Abstract
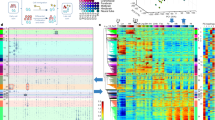
Regulated transcription controls the diversity, developmental pathways and spatial organization of the hundreds of cell types that make up a mammal. Using single-molecule cDNA sequencing, we mapped transcription start sites (TSSs) and their usage in human and mouse primary cells, cell lines and tissues to produce a comprehensive overview of mammalian gene expression across the human body. We find that few genes are truly ‘housekeeping’, whereas many mammalian promoters are composite entities composed of several closely separated TSSs, with independent cell-type-specific expression profiles. TSSs specific to different cell types evolve at different rates, whereas promoters of broadly expressed genes are the most conserved. Promoter-based expression analysis reveals key transcription factors defining cell states and links them to binding-site motifs. The functions of identified novel transcripts can be predicted by coexpression and sample ontology enrichment analyses. The functional annotation of the mammalian genome 5 (FANTOM5) project provides comprehensive expression profiles and functional annotation of mammalian cell-type-specific transcriptomes with wide applications in biomedical research.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Vickaryous, M. K. & Hall, B. K. Human cell type diversity, evolution, development, and classification with special reference to cells derived from the neural crest. Biol. Rev. Camb. Philos. Soc. 81, 425–455 (2006)
Lenhard, B., Sandelin, A. & Carninci, P. Metazoan promoters: emerging characteristics and insights into transcriptional regulation. Nature Rev. Genet. 13, 233–245 (2012)
Kanamori-Katayama, M. et al. Unamplified cap analysis of gene expression on a single-molecule sequencer. Genome Res. 21, 1150–1159 (2011)
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature http://dx.doi.org/10.1038/nature12787 (this issue)
The ENCODE Project Consortium An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012)
Su, A. I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl Acad. Sci. USA 101, 6062–6067 (2004)
Meehan, T. F. et al. Logical development of the cell ontology. BMC Bioinformatics 12, 6 (2011)
Mungall, C. J., Torniai, C., Gkoutos, G. V., Lewis, S. E. & Haendel, M. A. Uberon, an integrative multi-species anatomy ontology. Genome Biol. 13, R5 (2012)
Osborne, J. D. et al. Annotating the human genome with Disease Ontology. BMC Genomics 10 (Suppl 1). S6 (2009)
Severin, J. et al. Interactive visualization and analysis of large-scale NGS data-sets using ZENBU. Nature Biotechnol. http://dx.doi.org/10.1038/nbt.2840 (2014)
Oja, E., Hyvarinen, A. & Karhunen, J. Independent Component Analysis (John Wiley & Sons, 2001)
Affymetrix/Cold Spring Harbor Laboratory ENCODE Transcriptome Project Post-transcriptional processing generates a diversity of 5′-modified long and short RNAs. Nature 457, 1028–1032 (2009)
Carninci, P. et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nature Genet. 38, 626–635 (2006)
Ioshikhes, I., Hosid, S. & Pugh, B. F. Variety of genomic DNA patterns for nucleosome positioning. Genome Res. 21, 1863–1871 (2011)
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010)
Schug, J. et al. Promoter features related to tissue specificity as measured by Shannon entropy. Genome Biol. 6, R33 (2005)
Beissbarth, T. & Speed, T. P. GOstat: find statistically overrepresented Gene Ontologies within a group of genes. Bioinformatics 20, 1464–1465 (2004)
Velculescu, V. E. et al. Analysis of human transcriptomes. Nature Genet. 23, 387–388 (1999)
Schmidt, D. et al. Five-vertebrate ChIP-seq reveals the evolutionary dynamics of transcription factor binding. Science 328, 1036–1040 (2010)
