Abstract
Spatial patterning is a ubiquitous feature of biological systems. Meiotic crossovers provide an interesting example, defined by the classic phenomenon of crossover interference. Here we identify a molecular pathway for interference by analysing crossover patterns in budding yeast. Topoisomerase II plays a central role, thus identifying a new function for this critical molecule. SUMOylation (of topoisomerase II and axis component Red1) and ubiquitin-mediated removal of SUMOylated proteins are also required. The findings support the hypothesis that crossover interference involves accumulation, relief and redistribution of mechanical stress along the protein/DNA meshwork of meiotic chromosome axes, with topoisomerase II required to adjust spatial relationships among DNA segments.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kleckner, N., Zhang, L., Weiner, B. & Zickler, D. in Genome Organization and Function in the Cell Nucleus (ed. Rippe, K. ) 487–533 (John Wiley, 2011)
Zickler, D. & Kleckner, N. The leptotene-zygotene transition of meiosis. Annu. Rev. Genet. 32, 619–697 (1998)
Jones, G. H. & Franklin, F. C. Meiotic crossing-over: obligation and interference. Cell 126, 246–248 (2006)
Kleckner, N. et al. A mechanical basis for chromosome function. Proc. Natl Acad. Sci. USA 101, 12592–12597 (2004)
Muller, H. J. The mechanism of crossing over, parts I–IV. Am. Nat. 50, 193–434 (1916)
Sturtevant, A. H. The behavior of the chromosomes as studied through linkage. Z. indukt. Abstamm.-u. VererbLehre 13, 234–287 (1915)
Hillers, K. J. & Villeneuve, A. M. Chromosome-wide control of meiotic crossing over in C. elegans. Curr. Biol. 13, 1641–1647 (2003)
Zhang, L., Liang, Z., Hutchinson, J. & Kleckner, N. Crossover patterning by the Beam-Film model: analysis and implications. PLoS Genet. 10, e1004042 (2014)
King, J. S. & Mortimer, R. K. A polymerization model of chiasma interference and corresponding computer simulation. Genetics 126, 1127–1138 (1990)
Vecchiarelli, A. G., Hwang, L. C. & Mizuuchi, K. Cell-free study of F plasmid partition provides evidence for cargo transport by a diffusion-ratchet mechanism. Proc. Natl Acad. Sci. USA 110, E1390–E1397 (2013)
Blat, Y., Protacio, R. U., Hunter, N. & Kleckner, N. Physical and functional interactions among basic chromosome organizational features govern early steps of meiotic chiasma formation. Cell 111, 791–802 (2002)
Pan, J. et al. A hierarchical combination of factors shapes the genome-wide topography of yeast meiotic recombination initiation. Cell 144, 719–731 (2011)
Storlazzi, A. et al. Recombination proteins mediate meiotic spatial chromosome organization and pairing. Cell 141, 94–106 (2010)
Borner, G. V., Kleckner, N. & Hunter, N. Crossover/noncrossover differentiation, synaptonemal complex formation, and regulatory surveillance at the leptotene/zygotene transition of meiosis. Cell 117, 29–45 (2004)
Hunter, N. in Molecular Genetics of Recombination, Topics in Current Genetics (eds Aguilera, A. and Rothstein, R. ) 381–442 (Springer, 2006)
Henderson, K. A. & Keeney, S. Synaptonemal complex formation: where does it start? Bioessays 27, 995–998 (2005)
Bishop, D. K. & Zickler, D. Early decision; meiotic crossover interference prior to stable strand exchange and synapsis. Cell 117, 9–15 (2004)
Fung, J. C., Rockmill, B., Odell, M. & Roeder, G. S. Imposition of crossover interference through the nonrandom distribution of synapsis initiation complexes. Cell 116, 795–802 (2004)
Agarwal, S. & Roeder, G. S. Zip3 provides a link between recombination enzymes and synaptonemal complex proteins. Cell 102, 245–255 (2000)
Cheng, C. H. et al. SUMO modifications control assembly of synaptonemal complex and polycomplex in meiosis of Saccharomyces cerevisiae. Genes Dev. 20, 2067–2081 (2006)
