CRBN H4 and the preceding loop (197–205) form interactions with DDB1-BPA. The binding of G. gallus CRBN to BPA occurs through an H4 310 helix located at the central cavity of the WD40 propeller, on the narrow side of the WD40 propeller cone, where ligands typically bind. f, G. gallus CRBN binding to DDB1 differs from that previously seen for DCAF–DDB1 complexes, for which the majority of interactions outside the WD40 propeller domain originated from a consecutive helix–loop–helix (HLH) motif that predominantly contacts DDB1-BPC (golden-brown surface). Extensive interactions between a DCAF and the DDB1-BPA domain (red surface) have not been observed previously, in either the DDB1 co-crystal structures with DDB2 (PDB ID 3EI3; green) or CSA (PDB ID 4A11; dark blue) or other HLH motifs to which DDB1 binds. The novel DDB1-binding mode found in CRBN precluded prior sequence-based identification and suggests considerable plasticity of DCAF binding to DDB1.
Extended Data Figure 3 Thalidomide derivatives used in this study.
a–c, Skeletal formulae of thalidomide (a), lenalidomide (b) and pomalidomide (c). d–i, A thalidomide derivative containing a flexible pendant amine linker at the C4 position (4-OCHC(O)N(CH2)4NH2) of the phthalimide ring system was synthesized (d) as described in Supplementary Methods to derive Cy5-labelled thalidomide (e). Thalidomide analogues were synthesized, including the substitution of the C4 amino group with CH3 (f) or Cl (g), the C5 position with CH3 (h), or the C4 and C6 positions with CH3 (i).
Extended Data Figure 4 Detailed analysis of CRBN–compound interactions.
a, Thermal denaturation assays of thalidomide binding to human DDB1–wild-type human CRBN (CRBN wt) and to a human DDB1–human CRBN mutant harbouring the mutations Tyr384Ala and Trp386Ala (CRBNYW/AA). While the mutant showed no sign of binding to thalidomide, wild-type CRBN caused a shift in the thermal denaturation curve that was indicative of compound binding. b, c, As in a, but using lenalidomide (b) or pomalidomide (c). d, Increasing amounts of human DDB1–human CRBN were mixed with 20 nM Cy5-coupled thalidomide (compound 5). Protein–compound interactions were quantified using fluorescence polarization of the Cy5 dye. Curve fitting to a model assuming one binding site resulted in a dissociation constant (Kd) of 121.6 ± 23.2 nM. The data shown are the mean ± s.d. (n = 3). e–g, Competitive titration of thalidomide with human DDB1–human CRBN (50 nM) and Cy5-thalidomide (20 nM); half-maximum effective concentration (EC50) values were used to calculate a Ki inhibition constant (equivalent to the Kd of thalidomide for human DDB1–human CRBN) of 249.20 nM. Similar titrations were carried out for lenalidomide (f) and pomalidomide (g), resulting in Ki values of 177.80 nM and 156.60 nM, respectively. For all competitive titrations, the data shown are the mean ± s.d. (n = 3). h, Affinities of thalidomide analogues for DDB1–CRBN.
Extended Data Figure 5 Details of CRBN as a substrate receptor in the ubiquitin ligase complex CRL4CRBN.
a, Architecture of CRL4DCAF complexes: DDB1–DDB2 (PDB ID 3EI2), DDB1–CSA (PDB ID 4A11) and DDB1–CRBN are shown side by side. b, Overlay of the structure of DDB1–CRBN, with CRBN shown in cyan, and that of DDB1–DDB2, with DDB2 shown in green. The DDB2 footprint on DDB1 fully overlaps with that of CRBN, rendering binding to DDB1 mutually exclusive. The mutual exclusivity of DDB2 and CRBN binding has been noted previously10. As other WD40-DCAFs attach to DDB1 through similar HLH helical-binding motifs39 (see also Extended Data Fig. 2f), we expected CRBN to sterically exclude most, if not all, DCAFs from binding to DDB1. c, The PUA fold (CRBN-CTD in green) aligns with that of other PUA complexes, such as the MSS4–Rab8 complex (PDB ID 2FU5; r.m.s.d., 3.47 Å over 107 residues). The small helix from Rab8 occupies a binding grove on the PUA domain similar to thalidomide, lenalidomide and pomalidomide binding. d, As in c, but with the PUA surface coloured according to conservation within the CRBN orthologues (see Extended Data Fig. 1). The lenalidomide-binding groove on which Rab8 impinges is conserved. e, PUA-domain-containing proteins are also involved in RNA binding, as shown by the superposition of CRBN and RIG-I (PDB ID 3OG8). While thalidomide somewhat resembles a nucleotide (with the phthaloyl group chemically related to a purine), we note that the binding mode observed for the compound is probably not compatible with RNA or DNA binding to CRBN, owing to severe steric clashes. However, the PUA domain itself appears adaptable to a broad range of ligands. It is not clear at present whether CRBN recognizes a distinct post-translational modification on a protein that somewhat resembles thalidomide.