Barolo, S. Shadow enhancers: frequently asked questions about distributed cis-regulatory information and enhancer redundancy. Bioessays 34, 135–141 (2012)
Roach, J. C. et al. Transcription factor expression in lipopolysaccharide-activated peripheral-blood-derived mononuclear cells. Proc. Natl Acad. Sci. USA 104, 16245–16250 (2007)
Vaquerizas, J. M., Kummerfeld, S. K., Teichmann, S. A. & Luscombe, N. M. A census of human transcription factors: function, expression and evolution. Nature Rev. Genet. 10, 252–263 (2009)
Wingender, E., Schoeps, T. & Dönitz, J. TFClass: an expandable hierarchical classification of human transcription factors. Nucleic Acids Res. 41, D165–D170 (2013)
de Kok, Y. J. et al. Association between X-linked mixed deafness and mutations in the POU domain gene POU3F4. Science 267, 685–688 (1995)
Kiernan, A. E. et al. Sox2 is required for sensory organ development in the mammalian inner ear. Nature 434, 1031–1035 (2005)
Zheng, W. et al. The role of Six1 in mammalian auditory system development. Development 130, 3989–4000 (2003)
Paylor, R., Johnson, R. S., Papaioannou, V., Spiegelman, B. M. & Wehner, J. M. Behavioral assessment of c-fos mutant mice. Brain Res. 651, 275–282 (1994)
Trowe, M. O., Maier, H., Schweizer, M. & Kispert, A. Deafness in mice lacking the T-box transcription factor Tbx18 in otic fibrocytes. Development 135, 1725–1734 (2008)
Vahava, O. et al. Mutation in transcription factor POU4F3 associated with inherited progressive hearing loss in humans. Science 279, 1950–1954 (1998)
Chabchoub, E., Willekens, D., Vermeesch, J. R. & Fryns, J. P. Holoprosencephaly and ZIC2 microdeletions: novel clinical and epidemiological specificities delineated. Clin. Genet. 81, 584–589 (2012)
Pingault, V. et al. SOX10 mutations in patients with Waardenburg-Hirschsprung disease. Nature Genet. 18, 171–173 (1998)
Kapoor, S., Mukherjee, S. B., Shroff, D. & Arora, R. Dysmyelination of the cerebral white matter with microdeletion at 6p25. Indian Pediatr. 48, 727–729 (2011)
Murakami, T. et al. Signalling mediated by the endoplasmic reticulum stress transducer OASIS is involved in bone formation. Nature Cell Biol. 11, 1205–1211 (2009)
Acampora, D. et al. Craniofacial, vestibular and bone defects in mice lacking the Distal-less-related gene Dlx5. Development 126, 3795–3809 (1999)
Kieslinger, M. et al. EBF2 regulates osteoblast-dependent differentiation of osteoclasts. Dev. Cell 9, 757–767 (2005)
Funato, N. et al. Hand2 controls osteoblast differentiation in the branchial arch by inhibiting DNA binding of Runx2. Development 136, 615–625 (2009)
McIntyre, D. C. et al. Hox patterning of the vertebrate rib cage. Development 134, 2981–2989 (2007)
Driller, K. et al. Nuclear factor I X deficiency causes brain malformation and severe skeletal defects. Mol. Cell. Biol. 27, 3855–3867 (2007)
Lu, M. F. et al. prx-1 functions cooperatively with another paired-related homeobox gene, prx-2, to maintain cell fates within the craniofacial mesenchyme. Development 126, 495–504 (1999)
Ten Berge, D., Brouwer, A., Korving, J., Martin, J. F. & Meijlink, F. Prx1 and Prx2 in skeletogenesis: roles in the craniofacial region, inner ear and limbs. Development 125, 3831–3842 (1998)
Laclef, C. et al. Altered myogenesis in Six1-deficient mice. Development 130, 2239–2252 (2003)
Lee, M. S., Lowe, G. N., Strong, D. D., Wergedal, J. E. & Glackin, C. A. TWIST, a basic helix-loop-helix transcription factor, can regulate the human osteogenic lineage. J. Cell. Biochem. 75, 566–577 (1999)
Clement-Jones, M. et al. The short stature homeobox gene SHOX is involved in skeletal abnormalities in Turner syndrome. Hum. Mol. Genet. 9, 695–702 (2000)
He, G. et al. Inactivation of Six2 in mouse identifies a novel genetic mechanism controlling development and growth of the cranial base. Dev. Biol. 344, 720–730 (2010)
