Abstract
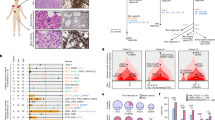
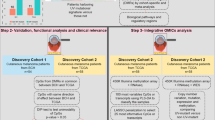
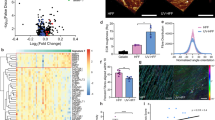
Intermittent intense ultraviolet (UV) exposure represents an important aetiological factor in the development of malignant melanoma1. The ability of UV radiation to cause tumour-initiating DNA mutations in melanocytes is now firmly established2, but how the microenvironmental effects of UV radiation3,4 influence melanoma pathogenesis is not fully understood. Here we report that repetitive UV exposure of primary cutaneous melanomas in a genetically engineered mouse model5 promotes metastatic progression, independent of its tumour-initiating effects. UV irradiation enhanced the expansion of tumour cells along abluminal blood vessel surfaces and increased the number of lung metastases. This effect depended on the recruitment and activation of neutrophils, initiated by the release of high mobility group box 1 (HMGB1) from UV-damaged epidermal keratinocytes and driven by Toll-like receptor 4 (TLR4). The UV-induced neutrophilic inflammatory response stimulated angiogenesis and promoted the ability of melanoma cells to migrate towards endothelial cells and use selective motility cues on their surfaces. Our results not only reveal how UV irradiation of epidermal keratinocytes is sensed by the innate immune system, but also show that the resulting inflammatory response catalyses reciprocal melanoma–endothelial cell interactions leading to perivascular invasion, a phenomenon originally described as angiotropism in human melanomas by histopathologists6. Angiotropism represents a hitherto underappreciated mechanism of metastasis7 that also increases the likelihood of intravasation and haematogenous dissemination. Consistent with our findings, ulcerated primary human melanomas with abundant neutrophils and reactive angiogenesis frequently show angiotropism and a high risk for metastases. Our work indicates that targeting the inflammation-induced phenotypic plasticity of melanoma cells and their association with endothelial cells represent rational strategies to specifically interfere with metastatic progression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chang, Y. M. et al. Sun exposure and melanoma risk at different latitudes: a pooled analysis of 5700 cases and 7216 controls. Int. J. Epidemiol. 38, 814–830 (2009)
Hodis, E. et al. A landscape of driver mutations in melanoma. Cell 150, 251–263 (2012)
Haass, N. K. & Herlyn, M. Normal human melanocyte homeostasis as a paradigm for understanding melanoma. J. Investig. Dermatol. Symp. Proc. 10, 153–163 (2005)
Zaidi, M. R. et al. Interferon-γ links ultraviolet radiation to melanomagenesis in mice. Nature 469, 548–553 (2011)
Gaffal, E. et al. Neonatal UVB exposure accelerates melanoma growth and enhances distant metastases in Hgf-Cdk4R24C C57BL/6 mice. Int. J. Cancer 129, 285–294 (2011)
Barnhill, R. L. & Lugassy, C. Angiotropic malignant melanoma and extravascular migratory metastasis: description of 36 cases with emphasis on a new mechanism of tumour spread. Pathology 36, 485–490 (2004)
Valastyan, S. & Weinberg, R. A. Tumor metastasis: molecular insights and evolving paradigms. Cell 147, 275–292 (2011)
Tronnier, M., Smolle, J. & Wolff, H. H. Ultraviolet irradiation induces acute changes in melanocytic nevi. J. Invest. Dermatol. 104, 475–478 (1995)
Bernard, J. J. et al. Ultraviolet radiation damages self noncoding RNA and is detected by TLR3. Nature Med. 18, 1286–1290 (2012)
Lange, S. S., Mitchell, D. L. & Vasquez, K. M. High mobility group protein B1 enhances DNA repair and chromatin modification after DNA damage. Proc. Natl Acad. Sci. USA 105, 10320–10325 (2008)
Yang, H. et al. A critical cysteine is required for HMGB1 binding to Toll-like receptor 4 and activation of macrophage cytokine release. Proc. Natl Acad. Sci. USA 107, 11942–11947 (2010)
Venereau, E. et al. Mutually exclusive redox forms of HMGB1 promote cell recruitment or proinflammatory cytokine release. J. Exp. Med. 209, 1519–1528 (2012)
Kirschmann, D. A., Seftor, E. A., Hardy, K. M., Seftor, R. E. & Hendrix, M. J. Molecular pathways: vasculogenic mimicry in tumor cells: diagnostic and therapeutic implications. Clin. Cancer Res. 18, 2726–2732 (2012)