Malkova, A. et al. Gene conversion and crossing over along the 405-kb left arm of Saccharomyces cerevisiae chromosome VII. Genetics 168, 49–63 (2004)
Bachant, J., Alcasabas, A., Blat, Y., Kleckner, N. & Elledge, S. J. The SUMO-1 isopeptidase Smt4 is linked to centromeric cohesion through SUMO-1 modification of DNA topoisomerase II. Mol. Cell 9, 1169–1182 (2002)
Rose, D. & Holm, C. Meiosis-specific arrest revealed in DNA topoisomerase II mutants. Mol. Cell. Biol. 13, 3445–3455 (1993)
Martini, E., Diaz, R. L., Hunter, N. & Keeney, S. Crossover homeostasis in yeast meiosis. Cell 126, 285–295 (2006)
Baldwin, M. & Bachant, J. Top2 SUMO conjugation in yeast cell lysates. Methods Mol. Biol. 582, 209–219 (2009)
Eichinger, C. S. & Jentsch, S. Synaptonemal complex formation and meiotic checkpoint signaling are linked to the lateral element protein Red1. Proc. Natl Acad. Sci. USA 107, 11370–11375 (2010)
Hooker, G. W. & Roeder, G. S. A Role for SUMO in meiotic chromosome synapsis. Curr. Biol. 16, 1238–1243 (2006)
Nagai, S., Davoodi, N. & Gasser, S. M. Nuclear organization in genome stability: SUMO connections. Cell Res. 21, 474–485 (2011)
Darst, R. P., Garcia, S. N., Koch, M. R. & Pillus, L. Slx5 promotes transcriptional silencing and is required for robust growth in the absence of Sir2. Mol. Cell. Biol. 28, 1361–1372 (2008)
Drouaud, J. et al. Sex-specific crossover distributions and variations in interference level along Arabidopsis thaliana chromosome 4. PLoS Genet. 3, e106 (2007)
Petkov, P. M., Broman, K. W., Szatkiewicz, J. P. & Paigen, K. Crossover interference underlies sex differences in recombination rates. Trends Genet. 23, 539–542 (2007)
Hou, Y. et al. Genome analyses of single human oocytes. Cell 155, 1492–1506 (2013)
Kleckner, N. Chiasma formation: chromatin/axis interplay and the role(s) of the synaptonemal complex. Chromosoma 115, 175–194 (2006)
Novak, I. et al. Cohesin Smc1beta determines meiotic chromatin axis loop organization. J. Cell Biol. 180, 83–90 (2008)
Klein, F. et al. Localization of RAP1 and topoisomerase II in nuclei and meiotic chromosomes of yeast. J. Cell Biol. 117, 935–948 (1992)
Moens, P. B. & Earnshaw, W. C. Anti-topoisomerase II recognizes meiotic chromosome cores. Chromosoma 98, 317–322 (1989)
Kleckner, N., Zickler, D. & Witz, G. Chromosome capture brings it all together. Science 342, 940–941 (2013)
Agostinho, M. et al. Conjugation of human topoisomerase 2α with small ubiquitin-like modifiers 2/3 in response to topoisomerase inhibitors: cell cycle stage and chromosome domain specificity. Cancer Res. 68, 2409–2418 (2008)
Lee, M. T. & Bachant, J. SUMO modification of DNA topoisomerase II: trying to get a CENse of it all. DNA Repair 8, 557–568 (2009)
Kawamura, R. et al. Mitotic chromosomes are constrained by topoisomerase II-sensitive DNA entanglements. J. Cell Biol. 188, 653–663 (2010)
Pope, L. H., Xiong, C. & Marko, J. F. Proteolysis of mitotic chromosomes induces gradual and anisotropic decondensation correlated with a reduction of elastic modulus and structural sensitivity to rarely cutting restriction enzymes. Mol. Biol. Cell 17, 104–113 (2006)
Libuda, D. E., Uzawa, S., Meyer, B. J. & Villeneuve, A. M. Meiotic chromosome structures constrain and respond to designation of crossover sites. Nature 502, 703–706 (2013)
Warsi, T. H. in Centromeric Functions and Dynamics of DNA Topoisomerase II in S. cerevisiae 130–187. Ph.D. thesis, Univ. California Riverside. (2009)
Chen, S. Y. et al. Global analysis of the meiotic crossover landscape. Dev. Cell 15, 401–415 (2008)
Mancera, E., Bourgon, R., Brozzi, A., Huber, W. & Steinmetz, L. M. High-resolution mapping of meiotic crossovers and non-crossovers in yeast. Cell 454, 479–485 (2008)
Cherry, J. M. et al. Genetic and physical maps of Saccharomyces cerevisiae. Nature 387, 67–73 (1997)
Argueso, J. L., Wanat, J., Gemici, Z. & Alani, E. Competing crossover pathways act during meiosis in Saccharomyces cerevisiae. Genetics 168, 1805–1816 (2004)