Freeman, T. C. et al. Construction, visualisation, and clustering of transcription networks from microarray expression data. PLoS Comput. Biol. 3, e206 (2007)
The FANTOM Consortium The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005)
Suzuki, H. et al. The transcriptional network that controls growth arrest and differentiation in a human myeloid leukemia cell line. Nature Genet. 41, 553–562 (2009)
Kawaji, H. et al. Comparison of CAGE and RNA-seq transcriptome profiling using a clonally amplified and single molecule next generation sequencing. Genome Res. http://dx.doi.org/10.1101/gr.156232.113 (2014)
Heffner, C. S. et al. Supporting conditional mouse mutagenesis with a comprehensive cre characterization resource. Nature Commun. 3, 1218 (2012)
Pringle, I. A. et al. Rapid identification of novel functional promoters for gene therapy. J. Mol. Med. 90, 1487–1496 (2012)
Pham, T. H. et al. Dynamic epigenetic enhancer signatures reveal key transcription factors associated with monocytic differentiation states. Blood 119, e161–e171 (2012)
Shulha, H. P. et al. Epigenetic signatures of autism; trimethylated H3K4 landscapes in prefrontal neurons. Arch. Gen. Psychiatry 69, 314–324 (2012)
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nature Immunol. 5, 730–737 (2004)
Shapira, S. D. et al. A physical and regulatory map of host-influenza interactions reveals pathways in H1N1 infection. Cell 139, 1255–1267 (2009)
Talukder, A. H. et al. Phospholipid scramblase 1 regulates Toll-like receptor 9-mediated type I interferon production in plasmacytoid dendritic cells. Cell Res. 22, 1129–1139 (2012)
Acknowledgements
FANTOM5 was made possible by a Research Grant for RIKEN Omics Science Center from MEXT to Y. Hayashizaki and a grant of the Innovative Cell Biology by Innovative Technology (Cell Innovation Program) from the MEXT, Japan to Y. Hayashizaki. It was also supported by Research Grants for RIKEN Preventive Medicine and Diagnosis Innovation Program (RIKEN PMI) to Y. Hayashizaki and RIKEN Centre for Life Science Technologies, Division of Genomic Technologies (RIKEN CLST (DGT)) from the MEXT, Japan. Extended acknowledgements are provided in the Supplementary Information.
Author information
Authors and Affiliations
Consortia
Contributions
The core members of FANTOM5 phase 1 were Alistair R. R. Forrest, Hideya Kawaji, Michael Rehli, J. Kenneth Baillie, Michiel J. L. de Hoon, Timo Lassmann, Masayoshi Itoh, Kim M. Summers, Harukazu Suzuki, Carsten O. Daub, Jun Kawai, Peter Heutink, Winston Hide, Tom C. Freeman, Boris Lenhard, Vladimir B. Bajic, Martin S. Taylor, Vsevolod J. Makeev, Albin Sandelin, David A. Hume, Piero Carninci and Yoshihide Hayashizaki. Samples were provided by: A. Blumenthal, A. Bonetti, A. Mackay-sim, A. Sajantila, A. Saxena, A. Schwegmann, A.G.B., A.J.K., A.L., A.R.R.F., A.S.B.E., B.B., C. Schmidl, C. Schneider, C.A.D., C.A.W., C.K., C.L.M., D.A.H., D.A.O., D.G., D.S., D.V., E.W., F.B., F.N., G.G.S., G.J.F., G.S., H. Kawamoto, H. Koseki, H. Morikawa, H. Motohashi, H. Ohno, H. Sato, H. Satoh, H. Tanaka, H. Tatsukawa, H. Toyoda, H.C.C., H.E., J. Kere, J.B., J.F., J.K.B., J.S.K., J.T., J.W.S., K.E., K.J.H., K.M., K.M.S., L.F., L.M.K., L.M.vdB., L.N.W., M. Edinger, M. Endoh, M. Fagiolini, M. Hamaguchi, M. Hara, M. Herlyn, M. Morimoto, M. Rehli, M. Yamamoto, M. Yoneda, M.B., M.C.F.C., M.D., M.E.F., M.O., M.O.H., M.P., M.vdW., N.M., N.O., N.T., P.A., P.G.Z., P.H., P.R., R.F., R.G., R.K.S., R.P., R.V., S. Guhl, S. Gustincich, S. Kojima, S. Koyasu, S. Krampitz, S. Sakaguchi, S. Savvi, S.E.Z., S.O., S.P.B., S.P.K., S. Roy., S.Z., T. Kitamura, T. Nakamura, T. Nozaki, T. Sugiyama, T.B.G., T.D., T.G., T.I., T.J.H., T.J.K., V.O., W.L., Y. Hasegawa, Y. Nakachi, Y. Nakamura, Y. Yamaguchi, Y. Yonekura, Y.I., Y.I.K., Y.M. and