Lugassy, C. et al. Pilot study on “pericytic mimicry” and potential embryonic/stem cell properties of angiotropic melanoma cells interacting with the abluminal vascular surface. Cancer Microenviron. 6, 19–29 (2013)
Schwitalla, S. et al. Intestinal tumorigenesis initiated by dedifferentiation and acquisition of stem-cell-like properties. Cell 152, 25–38 (2013)
Landsberg, J. et al. Melanomas resist T-cell therapy through inflammation-induced reversible dedifferentiation. Nature 490, 412–416 (2012)
Nagy, N. et al. Endothelial cells promote migration and proliferation of enteric neural crest cells via β1 integrin signaling. Dev. Biol. 330, 263–272 (2009)
Gupta, P. B. et al. The melanocyte differentiation program predisposes to metastasis after neoplastic transformation. Nature Genet. 37, 1047–1054 (2005)
Lugassy, C., Peault, B., Wadehra, M., Kleinman, H. K. & Barnhill, R. L. Could pericytic mimicry represent another type of melanoma cell plasticity with embryonic properties? Pigment Cell Melanoma Res. 26, 746–754 (2013)
Jensen, T. O. et al. Intratumoral neutrophils and plasmacytoid dendritic cells indicate poor prognosis and are associated with pSTAT3 expression in AJCC stage I/II melanoma. Cancer 118, 2476–2485 (2012)
Schmidt, H. et al. Pretreatment levels of peripheral neutrophils and leukocytes as independent predictors of overall survival in patients with American Joint Committee on Cancer Stage IV Melanoma: results of the EORTC 18951 Biochemotherapy Trial. J. Clin. Oncol. 25, 1562–1569 (2007)
Jablonska, J., Leschner, S., Westphal, K., Lienenklaus, S. & Weiss, S. Neutrophils responsive to endogenous IFN-β regulate tumor angiogenesis and growth in a mouse tumor model. J. Clin. Invest. 120, 1151–1164 (2010)
Eggermont, A. M. et al. Ulceration and stage are predictive of interferon efficacy in melanoma: results of the phase III adjuvant trials EORTC 18952 and EORTC 18991. Eur. J. Cancer 48, 218–225 (2012)
Mittal, D. et al. TLR4-mediated skin carcinogenesis is dependent on immune and radioresistant cells. EMBO J. 29, 2242–2252 (2010)
West, X. Z. et al. Oxidative stress induces angiogenesis by activating TLR2 with novel endogenous ligands. Nature 467, 972–976 (2010)
Kim, S. et al. Carcinoma-produced factors activate myeloid cells through TLR2 to stimulate metastasis. Nature 457, 102–106 (2009)
Hiratsuka, S. et al. Primary tumours modulate innate immune signalling to create pre-metastatic vascular hyperpermeability foci. Nature Commun. 4, 1853 (2013)
Huh, S. J., Liang, S., Sharma, A., Dong, C. & Robertson, G. P. Transiently entrapped circulating tumor cells interact with neutrophils to facilitate lung metastasis development. Cancer Res. 70, 6071–6082 (2010)
Cools-Lartigue, J. et al. Neutrophil extracellular traps sequester circulating tumor cells and promote metastasis. J. Clin. Invest. 123, 3446–3458 (2013)
Guo, Z. S., Liu, Z., Bartlett, D. L., Tang, D. & Lotze, M. T. Life after death: targeting high mobility group box 1 in emergent cancer therapies. Am. J. Cancer Res. 3, 1–20 (2013)
Kohlmeyer, J. et al. Complete regression of advanced primary and metastatic mouse melanomas following combination chemoimmunotherapy. Cancer Res. 69, 6265 (2009)
Vintersten, K. et al. Mouse in red: red fluorescent protein expression in mouse ES cells, embryos, and adult animals. Genesis 40, 241–246 (2004)
Gais, P. et al. Cutting edge: divergent cell-specific functions of MyD88 for inflammatory responses and organ injury in septic peritonitis. J. Immunol. 188, 5833–5837 (2012)
Tarutani, M. et al. Tissue-specific knockout of the mouse Pig-a gene reveals important roles for GPI-anchored proteins in skin development. Proc. Natl Acad. Sci. USA 94, 7400–7405 (1997)
Mollica, L. et al. Glycyrrhizin binds to high-mobility group box 1 protein and inhibits its cytokine activities. Chem. Biol. 14, 431–441 (2007)
Schiraldi, M. et al. HMGB1 promotes recruitment of inflammatory cells to damaged tissues by forming a complex with CXCL12 and signaling via CXCR4. J. Exp. Med. 209, 551–563 (2012)
Barnhill, R., Dy, K. & Lugassy, C. Angiotropism in cutaneous melanoma: a prognostic factor strongly predicting risk for metastasis. J. Invest. Dermatol. 119, 705–706 (2002)
Bloch, W. et al. The angiogenesis inhibitor endostatin impairs blood vessel maturation during wound healing. FASEB J. 14, 2373–2376 (2000)
Gaffal, E. et al. Cannabinoid 1 receptors in keratinocytes modulate proinflammatory chemokine secretion and attenuate contact allergic inflammation. J. Immunol. 190, 4929–4936 (2013)