de los Santos, T. et al. The Mus81/Mms4 endonuclease acts independently of double-Holliday junction resolution to promote a distinct subset of crossovers during meiosis in budding yeast. Genetics 164, 81–94 (2003)
Hollingsworth, N. M., Ponte, L. & Halsey, C. MSH5, a novel MutS homolog, facilitates meiotic reciprocal recombination between homologs in Saccharomyces cerevisiae but not mismatch repair. Genes Dev. 9, 1728–1739 (1995)
Chua, P. R. & Roeder, G. S. Zip2, a meiosis-specific protein required for the initiation of chromosome synapsis. Cell 93, 349–359 (1998)
Shinohara, M., Oh, S. D., Hunter, N. & Shinohara, A. Crossover assurance and crossover interference are distinctly regulated by the ZMM proteins during yeast meiosis. Nature Genet. 40, 299–309 (2008)
Jessop, L., Rockmill, B., Roeder, G. S. & Lichten, M. Meiotic chromosome synapsis-promoting proteins antagonize the anti-crossover activity of Sgs1. PLoS Genet. 2, e155 (2006)
Henderson, K. A. & Keeney, S. Tying synaptonemal complex initiation to the formation and programmed repair of DNA double-strand breaks. Proc. Natl Acad. Sci. USA 101, 4519–4524 (2004)
Sym, M. & Roeder, G. S. Crossover interference is abolished in the absence of a synaptonemal complex protein. Cell 79, 283–292 (1994)
Nishant, K. T. et al. The baker’s yeast diploid genome is remarkably stable in vegetative growth and meiosis. PLoS Genet. 6, e1001109 (2010)
Kauppi, L. et al. Numerical constraints and feedback control of double-strand breaks in mouse meiosis. Genes Dev. 27, 873–886 (2013)
Serrentino, M. E., Chaplais, E., Sommermeyer, V. & Borde, V. Differential association of the conserved SUMO ligase Zip3 with meiotic double-strand break sites reveals regional variations in the outcome of meiotic recombination. PLoS Genet. 9, e1003416 (2013)
Kim, K. P. et al. Sister cohesion and structural axis components mediate homolog bias of meiotic recombination. Cell 143, 924–937 (2010)
Koszul, R. & Kleckner, N. Dynamic chromosome movements during meiosis: a way to eliminate unwanted connections? Trends Cell Biol. 19, 716–724 (2009)
Loidl, J., Klein, F. & Engebrecht, J. Genetic and morphological approaches for the analysis of meiotic chromosomes in yeast. Methods Cell Biol. 53, 257–285 (1998)
Charles, D. R. The spatial distribution of cross-overs in X-chromosome tetrads of Drosophila melanogaster. J. Genet. 36, 103–126 (1938)
Zhang, L., Kim, K. P., Kleckner, N. E. & Storlazzi, A. Meiotic double-strand breaks occur once per pair of (sister) chromatids and, via Mec1/ATR and Tel1/ATM, once per quartet of chromatids. Proc. Natl Acad. Sci. USA 108, 20036–20041 (2011)
Imai, S., Armstrong, C. M., Kaeberlein, M. & Guarente, L. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature 403, 795–800 (2000)
Wu, C. S., Chen, Y. F. & Gartenberg, M. R. Targeted sister chromatid cohesion by Sir2. PLoS Genet. 7, e1002000 (2011)
Dhillon, N. & Kamakaka, R. T. A histone variant, Htz1p, and a Sir1p-like protein, Esc2p, mediate silencing at HMR. Mol. Cell 6, 769–780 (2000)
Derbyshire, M. K., Weinstock, K. G. & Strathern, J. N. HST1, a new member of the SIR2 family of genes. Yeast 12, 631–640 (1996)
Hong, S. et al. The logic and mechanism of homologous recombination partner choice. Mol. Cell 51, 440–453 (2013)
Acknowledgements
We thank M. Hochstrasser, J. Bachant, S. Jentsch, L. Pillus and M. Weinreich for plasmids, J. Fung for Zip2 focus data, D. Zickler for the image in Fig. 6a, and members of the Kleckner laboratory and D. Zickler for advice and discussions. This research, L.Z., S.W., S.Y. and N.K. were supported by a grant to N.K. from the National Institutes of Health (RO1 GM044794); S.H. and K.P.K. were supported by the National Research Foundation of Korea funded by the Ministry of Science, ICT and Future Planning (2012-M3A9C6050367).