Y.O. Analyses were carried out by: A. Mathelier, A. Meynert, A. Sandelin, A.C., A.D.D., A.P.G., A.H., A.J., A.M.B., A.P., A.R.R.F., A.S.K., A.T.K., A.V.F., B. Lenhard, B. Lilje, B.D., B.K., B.M., B.R.J., C. Schmidl, C. Schneider, C.A.S., C.F., C.J.M., C.O.D., C.P., C.V.C., D.A., D.A.M., D.C., E. Dalla, E. Dimont, E.A., E.A.S., E.J.W., E.M., E.V., Ev.N., F.D., G.J., G.J.F., G.M.A., H. Kawaji, H. Ohmiya, H. Shimoji, H.F., H.J., H.P., I.A., I.E.V., I.H., I.V.K., J.A.B., J.A.C.A., J.A.R., J.C.M., J.F.J.L., J.G., J.G.D.P., J.H., J.K.B., J.S., K. Kajiyama, K.I., K.L., L.H., L.L., M. Francescatto, M. Rashid, M. Rehli, M. Roncador, M. Thompson, M.B.R., M.C., M.C.F., M.J., M.J.L.dH., M.L., M.S.T., M.V., N.B., O.J.L.R., O.M.H., P.A.C.tH., P.J.B, R.A., R.S.Y., S. Katayama, S. Kawaguchi, S. Schmeier, S. Rennie, S.F., S.J.H.S., S.P., T. Sengstag, T.C.F., T.F.M., T.H., T.K., T.L., T.R., T.T., U.S., V.B.B., V.H., V.J.M., W.H., W.W.W., X.Z., Y. Chen, Y. Ciani, Y.A.M., Y.S., Z.T. Libraries were generated by: A. Kaiho, A. Kubosaki, A. Saka, C. Simon, E.S., F.H., H.N., J. Kawai, K. Kaida, K.N., M. Furuno, M. Murata, M. Sakai, M. Tagami, M.I., M.K., M.K.K., N.K., N.N., N.S., P.C., R.M., S. Kato, S.N., S.N.-S., S.W., S.Y., T.A., T. Kawashima. The manuscript was written by A.R.R.F. and D.A.H. with help from A. Sandelin, J.K.B., M. Rehli, H.K., M.J.L.dH., V.H., I.V.K., M.T. and K.M.S. with contributions, edits and comments from all authors. The project was managed by Y. Hayashizaki, A.R.R.F., P.C., M.I., M.S., J. Kawai, C.O.D., H. Suzuki, T.L. and N.K. The scientific coordinator was A.R.R.F and the general organizer was Y. Hayashizaki.
Corresponding authors
Ethics declarations
Competing interests
The author declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Decomposition-based peak identification (DPI).
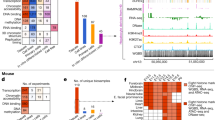
a, Schematic representation of each step in the peak identification. This starts from CAGE profiles at individual biological states (I), subsequently defines tag clusters (consecutive genomic region producing CAGE signals) over the accumulated CAGE profiles across all the states (II). Within each of the tag cluster, it infers up to five underlying signals (independent components) by using ICA independent component analysis (ICA) (III). It smoothens each of the independent components and finds peaks where signal is higher than the median (IV). The peaks along the individual components are finally merged if they are overlapping each other (V). b, c, Genomic view of actual examples (B4GALT1 locus) for human and mouse. CAGE profiles across the biological states (I) are shown as a greyscale plot, in which the x axis represents the genomic coordinates and individual rows represent individual biological states. Dark (or black) dots indicate frequent observation of transcription initiation (that is, larger number of CAGE read counts) and light dots (white) indicate less frequency. The blue histogram on the top indicates the accumulated CAGE read counts, and the entire region shown represents a single tag cluster (II). The histograms below the greyscale plot indicate the independent components of the CAGE signals inferred by ICA (III), and the resulting CAGE peaks are shown at the blue bars closest to the bottom (V). The bottom track indicates a gene model in RefSeq. The figures overall indicate that only one TSS is defined by RefSeq gene models in this locus, however, transcription starts from slightly different regions depending on the context, and the DPI method successfully captured the different initiation events. d, Breakdown of singleton and composite transcription initiation regions with homogenous or heterogeneous expression patterns according to likelihood ratio test (see Supplementary Methods).