Pflicke, H. & Sixt, M. Preformed portals facilitate dendritic cell entry into afferent lymphatic vessels. J. Exp. Med. 206, 2925–2935 (2009)
Kastenmüller, W., Torabi-Parizi, P., Subramanian, N., Lammermann, T. & Germain, R. N. A spatially-organized multicellular innate immune response in lymph nodes limits systemic pathogen spread. Cell 150, 1235–1248 (2012)
Nicosia, R. F. The aortic ring model of angiogenesis: a quarter century of search and discovery. J. Cell. Mol. Med. 13, 4113–4136 (2009)
Baker, M. et al. Use of the mouse aortic ring assay to study angiogenesis. Nature Protocols 7, 89–104 (2011)
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004)
Dunning, M. J., Smith, M. L., Ritchie, M. E. & Tavare, S. beadarray: R classes and methods for Illumina bead-based data. Bioinformatics 23, 2183–2184 (2007)
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007)
Huber, W., von Heydebreck A, Sültmann, H., Poustka, A. & Vingron, M. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 18,, (suppl. 1)S96–S104 (2002)
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A. affy–analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004)
Kholmanskikh, O. et al. Interleukins 1α and 1β secreted by some melanoma cell lines strongly reduce expression of MITF-M and melanocyte differentiation antigens. Int. J. Cancer 127, 1625–1636 (2010)
Supek, F., Bosnjak, M., Skunca, N. & Smuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011)
Lin, W. M. et al. Modeling genomic diversity and tumor dependency in malignant melanoma. Cancer Res. 68, 664–673 (2008)
Acknowledgements
We would like to thank the following individuals for their support: G. Merlino, M. Barbacid, A. Nagy, M. Gertsenstein and J. Takeda for generously providing genetically engineered mice; S. Bald, A. Sporleder, C. Lemke, T. Artz and P. Aymans for managing the mouse colony and performing tumour analyses. This research was funded in part by the following grants: Deutsche Krebshilfe P9 in the Melanoma Research Network and DFG A12 in the SFB832 as well as A22 in the SFB704 to T.T., BONFOR to J.L., J.K. and C.H.-H., DFG SFB832 core support and HO 4281/2-1 to M.H., DFG A7 in the SFB832 and A2 in the SFB829 to M.K., DFG A8 in the SFB704 to W. Ko., DFG A1 in the SFB704 to I.F., DFG SFB829 Z2 to W.B., NRW junior research group to D.W., Jürgen Manchot Stiftung to H.W. and AIRC to M.E.B. Additionally, the DFG supports I.F., W.Ka., W.Ko., M.H. and T.T. as members of the Excellence Cluster ImmunoSensation.
Author information
Authors and Affiliations
Contributions
T.B. coordinated and performed all animal experiments and tissue analyses. T.B. and T.Q. developed and performed transwell, 2D cell surface migration and ear tissue invasion assays. M.R. performed keratinocyte culture and melanoma-endothelial cell co-culture in aortic ring assays. J.L., N.G. and D.L.-R. helped with animal experiments and tissue analyses. J.L. and J.K. collected and analysed patient data. R.R. and M.K. performed static adhesion assays. D.W. and B.K.F. helped to design and interpret aortic ring assay experiments. W.Ka. helped with whole mount immunostaining, confocal microscopy and image analysis. W.B. performed transmission electron microscopy. M.R., J.K., S.R., D.v.d.B.-K. and C.H.-H. performed experiments with human cell lines. I.H. and D.S. provided human melanoma cell lines, helped to design experiments and interpret the data. M.E.B. helped to design and interpret experiments involving HMGB1. C.L. and R.B. helped to interpret angiotropism in mice and human histopathology. B.S., H.W. and I.F. helped to design and interpret experiments involving Myd88LSL mice. W.Ko. helped to design and interpret migration assays. M.H. coordinated human melanoma cell culture and performed microarray experiments and bioinformatic analyses. E.G. helped to design and coordinate all animal experiments. T.B., T.Q., M.R and J.L. helped to prepare figures and methods. W.Ko., M.H. and E.G. interpreted data and helped to write the manuscript. T.T. conceived and supervised all aspects of the project, designed experiments, interpreted the data and wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 UV radiation induces immune cell recruitment and melanocyte proliferation in the skin of HGF–CDK4(R24C) mice.