Author information
Authors and Affiliations
Contributions
L.Z. and N.K. conceived and designed experiments, analysed data and wrote the paper. L.Z., S.W., Y.S., S.H. and K.P.K. performed experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
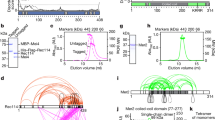
Extended Data Figure 1 Top2 protein level and localization on chromosomes in three top2 mutants.
a, Top2 protein levels shown as a function of time after entry into meiosis (t = 0). Top2 levels are severely reduced in pCLB2-TOP2 (middle panel) and are the same as WT in pCLB2-TOP2 top2YF (±20% relative to anti-Pgk1 control). Western blot analysis used anti-Top2 antibody (TopoGEN 2014) and anti-Pgk1 antibody (Abcam ab113687). b, Immunostaining of Top2 on meiotic chromosomes with the same antibody used for western blot analysis in a: at pachytene (shown) and at leptotene (data not shown). Top2 is undetectable on chromosomes in pCLB2-TOP2 and is present at similar levels to WT in pCLB2- and top2SNM. Chromosomes were concomitantly immunostained for Zip1 (Santa Cruz, sc-48716) as in text Fig. 1. Scale bars, 3 μm.
Extended Data Figure 2 Decreased crossover interference in pCLB2-TOP2 and sir2Δ, slx5Δ is confirmed on other chromosomes.
a, b, The same decreases in crossover interference (LCoC ≈ 0.2 µm versus ≈ 0.3 µm in WT) and corresponding increases crossover number observed for the indicated mutants on chromosome XV (Figs 2 and 3) are also observed on chromosomes IV and III in pCLB2-TOP2 and sir2Δ, and on chromosome IV in slx5Δ. Data for WT in black.
Extended Data Figure 3 Decreased crossover interference as revealed by modified coefficient of coincidence and tetrad analysis using the method of ref. 21, but synaptonemal complex length is the same as in WT.
a, By modified coefficient of coincidence analysis (Fig. 1; Methods), crossover interference can extend to about two intervals on either side of the reference interval (LMCoC ≈ 0.3 µm) in WT and in three sir2 mutants that exhibit WT crossover patterning by other criteria (LCoC ≈ 0.3 µm; Fig. 3 and Extended Data Fig. 2). In contrast, in all examined single and double mutants where crossover interference is defective (LCoC ≈ 0.2 µm; Figs 2, 3, 4), crossover interference extends only about 1.3 intervals (LMCoC ≈ 0.2 µm) (for top2 mutants, see also Fig. 2). Right column shows synaptonemal complex lengths for each of the analysed strains (average ± s.d.). There is no significant difference between strains exhibiting WT interference (average of averages is 3.25 ± 0.06 µm) and strains defective in the top2 interference pathway (average of averages is 3.27 ± 0.07 µm). b, Decreased crossover interference in slx5Δ and sir2RK as revealed by tetrad analysis. Each pair of intervals was tested, reciprocally, for the ratio of the map distances in one interval with and without crossovers in the other interval. Each number shows the average of the ratios for the two reciprocal cases. A value less than 1 indicates crossover interference. Solid and dotted lines indicate whether the level of interference is statistically (P < 0.05 by G-test) significant or not, respectively. Genetic crossover interference is greatly decreased in slx5Δ, and sir2RK relative to WT on each of three chromosomes. Tetrad data upon which this analysis is based are given in Supplementary Table 2.