Extended Data Figure 2 Broad and sharp promoters.
DPI peaks from the permissive set were aggregated by grouping neighbouring peaks less than 100 bp apart. Cumulative distribution of CAGE signal along each region was calculated and positions of 10th and 90th percentiles were determined. a, Schematic representation of CAGE signal within promoter region and calculation of interquantile width. Signal from CAGE transcription start sites (CTSS) is shown. Distance between these two positions (interquantile width) was used as a measure of promoter width. b, Distribution of promoter interquantile width across all 988 human samples. Individual grey lines show distribution in each sample and the average distribution is shown in yellow. For each sample only promoters with > = 5 TPM were selected. Distribution of obtained interquantile width was clearly bimodal and allowed us to set the empirical threshold at 10.5 bp that separates the best sharp from broad promoters. c, Distribution of expression specificity. The distribution of log ratios of expression in individual samples against the median expression across all samples is shown separately for sharp and broad promoters. Solid line shows the average distribution for all samples and the semi-transparent band denotes the 99% confidence interval. The dashed line corresponds to an expected log ratio if all samples contributed equally to the total expression. d, Average frequency of AA/AT/TA/TT (WW) dinucleotides around dominant TSS of sharp (red) and broad (blue) promoters across all human samples. Lines show the average signal and semi-transparent bands indicate the 99% confidence interval. Closer view of WW dinucleotide frequency displaying 10 bp periodicity is shown in the inset and indicates the likely position of the +1 nucleosome. For comparison, the signal aligned to randomly chosen TSS in broad promoters is shown in orange. e, As in a but for promoters in CD14+ monocytes. H2A.Z signal (subtracted coverage = plus strand coverage – minus strand coverage) around sharp and broad promoters is shown in corresponding semi-transparent colours (data from ref. 51). Transition point in subtracted coverage from positive to negative values indicates the most likely position of the nucleosome (shown as semi-transparent blue circle) centre. f, As in b but for promoters in frontal lobe. H3K4me3 signal (subtracted coverage = plus strand coverage – minus strand coverage) around sharp and broad promoters is shown in corresponding semi-transparent colours (data from ref. 52).
Extended Data Figure 3 Density plots of DPI peaks maximum and median expression.
a, Distribution for all human robust peaks. b, Distribution for all mouse robust peaks. Fraction on left of vertical dashed line corresponds to peaks with non-ubiquitous (cell-type-restricted) expression patterns (median <0.2 TPM). Fraction below the diagonal dashed line corresponds to ubiquitous-uniform (housekeeping) expression profiles (less than tenfold difference between maximum and median). Fraction in top-middle corresponds to ubiquitous-non-uniform expression profiles (maximum >tenfold median). c–e Show distibutions based on cell line, primary cell and tissue data, respectively. The mixture of cells in tissues may overestimate the fraction of ubiquitously expressed genes. f, Boxplot showing the number of peaks and detected > = 10 TPM in primary cells, cell lines or tissues. g, As in a but showing transcription factor p1 peaks only. h, Boxplot showing maximum expression of the main promoter for transcription factors or all coding genes. i, Density plots of human robust DPI peaks maximum and median expression for the main promoter of coding genes. j, As in d but showing the main promoter of transcription factors. Fraction on the left of the vertical dashed line corresponds to peaks with non-ubiquitous (cell-type-restricted) expression patterns (median <0.2 TPM). Fraction below the diagonal dashed line corresponds to ubiquitous-uniform (housekeeping) expression profiles (less than tenfold difference between max and median). Fraction above the diagonal and to the right of the vertical dashed lines corresponds to ubiquitous-non-uniform expression profiles (maximum > tenfold median). k, Distribution for peaks with CpG island only (n = 55,897). l, Distribution for peaks with only a TATA motif (n = 3,933). m, Distribution for peaks with both CpG islands and TATA box motifs (n = 834). n, Distribution for DPI peaks with neither a TATA motif nor CpG island (n = 124,152). Fraction on the left of the vertical dashed line corresponds to peaks with non-ubiquitous (cell-type-restricted) expression patterns (median <0.2 TPM). Fraction below the diagonal dashed line corresponds to ubiquitous-uniform (housekeeping) expression profiles (less than tenfold difference between max and median). Fraction above diagonal and to right of vertical dashed lines corresponds to ubiquitous-non-uniform expression profiles (maximum > tenfold median).