a, Experimental protocol for the short-term skin inflammation assay with UV (left) and representative CD45-immunostained skin sections of adult HGF–CDK4(R24C) mice exposed twice to 4.5 kJ m−2 UV compared to controls (right). b, Flow cytometry gating strategy to analyse the composition of skin-infiltrating inflammatory cells following UV exposure (upper and middle panel) and cumulative results (lower panel, n = 10 in each group, mean percentage ± s.e.m.; unpaired two-tailed Student’s t-test; *P < 0.05; **P < 0.01; ***P < 0.001). c, Flow cytometry gating strategy to analyse the composition of immune cells in the peripheral blood following UV exposure (upper and middle panel) and cumulative results (lower panel, n = 10 in each group, mean percentage ± s.e.m.; unpaired two-tailed Student’s t-test; P < 0.05; **P < 0.01). d, Experimental protocol for the chronic skin inflammation assay with UV (left) and representative H&E-stained skin sections of adult HGF–CDK4(R24C) mice exposed twice weekly to 4.5 kJ m−2 UV for 6 weeks compared to controls (right).
Extended Data Figure 2 Detailed morphologic and immunologic characterization of angiotropic growth and metastasis in mice with UV-irradiated primary HGF-CDK4(R24C) melanomas.
a, Photographs of the dermal interstitium showing macroscopically visible expansion of pigmented primary HGF-CDK4(R24C) melanomas (arrows) along blood vessels. b, Corresponding H&E-stained sections. c, Immunofluorescence images for melanoma cells (gp100, green) and endothelial cells (Meca32, red). d, Ultrastructural localization of melanoma cells (MC) on the abluminal surface of endothelial cells (EC). Middle picture shows a small cleft (arrows) with unstructured extracellular matrix separates MC and EC and indicates an activated, angiogenic phenotype. Lower picture shows a rare example of MC invading an opened EC–EC contact (border by dashed lines), reaching the vessel lumen. e, Angiotropism score of primary HGF-CDK4(R24C) melanomas exemplified by H&E stained sections (in analogy to procedures established in routine clinical pathology for human melanomas). f, Effect of UV irradiation on angiotropic growth of primary HGF-CDK4(R24C) melanomas (n = 20 in both cohorts; chi-square test **P < 0.01). g, Flow cytometric analyses of melanoma-infiltrating ímmune cells following UV exposure compared to controls (n = 20 in both cohorts, mean percentage ± s.e.m.; unpaired two-tailed Student’s t-test *P < 0.05; **P < 0.01). h, Macroscopic appearance of angiotropism near a draining lymph node (left) and corresponding H&E stained sections at low (100×, middle) and high magnification (400×, right). i, Macroscopic appearance of spontaneous lung metastases (left panel) and corresponding H&E stained sections (100×, right). j, Ultrastructural localization of melanoma cells (M) between the abluminal surface of lung capillaries (C) and alveolar epithelial cells (AE) type I and II, type II are recognizable by lamellar bodies (asterisk). Melanoma cells are separated from endothelial cells (EC) by a very thin layer of dense homogenous extracellular matrix (arrows). k, Number of macroscopically visible metastases on the lung surfaces of individual HGF–CDK4(R24C) mice (n = 20 per cohort, the bar indicates the mean, unpaired two-tailed Mann–Whitney U-test **P < 0.01). l, Flow cytometric analyses of inflammatory cells in peripheral blood and lungs of UV-exposed and control melanoma-bearing mice (n = 20 per cohort, mean percentage ± s.e.m.; unpaired two-tailed Student’s t-test *P < 0.05; **P < 0.01).
Extended Data Figure 3 MYD88-dependent TLR4-signalling predominantly in myeloid immune cells and the presence of neutrophils are required for the UV-induced skin inflammatory response.
a, Breeding scheme for the generation of conditional knock-in mice expressing MYD88 only in LysM+ myeloid immune cells or K5+ keratinocytes with appropriate global MYD88-deficient and wild-type littermate controls. b, ELISA measurement for TNF-release from bone marrow derived macrophages (top, n = 9 in each group, mean ± s.e.m.; unpaired two-tailed Student’s t-test **P < 0.01) and immunoblot for MYD88 and GAPDH in tissue homogenates of skin and spleen (bottom) derived from mice with the indicated genotypes. c, Skin sections of indicated mice exposed twice to UV and immunostained for K6 (upper panel) and CD45 (lower panel). d, Macroscopic appearance of blood vessels in the dermis (upper panel) and corresponding Meca32-stained skin sections (lower panel) of indicated mice exposed twice to UV. e, Number of vessel branching points (left) and mean number of small (5–10 µm) and larger (>10 µm) blood vessels (right) after UV irradiation (n = 10 in each group, mean ± s.e.m.; unpaired two-tailed Student’s t-test *P < 0.05; **P < 0.01). f, Experimental protocol for neutrophil or macrophage depletion. Groups of mice were treated as indicated with clodronate lipsomes (Clo Lip) or empty liposomes (Ctrl Lip), anti-Ly6G monoclonal antibody or control monoclonal antibody and exposed twice to 4.5 kJ m−2 UV. g, Flow cytometric analyses showing the percentage of myeloid immune cells in the skin (upper panel, left 4 graphs) and in the blood (upper panel, right 2 graphs) and relative increases of skin-infiltrating CD45+ immune cells and of epidermal thickness (lower panel, left 2 graphs) as well as of the corresponding mRNA expression levels in the skin for the depicted genes (lower panel, right 4 graphs) of mice treated as indicated (n = 10 in each group, mean percentage ± s.e.m.; unpaired two-tailed Student’s t-test *P < 0.05; ***P < 0.001).
