Abstract
Inhibitors against the p110δ isoform of phosphoinositide-3-OH kinase (PI(3)K) have shown remarkable therapeutic efficacy in some human leukaemias1,2. As p110δ is primarily expressed in leukocytes3, drugs against p110δ have not been considered for the treatment of solid tumours4. Here we report that p110δ inactivation in mice protects against a broad range of cancers, including non-haematological solid tumours. We demonstrate that p110δ inactivation in regulatory T cells unleashes CD8+ cytotoxic T cells and induces tumour regression. Thus, p110δ inhibitors can break tumour-induced immune tolerance and should be considered for wider use in oncology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Furman, R. R. et al. Idelalisib and rituximab in relapsed chronic lymphocytic leukemia. N. Engl. J. Med. 370, 997–1007 (2014)
Gopal, A. K. et al. PI3Kδ inhibition by idelalisib in patients with relapsed indolent lymphoma. N. Engl. J. Med. 370, 1008–1018 (2014)
Vanhaesebroeck, B. et al. P110δ, a novel phosphoinositide 3-kinase in leukocytes. Proc. Natl Acad. Sci. USA 94, 4330–4335 (1997)
Fruman, D. A. & Rommel, C. PI3K and cancer: lessons, challenges and opportunities. Nature Rev. Drug Discov. 13, 140–156 (2014)
Okkenhaug, K. et al. Impaired B and T cell antigen receptor signaling in p110Δ PI 3-kinase mutant mice. Science 297, 1031–1034 (2002)
Okkenhaug, K. Signaling by the phosphoinositide 3-kinase family in immune cells. Annu. Rev. Immunol. 31, 675–704 (2013)
Curiel, T. J. Regulatory T cells and treatment of cancer. Curr. Opin. Immunol. 20, 241–246 (2008)
Patton, D. T. et al. Cutting edge: the phosphoinositide 3-kinase p110δ is critical for the function of CD4+CD25+Foxp3+ regulatory T cells. J. Immunol. 177, 6598–6602 (2006)
Patton, D. T., Wilson, M. D., Rowan, W. C., Soond, D. R. & Okkenhaug, K. The PI3K p110δ regulates expression of CD38 on regulatory T cells. PLoS ONE 6, e17359 (2011)
Muranski, P. & Restifo, N. P. Adoptive immunotherapy of cancer using CD4+ T cells. Curr. Opin. Immunol. 21, 200–208 (2009)
Soond, D. R. et al. Pten loss in CD4 T cells enhances their helper function but does not lead to autoimmunity or lymphoma. J. Immunol. 188, 5935–5943 (2012)
Okkenhaug, K. et al. The p110δ isoform of phosphoinositide 3-kinase controls clonal expansion and differentiation of Th cells. J. Immunol. 177, 5122–5128 (2006)
Soond, D. R. et al. PI3K p110δ regulates T-cell cytokine production during primary and secondary immune responses in mice and humans. Blood 115, 2203–2213 (2010)
Pulaski, B. A., Smyth, M. J. & Ostrand-Rosenberg, S. Interferon-gamma-dependent phagocytic cells are a critical component of innate immunity against metastatic mammary carcinoma. Cancer Res. 62, 4406–4412 (2002)
Gabrilovich, D. I., Ostrand-Rosenberg, S. & Bronte, V. Coordinated regulation of myeloid cells by tumours. Nature Rev. Immunol. 12, 253–268 (2012)
Bunt, S. K., Sinha, P., Clements, V. K., Leips, J. & Ostrand-Rosenberg, S. Inflammation induces myeloid-derived suppressor cells that facilitate tumor progression. J. Immunol. 176, 284–290 (2006)
Youn, J. I., Nagaraj, S., Collazo, M. & Gabrilovich, D. I. Subsets of myeloid-derived suppressor cells in tumor-bearing mice. J. Immunol. 181, 5791–5802 (2008)
Putz, E. M. et al. PI3Kδ is essential for tumor clearance mediated by cytotoxic T lymphocytes. PLoS ONE 7, e40852 (2012)