Extended Data Figure 4 Additional aspects of crossover homeostasis analysis.
a, b, Crossover patterns along chromosome XV in TOP2 strains (a, black) and pCLB2-TOP2 strains (b, black) with WT or altered DSB levels as conferred by the indicated spo11/tel1 genotypes (for crossover homeostasis analysis; Fig. 2d and Methods). All experimental data sets were also subjected to beam-film simulation analysis (a and b, red). In all cases (a and b, red), best-fit simulations were obtained by using the same parameters as those that give the best-fit for SPO11 TEL1 meiosis (ref. 8; Fig. 2a) except that number of precursors (given by parameter N) was altered to account for alterations in DSB levels in the different strain backgrounds (LBF = 0.3 µm in TOP2 background versus 0.2 µm in PCLB2-TOP2 background; see Methods and below). For each spo11/tel1 genotype, the best-fit value of (N) is the same in pCLB2-TOP2 as in TOP2, thus confirming that the only change in various pCLB2-TOP2 strains examined is a change in precursor number, with no change in interference. The same results are also seen for beam-film simulations of analogous data for chromosome III (not shown). These results further illustrate the accuracy with which beam-film simulations can describe diverse crossover patterns. c, Comparison of rad50S DSB levels and beam-film-predicted precursor levels (N) for chromosome XV among strains with varying DSB levels due to different SPO11 TEL1 or carrying spo11 and/or tel1 mutant alleles. Top line: number of DSBs genome-wide, relative to WT = 100, as defined by rad50S analysis in TOP2 strains, either SPO11 TEL1 or carrying spo11 and/or tel1 mutant alleles (details in Methods). Middle line: number of DSBs predicted for chromosome XV. Number of DSBs in TOP2 SPO11 TEL1 was defined by several approaches (details in Methods). DSBs per chromosome XV as predicted for spo11/tel1 mutant strains by comparison of rad50S DSB levels with SPO11 TEl1 (top line). Bottom line: number of precursors predicted to be present by beam-film best-fit simulation analysis (given by parameter N, above). Predicted values are the same for TOP2 and pCLB-TOP2 strain series (from simulations in a and b). Note that in strains with lower total DSB levels, rad50S analysis gives lower DSB/precursor levels than beam-film simulations (discussion in Methods). Analogous results are obtained for chromosome III, as follows. (1) The predicted values of N are the same for both TOP2 and pCLB2-TOP2 strain series: N = 9 for tel1Δ, 6 for TEL1 SPO11, 5 for spo11-HA/spo11HA and 3 for spo11-HA/spo11YF. (2) These predicted values of N correspond well to DSB values predicted from rad50S analysis except at the lowest DSB levels: predicted DSBs = 9 for tel1Δ, 6 for TEL1 SPO11, 5 for spo11-HA/spo11HA and 2 for spo11-HA/spo11YF. d, Experimentally determined numbers of Zip3 foci from the analyses of chromosome XV in a and b are plotted as a function of either the number of precursors predicted by beam-film simulation analysis (left) or the number of DSBs predicted by rad50S DSB analysis (right) (values from c). e, Same as d, except that we analysed chromosome III. A slightly better match of experimental data to beam-film simulation predictions is obtained when the x axis metric is the predicted precursor number than when it is rad50S predicted DSB levels, suggesting that beam-film simulations are more accurate than rad50S DSB analysis, which is known to underestimate DSBs in several situations. Note that for each strain and chromosome, Zip3 foci were analysed in 200–300 cells. The average numbers of foci per bivalent ± s.d. as presented in d and e were as follows. TOP2 chromosome XV (d): tel1Δ 5.21 ± 0.93; tel1Δ spo11HA 4.92 ± 1.12; TEL1 SPO11 4.67 ± 1.16; spo11HA/spo11HA 4.11 ± 0.97; spo11HA/spo1DA 4.07 ± 1.07; spo11HA/spo11YF 3.51 ± 0.88. pCLB2-TOP2 chromosome XV (d): tel1Δ 6.46 ± 1.13; TEL1 SPO11 5.96 ± 1.1; spo11HA/spo11HA 5.29 ± 0.99; spo11HA/spo11DA 4.76 ± 0.94; spo11HA/spo11YF 3.71 ± 0.98. TOP2 chromosome III (e): tel1Δ 2.16 ± 0.59; TEL1 SPO11 1.82 ± 0.55; spo11HA/spo11HA 1.7 ± 0.62; spo11HA/spo11YF 1.31 ± 0.66. pCLB2-TOP2 chromosome III (e): tel1Δ 2.49 ± 0.82; TEL1 SPO11 2.1 ± 0.87; spo11HA/spo11HA 2.07 ± 0.75; spo11HA/spo11YF 1.51 ± 0.69.