Extended Data Figure 4 Genetic as well as pharmacologic blockade of the HMGB1/TLR4 signalling axis or depletion of neutrophils abrogates the ability of UV irradiation to promote spontaneous lung metastases in mice bearing serial HCmel12 skin transplants.
a, Angiotropism score in serial HCmel12 melanoma skin transplants exemplified by H&E stained sections. b, Effect of UV irradiation on angiotropic growth (n = 15 in both cohorts; Chi-square test *P < 0.05). c, Flow cytometric analyses of melanoma-infiltrating myeloid immune cells in UV-exposed mice compared to controls (n = 15 in both cohorts, mean percentage ± s.e.m.; unpaired two-tailed Student’s t-test *P < 0.05; ***P < 0.001). d, Number of lung metastases in individual mice bearing serial HCmel12 melanoma skin transplants (n = 15 in each cohort, the bar indicates the mean; unpaired two-tailed Mann–Whitney U test **P < 0.01). e, Flow cytometric analyses of myeloid immune cells in the peripheral blood following UV exposure compared to controls (n = 15 in both cohorts, mean percentage ± s.e.m.; unpaired two-tailed Student’s t-test **P < 0.01). f, Growth kinetics of HCmel12 melanoma skin transplants in groups of UV-irradiated Tlr4−/− and Myd88−/− mice compared to controls. g, Left, number of lung metastasis in individual UV-exposed mice compared to untreated controls (n = 12 in each cohort). Right, flow cytometric analyses of melanoma-infiltrating myeloid immune cells following UV exposure compared to controls (n = 12 in each cohort, mean percentage ± s.e.m.). h, Growth kinetics of HCmel12 melanoma skin transplants in groups of UV-irradiated in Myd88MYEL and Myd88Litt mice. i, Left, Corresponding number of lung metastasis (n = 12 in each cohort, the bar indicates the mean; unpaired two-tailed Mann–Whitney U-test *P < 0.05; **P < 0.01). Right, corresponding flow cytometric analyses of melanoma-infiltrating myeloid immune cells (n = 12 in both cohorts, mean percentage ± s.e.m. unpaired two-tailed Student’s t-test *P < 0.05; **P < 0.01). j, Growth kinetics of HCmel12 skin transplants in groups of UV-irradiated mice treated with anti-Ly6G monoclonal antibody or control monoclonal antibody. k, Left, corresponding number of lung metastasis (n = 12 in each cohort, the bar indicates the mean; unpaired two-tailed Mann–Whitney U-test **P < 0.01). Right, corresponding flow cytometric analyses of melanoma-infiltrating myeloid immune cells (n = 12 in both cohorts, mean percentage ± s.e.m.). l, Growth kinetics of HCmel12 skin transplants in groups of UV-irradiated mice treated with glycyrrhizin (Gly), CLI-095 (CLI) or vehicle as indicated compared to controls. m, Top, corresponding number of lung metastasis (n = 12 in each cohort, the bar indicates the mean; unpaired two-tailed Mann–Whitney U-test *P < 0.05; **P < 0.01). Bottom, corresponding melanoma-infiltrating myeloid immune cells in mice treated as indicated (n = 12 in each cohort, mean percentage ± s.e.m.). Cumulative results of 3 independent experiments with 4 or 5 mice per group are shown.