Gattinoni, L., Klebanoff, C. A. & Restifo, N. P. Paths to stemness: building the ultimate antitumour T cell. Nature Rev. Cancer 12, 671–684 (2012)
Schmid, M. C. et al. Receptor tyrosine kinases and TLR/IL1Rs unexpectedly activate myeloid cell PI3Kγ, a single convergent point promoting tumor inflammation and progression. Cancer Cell 19, 715–727 (2011)
Soler, A. et al. Inhibition of the p110α isoform of PI 3-kinase stimulates nonfunctional tumor angiogenesis. J. Exp. Med. 210, 1937–1945 (2013)
Murray, J. M. et al. Potent and highly selective benzimidazole inhibitors of PI3-kinase delta. J. Med. Chem. 55, 7686–7695 (2012)
Safina, B. S. et al. Discovery of novel PI3-kinase δ specific inhibitors for the treatment of rheumatoid arthritis: taming CYP3A4 time-dependent inhibition. J. Med. Chem. 55, 5887–5900 (2012)
Acknowledgements
This research was supported by Cancer Research UK (C23338/A10200; C23338/A15965 to B.V. and C18270/A12888 to T. Hagemann), Biotechnology and Biological Sciences Research Council (BB/E009867/1 to K.O.) and Wellcome Trust (095691/Z/11/Z) (to K.O.). We thank D. Sutherlin, J. Nonomiya, J. Lesnick and K. Reif (Genentech) for technical assistance, M. Whitehead, D. Dubuisson, D. Patton, E. Slack, N. Harrington, G. Rosignoli, W. P. Day, A. Brown, P. Depledge, F. Leenders, J. Pang, L. Salphati and X. Zhang for experimental help, D. Fearon and J. Arnold for LLC-OVA cells, and A. Rudensky for FoxP3YFP-Cre mice.
Author information
Authors and Affiliations
Contributions
K.A., D.R.S., R.P., E.L.L, H.B. and W.P. performed experiments and data analyses with input from K.O. and B.V. D.R.S. and K.O. generated the FoxP3YFP-Cre×δflox/flox mice and helped to design, perform and interpret experiments. R.P. and W.P. helped design experiments. T. Hagemann and his team carried out the KPC experiments and performed analysis with help of K.A. M.T. generated the p110δflox/flox mice. T. Hancox performed chemistry for small molecule inhibitor development. H.M. helped to design and performed in vivo pharmacological cancer experiments. L.F. helped design and interpret pharmacological data. C.L.S. performed and interpreted histopathology. K.A., K.O. and B.V. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
K.O. is consultant to GSK (Stevenage) and B.V. is a consultant to Retroscreen (London, UK) and Karus Therapeutics (Oxford, UK).
Extended data figures and tables
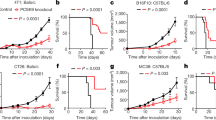
Extended Data Figure 1 Impact of p110δ inactivation on CD4 and CD8 T cells in mice with 4T1 or EL4 tumours.
a, Levels of CD44highCD4+ and CD44highCD8+ T cells in the indicated immune compartments of naive and 4T1 tumour-bearing on day 26 after inoculation in wild-type or δD910A mice. b, Distribution of cells on day 5 of culture of splenocytes, isolated from 4T1 tumour-bearing wild-type and δD910A mice 21 days after inoculation, in the presence of mitomycin-treated 4T1 cells. c, Gene expression in CTLs derived from splenocytes from wild-type and δD910A OT-I mice, cultured in the presence of SIINFEKL OVA peptide and IL-2. GzmA, granzyme A; GzmB, granzyme B, Prf1, perforin and (FasL or CD95L) Fas ligand. Expression levels are presented relative to β2-microglobulin. *P < 0.05, **P < 0.01, ***P < 0.001 (non-parametric Mann–Whitney t-test). Numbers in brackets indicate the number of mice used per experiment. Each dot represents an individual mouse.