Extended Data Figure 4 Cross-species projected super-clusters.
a, The number of mouse and human TSSs (both permissive and robust) per projected super-cluster. b, Same data as presented in panel a, with the y axis on a log scale. There is a slight tendency for more human TSSs per super-cluster than mouse TSSs. c, The number of human and mouse TSSs per projected super-cluster, density of data points indicated by log-scaled colour gradient shown on the right. Most super-clusters contain < = 4 DPI defined TSSs in both species. d, Evaluating the conservation of TSS annotation between species. Projected super-clusters are annotated by the most functional contributing TSS from each species (see Methods). Grey shading in the margins summarizes the proportion of super-clusters with each category of annotation in both mouse (y axis) and human (x axis). Numbers and volumes of circles represent counts of projected super-clusters, for example there are 34,868 super-clusters in which > = 1 human and > = 1 mouse component TSS are annotated as protein coding and 719 super-clusters in which the human TSSs are unannotated and at least one of the mouse TSSs are annotated as the 5′ end of a non-coding transcript.
Extended Data Figure 5 De novo derived, cell-state-specific motif signatures.
a–c, The de novo motif discovery tools DMF, HOMER, ChIPMunk and ScanAll were applied to detect sequence motifs enriched in the vicinity of sample-specific peaks (a), yielding 8,699 de novo motifs (b). The coverage of known motif space by the de novo motifs was evaluated by comparing them to the SWISSREGULON, HOCOMOCO, TRANSFAC, HOMER, JASPAR, and ENCODE LEXICON motif collections. c, The remaining 1,221 de novo motifs that were not similar to known motifs were then clustered using MACRO-APE, resulting in 169 unique novel motifs. d, Known motifs from the HOMER database were annotated and counted in around cell-type-specific TSSs (−300 to +50 bp) associated with CpG islands (CGI) or non-CGI regions. e–g, RNA Pol II ChIP-seq signal and motif finding in ‘housekeeping gene’ promoters with different absolute expression levels. Human housekeeping gene promoters were defined as (log10(max + 0.1) − log10(median + 0.1) < = 1). The resulting clusters were then extended by −300 and +50. Overlapping extended clusters were removed by only keeping those with the highest expression. e, Extended clusters were then split into 5 equal sized bins with decreasing absolute expression. f, RNA Pol II occupancy at binned clusters in ENCODE cell lines (highly expressed genes show the highest occupancy, but even bin5 clusters showing very low tag counts are still highly occupied). g, Bubble plot representation comparing known motif enrichments in bin1 (high expression) and bin5 (low expression) extended CAGE clusters. The bubble plots encode two quantitative parameters per motif: difference in motif occurrence between bin1 (x axis) and bin5 (y axis) as well as the adjusted P values for enrichment (bubble diameter). Colouring indicates significantly differentially distributed motifs (5% FDR). The right panel additionally summarizes the fraction of clusters in each bin that contain the indicated motifs along with the Benjamini Hochberg adjusted hypergeometric P value for differential enrichment.