Extended Data Figure 5 Inflammation promotes the ability of mouse melanoma cells to migrate towards and spread along blood vessel endothelial cell surfaces in aortic ring and ear tissue explants.
a, Cytokine levels measured by ELISA in UV-irradiated skin and in medium conditioned by TLR4-activated neutrophils (Neu), n = 3; mean ± s.e.m.; b, In vitro morphology of HCmel12 cells (left) and immunoblot analyses for TRP2 and gp100 (right). c, Preferential 2D migration of HCmel12 cells on surfaces of bEND.5 endothelial cells compared to surfaces of SP1 keratinocytes or the matrix proteins fibronectin (Fn1) and collagen IV (Col4) under inflammatory conditions. Shown are migration velocities (left) and average accumulated distances (right) of individual cells in representative experiments (n = 70 in each group, bars indicate the mean; unpaired two-tailed Mann–Whitney U test; **P < 0.01; ***P < 0.001). d, Aortic ring endothelial sprouting at day 7 in response to inflammatory activation with neutrophil conditioned medium (Neu), impact of TNFR2-IgG:Fc and effect of 1000 U ml−1 recombinant mouse TNF. e. Quantification of sprouts using aortic rings from Actb-DsRed or wild-type mice (n = 12 in each group; mean ± s.e.m.; unpaired two-tailed Student’s t-test; **P < 0.01). f, Effect of neutrophil conditioned medium (Neu) and of recombinant mouse TNF on the interaction between EGFP-expressing HCmel12 cells and aortic ring endothelial sprouts at days 4 and 7 compared to control co-cultures. Shown are merged phase contrast and fluorescence microscopic images for the indicated conditions and time points. g, Confocal images of EGFP-expressing HCmel12 melanoma cells (green) expanding along an aortic ring endothelial sprout from Actb-DsRed mice (red) in a pericyte-like manner under inflammatory conditions. All images are representative of at least 3 independent experiments with 2 or more replicates. h, Experimental protocol for the generation of UV-inflamed ear tissue explants. i, Confocal immunofluorescence image of whole mount ear tissue showing increased immune cell infiltration and dilation of CD31+ blood vessels in the dermal interstitium following UV exposure. j, Experimental protocol for ear tissue invasion assays with EGFP-expressing melanoma cells. k, Confocal immunofluorescence images of whole mount ear tissue explants from UV-irradiated mice showing EGFP-expressing HCmel31 melanoma cells cuffing CD31+ blood vessel endothelial cells. l, Volume-rendered, 3D reconstruction of a magnification from the indicated area in k.
Extended Data Figure 6 TNF shifts human melanoma cells towards a migratory phenotype that uses selective motility cues on endothelial cell surfaces.
a, In vitro morphology of the indicated human melanoma cells (left panel), effect of inflammatory activation with TNF on f-actin distribution determined by phalloidin immunostaining, indicating a shift towards a migratory phenotype (right panel). b. Migration of control and TNF-activated human melanoma cells towards control or TNF-activated HUVEC in transwell assays. Shown are mean number of migrated cells in triplicate determinations ± s.e.m. of 1 representative experiment out of 4 (Med = medium only, unpaired two-tailed Mann–Whitney U-test; *P < 0.05; **P < 0.01; ***P < 0.001). c, Static adhesion to the indicated extracellular matrix components. Shown are means of triplicate determinations ± s.e.m. of 1 representative experiment out of 3 (unpaired two-tailed Student’s t-test; *P < 0.05; ***P < 0.001). d, 2D migration of control and TNF-activated melanoma cells on HUVEC surfaces. e, 2D migration of TNF-activated melanoma cells on surfaces of HUVEC or Collagen IV (Col4) alone. Shown are average accumulated distances (left) and migration velocities (right) of individual cells in 1 representative experiment out of 3 (n = 70 in each group, bars indicate the mean; unpaired two-tailed Mann–Whitney U-test; *P < 0.05; **P < 0.01; ***P < 0.001). f, Confocal immunofluorescence images of EGFP-expressing human melanoma cells and CD31+ endothelial cells in ear tissue invasion assays confirming the ability of human melanoma cells to migrate towards and closely associate with endothelial cells in a physiological environment.
Extended Data Figure 7 TNF induces genes in human melanoma cells related to the biological processes angiogenesis, cell migration and cell adhesion.