Extended Data Figure 2 Impact of p110δ inactivation on myeloid cells in 4T1 tumours.
a, Gating strategy used to identify myeloid cell subsets. Splenic cells were gated on CD11bhigh cells followed by Ly6C and Ly6G gating. FSC, forward scatter; SSC, side scatter (top). Frequency of CD11b+ cells in the spleen of wild-type and δD910A naïve mice and in 4T1 tumour-bearing mice on day 21 after inoculation (bottom). b, [3H]-Thymidine incorporation in co-cultures of splenocytes and purified myeloid cells, in combinations as indicated, with or without stimulation with anti-CD3 antibodies. Cultures were made using cells derived from individual mice. Error bars represent standard deviation from the mean of biological replicates. *P < 0.05, **P < 0.01 (non-parametric Mann–Whitney t-test). Numbers in brackets indicate the number of mice used per experiment. Each dot represents an individual mouse.
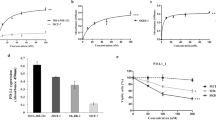
Extended Data Figure 3 Characterization of the p110δ-selective inhibitor PI-3065.
a, PI-3065 structure and in vitro IC50 on selected PI3K family members. No significant activity against 72 protein kinases was observed at ≤ 10 μM in a KinaseProfiler assay (Millipore). b, Pharmacokinetic parameters of PI-3065. Mean (± s.d.) plasma concentration profile of PI-3065 following a single oral dose (75 mg kg−1) administred per os (po) to female BALB/c mice. AUCinf, area under the curve, extrapolated to infinity; Cmax, highest observed plasma concentration; tmax, time at which Cmax occurred, QD, quaque die (every day). c, Growth of primary 4T1 tumours, inoculated in the breast fat pad, measured by calipers and expressed as tumour volume. Mice were dosed per os with vehicle or PI-3065 (75 mg kg−1, daily) for 36 days. 105 tumour cells were inoculated 12 h post first dosing. d, Percentage of tumour-free mice upon continuous per os treatment of mice with vehicle or PI-3065 (25 mg kg−1, twice daily) for 37 days, with tumour cells inoculated on day 7 of PI-3065 dosing. 15 mice were used for each genotype. e, Class I PI3K isoform expression in 4T1 cells. f, Proliferation of 4T1 cells following a 4-h treatment with the indicated p110δ inhibitors, washing and (3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxy phenyl)-2-(4-sulphophenyl)-2H-tetrazolium salt (MTS) staining after 48 h culture. g, Percentage body weight change (from day 0) of 4T1 tumour-bearing mice upon daily per os administration of PI-3065 (75 mg kg−1) or vehicle for 36 consecutive days. *P < 0.05, **P < 0.01 (non-parametric Mann–Whitney t test). Numbers in brackets indicate the number of mice used per experiment.
Supplementary information
Supplementary information
This file contains Supplementary Methods and additional references. (PDF 328 kb)
Rights and permissions
About this article
Cite this article
Ali, K., Soond, D., Piñeiro, R. et al. Inactivation of PI(3)K p110δ breaks regulatory T-cell-mediated immune tolerance to cancer. Nature 510, 407–411 (2014). https://doi.org/10.1038/nature13444
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13444
This article is cited by
-
Regulatory cells and the effect of cancer immunotherapy
Molecular Cancer (2023)
-
Circulating Th17 T cells at treatment onset predict autoimmune toxicity of PI3Kδ inhibitors
Blood Cancer Journal (2023)
-
Metabolic profiles of regulatory T cells and their adaptations to the tumor microenvironment: implications for antitumor immunity
Journal of Hematology & Oncology (2022)
-
Immunosuppressive cells in cancer: mechanisms and potential therapeutic targets
Journal of Hematology & Oncology (2022)
-
Signaling pathways and targeted therapies in lung squamous cell carcinoma: mechanisms and clinical trials
Signal Transduction and Targeted Therapy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.