Extended Data Figure 6 Features of cell-type-specific promoters.
a, The distribution of expression log ratios of all individual samples against the median of all samples is shown separately for CGI-associated and non-CGI-associated CAGE clusters. The dashed line corresponds to an expected log ratio if all samples contribute equally to the total expression. b, Histograms for genomic distance distributions of HepG2 DNase I hypersensitivity, H3K4me3, H2A.Z, POL2, P300, GABP, YY1, HNF4A, FOXA1 and FOXA2 ChIP-seq tag counts centred across CGI-associated and non-CGI-associated CAGE clusters (separated according to expression specificities) across a 2 kilobase (kb) genomic region. Expression specificity bins are colour-coded (as indicated in the DNase I panel) with blue representing the highest degree of specificity. Numbers of regions in bins are given in the GABP panel (CGI no. / nCGI no., colour coding as above). c, Histograms for genomic distance distributions of ChIP-seq-derived sequence motifs for GABP, YY1, HNF4A, FOXA1 and FOXA2 (corresponding to the samples in the lower panel of c) centred across CGI-associated and non-CGI-associated CAGE clusters (separated according to expression specificities) across a 2 kb genomic region. Motifs are shown on top. The percentage of promoters overlapping with ChIP-seq peaks (b) or consensus sequences (c) for transcription factors binding the highest specificity clusters (HNF4A, FOXA2, TCF7L2) is also given in blue. d, Plots showing mean expression specificity (high values indicate more constrained expression over cells, see the accompanying manuscript4) in enhancers close to RefSeq promoters as a function of promoter CpG content and three classes of promoter expression specificity.
Extended Data Figure 7 Extended features of cell-type-specific promoters.
a, Distribution of global expression specificity estimated using primary cells, cell lines or tissues only. b, Distribution of expression specificity for HepG2, GM12878, HeLaS3, K562 and CD14+monocytes (distribution of expression log ratios of all individual samples against the median of all samples is shown separately for CGI-associated and nonCGI-associated CAGE clusters. The dashed line corresponds to an expected log ratio if all samples contribute equally to the total expression). c, Histograms for genomic distance distributions of K562 DNase I hypersensitivity, H3K4me3, H2A.Z, POL2, P300, GATA1 ChIP-seq tag counts centred across CGI-associated and non-CGI-associated CAGE clusters (separated according to expression specificities) across a 2 kb genomic region. Expression specificity bins are colour-coded with blue representing the highest degree of specificity. d, DNase I hypersensitivity, H3K4me3, H2A.Z, POL2, P300 and IRF4 in GM12878. e, DNase I hypersensitivity, H3K4me3, H2A.Z in HeLaS3. f, DNase I hypersensitivity, H3K4me3, H2A.Z, PU.1 and CEBPB in CD14+ monocytes.
Extended Data Figure 8 Transcription factor promoter expression profile clustering.
a, Biolayout visualization of transcription factor coexpression in human primary cells (3,775 nodes, 54,892 edges r > 0.70, MCL2.2). b, Hierarchical coexpression clustering and heatmap of ETS family transcription factors across the entire human collection (only promoter1(p1) data shown).
Extended Data Figure 9 Collapsed coexpression network for mouse coexpression groups.
One node is one group of promoters. Derived from expression profiles of 116,277 promoters across 402 primary cell types, tissues and cell lines (r > 0.75, MCLi = 2.2). For display, each group of promoters is collapsed into a sphere, the radius of which is proportional to the cube root of the number of promoters in that group. Edges indicate r > 0.6 between the average expression profiles of each cluster. Colours indicate loosely-associated collections of coexpression groups (MCLi = 1.2). Labels show representative descriptions of the dominant cell type in coexpression groups in each region of the network, and a selection of highly-enriched pathways (FDR < 10−4) from KEGG (K), WikiPathways (W), Netpath (N) and Reactome (R).
Extended Data Figure 10 Annotated expression profiles of alternative promoters.