Extended Data Figure 6 Mutations in CRBN are found in mental retardation and cancer.
a, Mutations in CRBN that are associated with mental retardation (indicated in red) and found in cancer (shown as sticks in magenta). CRBN was first identified as a gene that was mutated in a mild form of mental retardation11. The identified nonsense mutation causes a premature stop codon after Arg 419, resulting in a truncated form of CRBN. When tested in cells, the truncated CRBN bound to DDB1 but exhibited a higher rate of autoubiquitination40. b, In analogy, we found that a recombinant G. gallus CRBN construct (1–426) apparently remained properly folded in vitro, with no obvious association of heat-shock factors. This construct was also capable of forming complexes with DDB1. Two structural rationales that are not mutually exclusive can be proposed for the detrimental effect of the truncation seen in non-syndromic mental retardation. First, the conserved C terminus may be involved in substrate binding; in addition, the absence of the C terminus could also give rise to temporary unfolding that impinges on the ubiquitination zone and thus renders CRBN subject to CRL4CRBN ubiquitination. Second, the removal of the CRBN C-terminal tail could also delete the binding site for a potential de-ubiquitination enzyme that frequently appears bound at the tails of CRL4 substrate receptors, thus counteracting receptor autoubiquitination; DDB2 is complexed to USP24 (ref. 41) and CSA in conjunction with USP7 (ref. 42). c, The Catalogue of Somatic Mutations in Cancer (COSMIC) database43 reports a number of somatic mutations in the CRBN gene in various tumours: one nonsense mutation (p.Glu 106*) and seven missense substitutions have been reported. The structure now provides a molecular rationale for the reported mutations. The premature stop codon after Glu 106 probably leads to an unfolded protein or at least abolishes the interaction with DDB1, resulting in a phenotype similar to a total loss of CRBN expression. Trp224Leu and Asn236Thr are surface-exposed residues involved in DDB1 binding, probably weakening this interaction. The Thr119Ala and Asp249Asn mutations affect the core of the protein and probably impair correct folding. The mutations Glu132Lys, Leu168Pro and Thr403Met are surface mutations and could be involved in substrate binding.
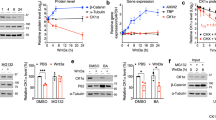
Extended Data Figure 7 Biochemical characterization of thalidomide derivatives and their effect on CRL4CRBN autoubiquitination.
a, In vitro autoubiquitination assays were performed with neddylated CRL4CRBN, E1 (UBA1), E2 (UBCH5A) and ubiquitin. CRBN ubiquitination was detected by anti-CRBN immunoblotting. The use of ubiquitin resulted in a high molecular weight smear, similar to that observed for CRBN autoubiquitination in cells10. b, This is in contrast to the use of K0 ubiquitin, which gave rise to a defined banding pattern. The amount of non-ubiquitinated CRBN remained largely constant, as judged by anti-CRBN immunoblotting, with no gross overall inhibition of autoubiquitination observed following addition of thalidomide (Thal) (lane 3), lenalidomide (Len) (lane 4) or pomalidomide (Pom) (lane 5). Autoubiquitination was inhibited by the addition of CSN (lane 6). c, Autoubiquitination of CRBN within the ligase CRL4CRBN was monitored in the presence of ATP, E1 (UBA1) and E2 (UBCH5A) (lane 2). We found that CRBN autoubiquitination was repressed in a dose-dependent manner in the presence of CSN (lanes 3–5). d, Mass-spectrometry-based identification of ubiquitin modification on CRBN using K0 ubiquitin. One phosphorylation site (Ser 25), as well as three ubiquitination sites, were detected in recombinant CRBN. The location of the ubiquitin sites on the G. gallus CRBN structure is indicated: Lys 39 and Lys 43 are located on the unstructured CRBN N-terminal tail, which is not ordered in the structure and is indicated as a dashed line. The Lys-412 site is also indicated in magenta and is found within 10–15 Å of the presumed radius of gyration of the disordered residues Lys 39/Lys 43. e, Immobilized artificial membrane chromatography experiments predicted similar cell permeability properties for the compounds used in this study.
Extended Data Figure 8 Details of protein array CRL4CRBN ligase substrate profiling.
a, Assessment of intra-array reproducibility: comparison of the fluorescence intensities of the two replicate spots for each protein for an array probed with E1, E2, CRL4CRBN and lenalidomide. b, Assessment of between-array reproducibility: a pairwise correlation plot showing Pearson correlation coefficients between spot intensities for all pairs of arrays, reordered according to similarity. c, A heatmap of cluster averages (the mean across all genes in a cluster). Cluster 42 contains MEIS2. d, A heatmap of individual genes in cluster 42, which contains MEIS2. e, A heatmap of individual genes of interest, including MEIS2 and the IKAROS transcription factors IKZF1 and IKZF3. f, The top four candidate genes were transiently overexpressed in HEK 293T cells and assessed for stabilization on lenalidomide treatment (40 µM for 4 h).