Extended Data Figure 5 Increased level of SUMO–protein conjugates in slx5Δ.
a, Western blots for whole protein extracts in WT and slx5Δ probed with anti-Smt3 antibody (Santa Cruz, sc-28649) and anti-Pgk1 antibody (Abcam ab113687) as a function of time after entry into meiosis (t = 0). Abundance of SUMO conjugates is increased in the mutant, especially in regions of high molecular mass. b, Quantification of the gel in a.
Extended Data Figure 6 The role of Sir2 in crossover interference is specific to its interaction with Slx5.
WT crossover interference is seen in diverse sir2 non-null mutants affecting specific sub-functions (other than sir2RK; Fig. 3) and in mutants deleted for various interaction partners. sir2-345 is defective in histone deacetylase activity63; sir2ΔC500 lacks a Sir2 cohesion role64. sir3Δ, sir4Δ, esc2Δ and esc8Δ eliminate Sir2 interaction partners involved in silencing43,65; hst1Δ eliminates a Sir2 homologue66.
Extended Data Figure 7 Mutant coefficient of coincidence and crossover number phenotypes cannot be explained by increased DSBs or by prolongation of the crossover-designation stage.
Mutants in the described crossover interference pathway all confer coordinate changes in crossover interference, which is reduced, and the total number of crossovers, which is increased, by about 20% on chromosome XV. There are the expected consequences of a single defect in crossover interference, as illustrated by corresponding beam-film simulations, which quantitatively explain these results by a change in a single parameter, the interference length (LBF) (Figs 2 and 3). This interference defect could comprise a defect in generation and spreading of the inhibitory signal and/or of the ability of unreacted precursors to respond to that signal (see text and Methods (section ‘Beam-film simulations’)). An increase in the number of crossovers can also occur as the result of either (1) prolongation of the crossover-designation period or (2) an increase in the number of DSBs8. Neither of these effects can explain the mutant phenotypes described in the text. (1) Crossover designation precedes synaptonemal complex formation and thus the pachytene stage14. Time-course analysis of representative mutant strains reveals that, in sir2 mutants and in top2SNM, meiosis proceeds through pachytene and the two meiotic divisions normally (Extended Data Fig. 8a; ref. 14; data not shown). slx5/8 mutants and PCLB2-TOP2 mutants show no delay in progressing through prophase to pachytene (data not shown) but show a delay in meiosis I (slx5) or pachytene arrest (PCLB2-TOP2) (Extended Data Fig. 8a; data not shown). The pCLB2-TOP2 top2YF mutant does show a delay in achieving pachytene, as well as pachytene arrest, but exhibits the same crossover patterning phenotype as all other mutants, which show no pre-pachytene delay. Thus, prolonged crossover designation is not the basis for these phenotypes. (2) An increase in DSBs, without any change in crossover interference, does increase the number of crossovers; however, it has very little effect on crossover interference relationships (coefficient of coincidence curves) in budding yeast8. Correspondingly, two lines of evidence show that the mutant defects described here cannot be attributed to an increase in DSBs. a, A tel1Δ mutant exhibits increased DSBs but no change in coefficient of coincidence relationships. TEL1 encodes the yeast homologue of ATM. Absence of Tel1 confers a 50% increase in DSBs62 and a 10% increase in number of Zip3 foci (Supplementary Fig. 7 in ref. 8; reproduced in Extended Data Fig. 7a left, red colour). However, (1) there is no change in coefficient of coincidence relationships relative to WT (Extended Data Fig. 7a left), (2) the increase in crossovers is precisely that predicted on the basis of crossover homeostasis (ref. 8; text Fig. 2d, filled black circle at 19 DSBs/precursors per chromosome XV) and (3) beam-film simulation accurately describes the tel1Δ phenotype, relative to WT, by a change in a single parameter: the level of DSBs (n = 19, grey, versus 13, gold, in WT). The last point is documented in Extended Data Fig. 7a middle