Overlay of coexpression groups enriched for genes involved in the KEGG pathway for influenza A pathogenesis (hsa:05164; FDR < 0.1, n > 2). a, Collapsed coexpression network showing 5 groups enriched for influenza pathogenesis genes: C0 (blue), C26 (purple), C61 (yellow), C187 (green) and C413 (red). b, Excerpt from KEGG pathway diagram showing positions of genes in each coexpression group (background colours as in a). Pathway entities that map to two coexpression groups have the background colour of the smaller group, and the text/border colour of the larger group. Details and promoter-level displays (edges indicate r > 0.75) for two coexpression groups are displayed with transcripts mapping to KEGG pathway highlighted (inset). In this example the KEGG pathway for influenza A pathogenesis (hsa:05164) was strikingly over-represented in one small coexpression group in particular (C413, P value <10−11, FDR = 4.5 × 10−10). Of 19 promoters in coexpression group 413, eight were present in the KEGG pathway, including RIG-I (DDX58), the gene encoding the receptor for the mitochondrial antiviral signalling pathway53. Four of the remaining genes (TRIM21, TRIM22, RTP4 and XAF1) were found to be key host determinants of influenza virus replication in a high-throughput short interfering RNA (siRNA) screen54, whereas another, PLSCR1, is required for a normal interferon response to influenza A55. The top five transcription factor expression profiles most correlated with C413 were IRF7, IRF9, STAT1, SP100 and ZNFX1, and from motif enrichment analysis, the most frequent motifs found in promoters of cluster C413 were potential IRF-binding motifs. c, p1@IRF9 and p2@IRF9 expression ranked by the ubiquitously expressed p1@IRF9 promoter. d, As in a but ranked by expression of p2@IRF9. e, f, Similar to a and b but showing expression of p1@TRMT5 (housekeeping profile) and p2@TRMT5 (expressed in pathogen challenged monocytes). g, Histogram showing the number of different coexpression clusters (see Fig. 4) in which named genes with alternative promoters participate. The majority of genes with alternative promoters participate in more than one cluster; 17 genes participate in more than 10 different clusters and are not shown on this graph.
Extended Data Figure 11 Sample ontology enrichment analysis (SOEA).
Expression profile-sample ontology associations were tested by Mann–Whitney rank sum test to identify cell, disease or anatomical ontology terms over-represented in ranked lists of samples expressing each peak. a, p1@CXCL6 enriched in vascular associated smooth muscle cells. b, p5@ST8SIA3 enriched in brain tissues. c, Novel peak enriched in mast cells. d, p1@KIAA0125 enriched in myeloid leukaemia. e, p1@BRI3 enriched in myeloid leukaemia. f, p1@BDNF enriched in fibroblasts. g, Novel peak enriched in leukocytes. h, Novel peak enriched in classical monocytes. i, j, Venn diagrams showing degree of overlap between peaks associated to known genes (blue), cell ontology enriched (yellow), Uberon anatomical ontology enriched (green) and disease ontology (red). i, At a threshold of 10−20 (Mann–Whitney rank sum test), 64% (59,835 out of 93,558) of the expression profiles of human known transcripts and 74% (67,810 out of 91,269) of the expression profiles for novel transcripts show enrichment for one or more sample ontologies. j, Mouse sample ontology enrichment 10−20 threshold. 30% (18,273 out of 61,134) known are enriched and 47% (26,176 out of 55,143) novel are enriched.
Extended Data Figure 12 Sample-to-sample correlation graph.
821 nodes are shown, 21,821 edges shown (r>0.75). a, Samples are coloured by sample type (primary cell, cell line or tissue). Note the separation of cell lines and primary cells. b, As in a, except major subgroups are coloured and labelled separately.
Supplementary information
Supplementary Information
This file contains Acknowledgements, Supplementary Methods, Supplementary Notes 1-7, Supplementary Figures 1-24, and additional references (see page 1 for more details). Supplementary Tables 1-16 are in a separate excel fie. (PDF 5780 kb)
Supplementary Tables
This file contains Supplementary Tables 1-16.Supplementary Table 2 in the original file was truncated and was replace online on 19 October 2015. (XLSX 19828 kb)
Rights and permissions
About this article
Cite this article
The FANTOM Consortium and the RIKEN PMI and CLST (DGT). A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014). https://doi.org/10.1038/nature13182
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13182
This article is cited by
-
A type 1 immunity-restricted promoter of the IL−33 receptor gene directs antiviral T-cell responses
Nature Immunology (2024)
-
Quantitative analysis of transcription start site selection reveals control by DNA sequence, RNA polymerase II activity and NTP levels
Nature Structural & Molecular Biology (2024)
-
AIRE relies on Z-DNA to flag gene targets for thymic T cell tolerization
Nature (2024)
-
Biological basis of extensive pleiotropy between blood traits and cancer risk
Genome Medicine (2024)
-
Combining a prioritization strategy and functional studies nominates 5’UTR variants underlying inherited retinal disease
Genome Medicine (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.