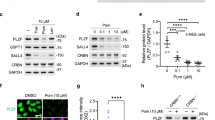
Extended Data Figure 9 Thalidomide and its derivatives modulate the cellular abundance of MEIS2.
a, MEIS2 ubiquitination by CRL4CRBN in vitro was inhibited by thalidomide in a dose-dependent manner. b, A CRL4CRBN complex harbouring a CRBN variant carrying Tyr384Ala and Trp386Ala substitutions was found severely impaired in its ability to ubiquitinate MEIS2 in vitro. c, CRL4CRBN autoubiquitination (lane 2) was inhibited by CSN (lane 3). The inhibition of CRBN autoubiquitination was partially overcome by the addition of MEIS2 (lane 4). d, E3 ligase specificity control showing that MEIS2 is ubiquitinated by CRL4CRBN (lane 1) but not by CRL4CDT2 (lane 2) or CRL4CSA (lane 3). The specificity was further improved in the presence of CSN: MEIS2 ubiquitination by CRL4CRBN did not change (compare lanes 1 and 4), while MEIS2 ubiquitination by CRL4CDT2 (lane 5) and CRL4CSA (lane 6) was further suppressed. e, All immunoblotting was carried out in a quantitative fashion with infrared detection on a LI-COR Odyssey reader. A representative example of a linearity control included on every blot is shown in the figure. f, g, Overexpressed DDK (residues DYKDDDDK)-tagged MEIS2 was stabilized on treatment of HEK 293T cells with 40 µM lenalidomide (f) but not on treatment of HEK 293T cells that stably overexpressed a CRBNYW/AA mutant protein (g). We found that overexpressing wild-type CRBN but not the CRBNYW/AA mutant protein resulted in a severe growth defect in HEK 293T cells (data not shown). h, Similar to what was observed with lenalidomide (Fig. 4b), MLN4924 was found to stabilize MEIS2 protein levels in a CHX chase experiment. i, Endogenous MEIS2 levels in M059J cells (DMSO control, lane 1). Anti-ERK2 immunoblotting was used as a loading control. MEIS2 levels higher than in the DMSO control were seen following the addition of 1–30 µM lenalidomide (lanes 2–4), 5 µM bortezomib or 5 µM MLN4924 following a 4 h incubation. j, The average increase in the MEIS2 protein level following treatment with the indicated amounts of lenalidomide for 4 h. The data are shown as mean ± s.e.m. (n = 3); *, P < 0.05; **, P < 0.01 (two-tailed, unpaired Student’s t-test). k, Thalidomide and pomalidomide led to an increase in MEIS2 protein levels similar to that observed for lenalidomide. An anti-H4 immunoblot was used as a loading control. l, Zebrafish embryos were treated with the indicated amounts of thalidomide 2 h post fertilization and subjected to immunoblotting analysis after 22 h incubation. An up to 1.5-fold increase in MEIS2 protein levels was observed in whole embryo lysate. The data are shown as mean ± s.e.m. (n = 3); *, P < 0.05 (two-tailed, unpaired Student’s t-test). m, Lenalidomide treatment did not influence the accumulation of MEIS2 mRNA. HEK 293T cells treated with the indicated amounts of lenalidomide for 12 h were subjected to quantitative RT–PCR to assess the levels of MEIS2 mRNA. The mRNA levels were normalized to those of GAPDH mRNA. The MEIS2 mRNA level remained stable or even decreased, which does not explain the increase in MEIS2 protein levels observed. n, SDS–PAGE analysis of CRL4CRBN (lane 1) and neddylated CRL4CRBN (lane 2). o, p, Immunoblots with anti-MEIS2 (Abnova) (o) and anti-CRBN (Novus Biologicals) (p) antibodies for endogenous proteins in SK-N-DZ cells displayed no significant nonspecific signal.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, a Supplementary Discussion and Supplementary References. (PDF 714 kb)
Rights and permissions
About this article
Cite this article
Fischer, E., Böhm, K., Lydeard, J. et al. Structure of the DDB1–CRBN E3 ubiquitin ligase in complex with thalidomide. Nature 512, 49–53 (2014). https://doi.org/10.1038/nature13527
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13527
This article is cited by
-
MYC induces CDK4/6 inhibitors resistance by promoting pRB1 degradation
Nature Communications (2024)
-
DNA-encoded library-enabled discovery of proximity-inducing small molecules
Nature Chemical Biology (2024)
-
An orthogonalized PYR1-based CID module with reprogrammable ligand-binding specificity
Nature Chemical Biology (2024)
-
Possible New Candidates Involved to Thalidomide-Related Limbs and Cardiac Defects: A Systems Biology Approach
Biochemical Genetics (2024)
-
The CB1 receptor interacts with cereblon and drives cereblon deficiency-associated memory shortfalls
EMBO Molecular Medicine (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.