a, Gene ontology (GO) analysis of TNF induced genes (>twofold in 4/5 cell lines) using a publicly available gene expression data set (GSE19428) of five human melanoma cell lines treated with TNF. GO enrichment analysis was done by hypergeometric testing and P values were corrected for multiple testing by the Benjamini & Hochberg (B & H) method (Supplementary Table 1). b, Removal of GO term redundancy and semantic visualization of the identified top scoring GO terms using REViGO (http://revigo.irb.hr/). Positioning in the semantic space indicates semantic and hence functional similarity of GO terms, but the semantic space units have no intrinsic meaning. B & H corrected P values are indicated by colour coding. The central GO terms angiogenesis (GO:0001525), cell migration (GO:0016477) and cell adhesion (GO:0007155) are highlighted in red letters. c, A cluster of gene probes (orange) out of the GO terms cell migration, cell adhesion and angiogenesis is highly expressed at baseline (without TNF treatment) in a subset of human melanoma cell lines (yellow cluster). There was no significant association of the yellow cluster with BRAF or NRAS mutations (two-sided Fisher’s exact test, P = 0.51 and P = 0.43). Gene expression data was downloaded from the BROAD melanoma portal (http://www.broadinstitute.org/melanoma/) and visualized by a heatmap of mean centred log2-normalized gene expression values as indicated by the colour key. Heat map legend key: WT, wild-type; MUT, mutated; NA, not available. d, TNF responsiveness of gene probes (Mean log2-fold change of five melanoma cell lines, GSE19428) is matched to the baseline expression in the BROAD melanoma cell line panel shown in the heat map (in c). Affymetrix microarray platforms were used for both data sets (Hgu133A and Hgu133A 2.0) with matched gene probes. Genes probes of the orange cluster are predominantly TNF inducible and show high baseline expression in the subset of melanoma cell lines highlighted in yellow in panel c. Representative genes of the orange cluster are shown in the orange box. Gene probe lists are provided in Supplementary Table 2.
Extended Data Figure 8 Similarity of transcriptional responses to TNF in RAS and BRAF mutant human melanoma cell lines and conservation of a core set of TNF-regulated genes involved in tumour–endothelial interactions and angiogenesis in human and mouse melanomas.
a, A panel of human melanoma cell lines with known RAS and BRAF mutation status was either treated with TNF for 72 h or left untreated. Schematic view of experimental setup and outline of bioinformatic analysis. b, Comparison of TNF regulated genes (gene probes) in RAS mutant (n = 7) versus BRAF mutant (n = 8) human melanoma cells. The mean log2-fold changes caused by TNF treatment in the respective subgroups were plotted against each other. The dashed diagonal depicts identical changes by TNF in both subgroups. The smoothed blue coloured density plot indicates that the majority of the probes were not regulated by TNF (log2-fold changes close to zero in both groups). Grey dots represent TNF regulated genes and red dots represent TNF regulated genes that belong to the GO categories angiogenesis (GO:0001525), cell migration (GO:0016477) and cell adhesion (GO:0007155). Pearson’s product-moment correlation coefficient (r) and P values (two-sided correlation test) were calculated for TNF mediated transcriptional changes in RAS versus BRAF mutant melanoma cells and indicated at the top of the plot either for all TNF regulated gene probes or for those belonging to the three selected GO categories (t, correlation test statistic; d.f., degree of freedom). Representative TNF regulated genes out of the three GO categories are highlighted by red dots with black circles and their gene symbols are indicated. Gene probe lists are provided as Supplementary Table 3. c, Relative change of mRNA expression levels for the indicated genes as determined by qRT–PCR in human and mouse melanoma cell lines following 72 h of exposure to 1,000 U ml−1 human or mouse TNF. Mean log2 fold change ± s.e.m. of n = 3 biological replicates are shown.
Extended Data Figure 9 Association of angiotropism with tumour thickness, nodular growth pattern, ulceration, positive sentinel node status and metastasis-free survival in an unselected and representative cohort of 178 patients that underwent excision of their primary cutaneous melanoma and sentinel lymph node biopsy at the University Hospital of Bonn Dermatology Department (2000–2010).
a, Clinico-pathologic characteristics of patients with primary melanomas with and without angiotropism. Statistical evaluation was performed with the indicated tests. b, Representative CD31 immunostained section of an ulcerated and angiotropic primary melanoma. c, Kaplan–Meier survival curves for metastasis free survival in patient cohorts with and without ulceration (left) or with and without angiotropism (right) of their primary melanoma.
Extended Data Figure 10 Graphical abstract illustrating how UV irradiation induces a neutrophilic skin inflammatory response that drives angiotropic growth and systemic metastatic spread of cutaneous melanoma.
a, UV irradiation of the skin causes DNA damage in epidermal keratinocytes leading to release of HMGB1. b, HMGB1 initiates the recruitment of immune cells, most notably neutrophils and inflammatory monocytes, in a TLR4/MYD88-dependent manner. c, The neutrophilic inflammatory environment shifts melanoma cells towards a migratory phenotype and activates angiogenesis. d, The local release of additional endogenous TLR4 ligands such as s100a8/a9 by activated immune cells fuels autocrine and paracrine feed-forward signalling loops that further amplify the neutrophilic inflammatory cytokine–chemokine cascade and drive the production of TNF, IL1 and CXCL2. e, Melanoma cells closely associate and interact with endothelial cells in a pericyte-like manner. f, Melanoma cells use motility and guidance cues on endothelial surfaces for perivascular expansion. This increases the likelihood of intravasation and hematogenous dissemination. g, Melanoma cells preferentially colonise the lung where activated neutrophils may provide a metastatic niche. h, Melanoma cells again closely interacting with endothelial cells on their luminal and abluminal surfaces.