and right. The middle panel in Extended Data Fig. 7a shows the beam-film best-fit simulation for WT chromosome XV, where n = 13 (gold), compared with the experimental coefficient of coincidence curve (black; from Fig. 1); the right panel shows the beam-film best-fit simulation for tel1Δ chromosome XV, where n = 19 (grey) and all other parameters are the same as for WT, compared with the experimental coefficient of coincidence curve (red) from the left panel. b, Beam-film simulations predict no/little change in coefficient of coincidence with increasing DSBs for yeast chromosome XV (data not shown). More specifically, to explain the increased number of crossovers observed in the analysed mutants, for example pCLB2-TOP2, the value of N required for beam-film simulations of chromosome XV would be 26 (double the WT value of N = 13). If beam-film simulations are performed under the same parameter values used for WT except that N = 26 instead of N = 13, the predicted coefficient of coincidence curve is unchanged compared with that predicted for WT (left panel, compare gold for N = 13 with green for N = 26). Correspondingly, the coefficient of coincidence curve predicted for N = 26 (green) matches the WT coefficient of coincidence curve (black) and is unlike the coefficient of coincidence curve for the mutant (pink) (right panel). Additional evidence that DSB number is not altered in pCLB2-TOP2 versus TOP2 is presented in Extended Data Figs 4 and 8.
Extended Data Figure 8 Progression of meiosis and of recombination in interference-defective mutants.
Representative mutants were examined for progression of meiotic divisions and for recombination at the previously characterized HIS4LEU2 locus67 (strains in Extended Data Table 1). a, Meiotic divisions. The first meiotic division occurs normally in sir2RK (defective in interaction with Slx5); it is delayed in slx5Δ and is completely absent in PCLB2-TOP2 and PCLB-TOP2 top2YF due to arrest at pachytene23 (L.Z., unpublished observations). b, c, DNA events. The HIS4LEU2 locus probably provides a direct readout of DNA events independent of the effects of interference. HIS4LEU2 does not exhibit crossover homeostasis24, which implies that it is not sensitive to crossover interference8. This feature presumably reflects the fact that this locus is a very strong DSB hot spot. A DSB occurs at this site in virtually every nucleus with a concomitant reduction in DSBs (and thus crossover precursors) at other positions in its vicinity (N.K., unpublished observations). This locus may also undergo early crossover designation, thus also dominating crossover interference patterns per se. Importantly, Zip3 foci are used for diagnosis of crossover interference relationships8. Zip3 foci form as a specific consequence of programmed crossover designation; they do not mark the sites of non-interfering crossovers, which exhibit an entirely different pattern along the chromosomes8. Furthermore, formation of Zip3 foci is upstream of, and thus insensitive to, defects in later events, including (1) major perturbations in the kinetics of recombination or the fidelity with which initiated events (crossover-fated and/or non-crossover-fated) proceed to their assigned fates (see, for example, ref. 14) or (2) the potential occurrence of additional DSBs due to delayed synaptonemal complex formation (discussion in refs 8 and 56). Thus, none of the recombination aberrancies detected by physical analysis of recombination in the analysed mutants (below) is relevant to their crossover interference phenotypes. Correspondingly, although all mutants give exactly the same crossover patterns (interference and crossover number) as defined by Zip3 foci, the mutants vary widely with respect to DNA recombination phenotypes. The results below can be summarized to say that (1) absence of Slx5/8-Sir2 STUbL activity has little, or only subtle, effect(s) on recombination, whereas (2) absence of TopoII or TopoII catalytic activity confers delays and aberrancies. b, DSBs, SEIs and dHJs. Progression through recombination is very similar to WT in sir2RK and slx5Δ. Both PCLB2-TOP2 and PCLB-TOP2 top2YF exhibit