Supplementary information
Supplementary Table 1
This file contains the gene ontology terms enriched in TNF stimulated human melanoma cell lines (n=5) using the published dataset GSE19428 as identified by hypergeometric testing. Raw and corrected p-values (Benjamini&Hochberg) are given within the table. Corresponds to Extended Data Figure 7. (XLS 136 kb)
Supplementary Table 2
This file contains a list of gene probes out of gene ontology terms angiogenesis (GO:0001525), cell migration (GO:0016477) and cell adhesion (GO:0007155) that are represented in the heatmap and the barplot of Extended Data Figure 7c,d. The position of the probes in the heatmap and their mean responsiveness to TNF (GSE19428) are indicated. (XLS 104 kb)
Supplementary Table 3
The file shows the core set of genes/gene probes induced by TNF in the „Ma.Mel“ melanoma cell line panel (n=17, GSE51221). Two separate lists are provided for genes that belong to the GO terms angiogenesis, cell migration and cell adhesion and for genes that are not annotated within these GO terms. (XLS 146 kb)
Supplementary Table 4
This file contains the PCR primer sequences. (XLS 35 kb)
2D migration of HCmel12 melanoma cells on endothelial cell surfaces
EGFP expressing HCmel12 cells were seeded on a confluent layer of murine bEND.5 blood endothelial cells in 8µ chamber slides. Melanoma cell migration was analysed by time lapse video microscopy over a period of 12h (one frame / 5minutes). Motile cells were tracked automatically using Imaris Software (Bitplane). Coloured dots represent individual cells, coloured lines represent individual tracks of motile cells. (MPG 2330 kb)
2D migration of TNF-stimulated HCmel12 melanoma cells on endothelial cell surfaces
EGFP expressing HCmel12 cells were stimulated for 72 hours with 1000U/ml recombinant TNF and seeded on a confluent layer of murine bEND.5 blood endothelial cells in 8µ chamber slides. Melanoma cell migration was analysed by time lapse video microscopy over a period of 12h (one frame / 5minutes). Motile cells were tracked automatically using Imaris Software (Bitplane). Coloured dots represent individual cells, coloured lines represent individual tracks of motile cells. (MPG 2330 kb)
3D-rendering of HCmel12 cells in a pericyte-like location interacting with blood vessels in ear tissue explants
EGFP expressing HCmel12 melanoma cells were seeded on the ventral side of inflamed ear tissue explants from UV-irradiated C57BL/6 mice. HCmel12 cells were allowed to adhere for two hours and invade the ear tissue for 16 hours. Ears were fixed and blood endothelial cells were stained with an anti-CD31 antibody followed by an Alexa594-conjugated secondary antibody. Images were acquired with an upright LSM780 confocal laser-scanning microscope (Carl Zeiss Microimaging). Volume-rendered 3-D reconstruction and animation performed on the z-series, was performed using Imaris software (Bitplane). (MPG 1722 kb)
3D-rendering of human MZ7-MEL cells in a pericyte-like location interacting with blood vessels in ear tissue explants
EGFP expressing MZ7-MEL melanoma cells were seeded on the ventral side of inflamed ear tissue explants from UV-irradiated C57BL/6 mice. MZ7-MEL cells were allowed to adhere for two hours and invade the ear tissue for 16 hours. Ears were fixed and blood endothelial cells were stained with an anti-CD31 antibody followed by an Alexa594-conjugated secondary antibody. Images were acquired with an upright LSM780 confocal laser-scanning microscope (Carl Zeiss Microimaging). Volume-rendered 3-D reconstruction and animation performed on the z-series, was performed using Imaris software (Bitplane). (MPG 1722 kb)
Rights and permissions
About this article
Cite this article
Bald, T., Quast, T., Landsberg, J. et al. Ultraviolet-radiation-induced inflammation promotes angiotropism and metastasis in melanoma. Nature 507, 109–113 (2014). https://doi.org/10.1038/nature13111
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13111
This article is cited by
-
Dormancy of cutaneous melanoma
Cancer Cell International (2024)
-
Exosomes from hypoxic pretreated ADSCs attenuate ultraviolet light-induced skin injury via GLRX5 delivery and ferroptosis inhibition
Photochemical & Photobiological Sciences (2024)
-
The evolution and heterogeneity of neutrophils in cancers: origins, subsets, functions, orchestrations and clinical applications
Molecular Cancer (2023)
-
The multifunctional protein HMGB1: 50 years of discovery
Nature Reviews Immunology (2023)
-
The journey from melanocytes to melanoma
Nature Reviews Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.