a phenotype corresponding to delayed progression beyond the point of crossover designation: DSBs appear on time; however, DSBs, single-end invasions (SEIs) and double Holliday junctions (dHJs) all accumulate to higher than normal levels at later than normal times, implying delayed progression of crossover-designated DSBs to SEIs, and of SEIs to dHJs, where SEIs and dHJs are both crossover-specific intermediates14. There is no significant alteration in homologue-versus-sister bias in any of the four mutants, with inter-homologue dHJs predominating over inter-sister dHJs similarly to WT in all cases. c, Inter-homologue crossover (CO) and non-crossover (NCO) products. Inter-homologue crossover and non-crossover levels are very similar to WT in PCLB2-TOP2 and show variations relative to WT in the other mutants. A differential deficit of crossovers versus non-crossovers in PCLB2-TOP2 top2YF suggests a specific defect in crossover maturation in this mutant.
Extended Data Figure 9 The metric of crossover interference is physical axis length (micrometres).
a, This study considered two different condensin mutants, ycs4S and pCLB2-BRN1. Axis length is normal in ycs4S and longer than normal in pCLB2-BRN1. Analysis presented for chromosome XV in pCLB2-BRN1 (Fig. 5) was also done on chromosome III in that mutant background (right column), confirming that coefficient of coincidence relationships are WT when the metric is physical chromosome length but not when the metric is genomic distance. We similarly analysed chromosomes III and XV in the ycs4S background (left and middle columns), confirming WT coefficient of coincidence relationships by both metrics. b, Zip3 focus analysis for chromosome XV in the indicated strains (red; from Fig. 5) and beam-film simulation analysis (green). Best-fit simulations could be obtained for all strains using the same parameter values as for WT meiosis, including interference distance (LBF ≈ 0.3 µm), except that the number of precursors (N) had to be varied linearly with axis length. For the indicted strains, from left to right, N = 17, 13, 12, 10, 9 and 8. This result implies direct interplay between physical chromosome length (micrometres of synaptonemal complex) and DSB probability, as discussed elsewhere. c, d, For the mutant cases described in b, experimentally observed average numbers of Zip3 foci vary linearly with axis length (c). In contrast, different numbers of Zip3 foci are observed for the different strains despite the fact that chromosome XV has the same genomic length in all cases (d). We also note that the best fit simulation for BR zip1Δ had to include a 10% decrease in the ‘efficiency of maturation of crossover-designated interactions’, which, in the present context, implies that in a zip1Δ background there is a 10% reduction in either (1) the stability of a Zip3 focus under cytological spreading conditions at the absence of synaptonemal complex or (2) the probability that a crossover designation will give a Zip3 focus.
Supplementary information
Supplementary Information
This file contains the Supplementary Discussion and Supplementary Table 2. (PDF 262 kb)
Supplementary Data
This file contains Supplementary Table 1. (XLS 1280 kb)
Rights and permissions
About this article
Cite this article
Zhang, L., Wang, S., Yin, S. et al. Topoisomerase II mediates meiotic crossover interference. Nature 511, 551–556 (2014). https://doi.org/10.1038/nature13442
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13442
This article is cited by
-
Predicting recombination suppression outside chromosomal inversions in Drosophila melanogaster using crossover interference theory
Heredity (2023)
-
Learning to tango with four (or more): the molecular basis of adaptation to polyploid meiosis
Plant Reproduction (2023)
-
Crossover patterning in plants
Plant Reproduction (2023)
-
Yeast polyubiquitin unit regulates synaptonemal complex formation and recombination during meiosis
Journal of Microbiology (2022)
-
Getting there: understanding the chromosomal recruitment of the AAA+ ATPase Pch2/TRIP13 during meiosis
Current Genetics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.