Abstract
The seasonal human influenza A/H3N2 virus undergoes rapid evolution, which produces significant year-to-year sequence turnover in the population of circulating strains. Adaptive mutations respond to human immune challenge and occur primarily in antigenic epitopes, the antibody-binding domains of the viral surface protein haemagglutinin. Here we develop a fitness model for haemagglutinin that predicts the evolution of the viral population from one year to the next. Two factors are shown to determine the fitness of a strain: adaptive epitope changes and deleterious mutations outside the epitopes. We infer both fitness components for the strains circulating in a given year, using population-genetic data of all previous strains. From fitness and frequency of each strain, we predict the frequency of its descendent strains in the following year. This fitness model maps the adaptive history of influenza A and suggests a principled method for vaccine selection. Our results call for a more comprehensive epidemiology of influenza and other fast-evolving pathogens that integrates antigenic phenotypes with other viral functions coupled by genetic linkage.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bao, Y. et al. The Influenza Virus Resource at the National Center for Biotechnology Information. J. Virol. 82, 596–601 (2008)
Wiley, D. C., Wilson, I. A. & Skehel, J. J. Structural identification of the antibody-binding sites of Hong Kong influenza haemagglutinin and their involvement in antigenic variation. Nature 289, 373–378 (1981)
Bush, R. M., Bender, C. A., Subbarao, K., Fox, N. J. & Fitch, W. M. Predicting the evolution of human influenza A. Science 286, 1921–1925 (1999)
Plotkin, J. B., Dushoff, J. & Levin, S. A. Hemagglutinin sequence clusters and the antigenic evolution of influenza A virus. Proc. Natl Acad. Sci. USA 99, 6263–6268 (2002)
Koelle, K., Cobey, S., Grenfell, B. & Pascual, M. Epochal evolution shapes the phylodynamics of interpandemic influenza A (H3N2) in humans. Science 314, 1898–1903 (2006)
Wolf, Y. I., Viboud, C., Holmes, E. C., Koonin, E. V. & Lipman, D. J. Long intervals of stasis punctuated by bursts of positive selection in the seasonal evolution of influenza A virus. Biol. Direct 1, 34 (2006)
Shih, A. C.-C., Hsiao, T.-C., Ho, M.-S. & Li, W.-H. Simultaneous amino acid substitutions at antigenic sites drive influenza A hemagglutinin evolution. Proc. Natl Acad. Sci. USA 104, 6283–6288 (2007)
Bhatt, S., Holmes, E. C. & Pybus, O. G. The genomic rate of molecular adaptation of the human influenza A virus. Mol. Biol. Evol. 28, 2443–2451 (2011)
Strelkowa, N. & Lässig, M. Clonal interference in the evolution of influenza. Genetics 192, 671–682 (2012)
Illingworth, C. J. R. & Mustonen, V. Components of selection in the evolution of the influence virus: linkage effects beat inherent selection. PLoS Pathog. 8, e1003091 (2012)
Meyer, A. G., Dawson, E. T. & Wilke, C. O. Cross-species comparison of site-specific evolutionary-rate variation in influenza haemagglutinin. Phil. Trans. R. Soc. B 368, 20120334 (2013)
Smith, D. J. et al. Mapping the antigenic and genetic evolution of Influenza virus. Science 305, 371–376 (2004)
Bloom, J. D. & Glassman, M. J. Inferring stabilizing mutations from protein phylogenies: application to influenza hemagglutinin. PLOS Comput. Biol. 5, e1000349 (2009)
Wylie, C. S. & Shakhnovich, E. I. A biophysical protein folding model accounts for most mutational fitness effects in viruses. Proc. Natl Acad. Sci. USA 108, 9916–9921 (2011)
Holmes, E. C. et al. Whole-genome analysis of human inuenza A virus reveals multiple persistent lineages and reassortment among recent H3N2 viruses. PLoS Biol. 3, e300 (2005)
Grenfell, B. T. et al. Unifying the epidemiological and evolutionary dynamics of pathogens. Science 303, 327–332 (2004)
Rambaut, A. et al. The genomic and epidemiological dynamics of human influenza A virus. Nature 453, 615–619 (2008)
Russell, C. A. et al. The global circulation of seasonal influenza A (H3N2) viruses. Science 320, 340–346 (2008)
Gog, J. R. & Grenfell, B. T. Dynamics and selection of many-strain pathogens. Proc. Natl Acad. Sci. USA 99, 17209–17214 (2002)
Tria, F., Lässig, M., Peliti, L. & Franz, S. A minimal stochastic model for influenza evolution. J. Stat. Mech. P07008 (2005)
Kryazhimskiy, S., Dieckmann, U., Levin, S. A. & Dushoff, J. On state-space reduction in multi-strain pathogen models, with an application to antigenic drift in influenza A. PLOS Comput. Biol. 3, e159 (2007)
Minayev, P. & Ferguson, N. Improving the realism of deterministic multi-strain models: implications for modelling influenza A. J. R. Soc. Interface 6, 509–518 (2009)
Rasmussen, D. A., Ratmann, O. & Koelle, K. Inference for nonlinear epidemiological models using genealogies and time series. PLOS Comput. Biol. 7, e1002136 (2011)
Kryazhimskiy, S., Dushoff, J., Bazykin, G. A. & Plotkin, J. B. Prevalence of epistasis in the evolution of influenza A surface proteins. PLoS Genet. 7, e1001301 (2011)
Bogner, P. et al. A global initiative on sharing avian flu data. Nature 442, 981 (2006)
Huang, J.-W., Lin, W.-F. & Yang, J.-M. Antigenic sites of H1N1 influenza virus hemagglutinin revealed by natural isolates and inhibition assays. Vaccine 30, 6327–6337 (2012)
WHO Recommendations for Influenza Vaccine Composition. Retrieved from http://www.who.int/influenza/vaccines/vaccinerecommendations1/en
Mustonen, V. & Lässig, M. From fitness landscapes to seascapes: non-equilibrium dynamics of selection and adaptation. Trends Genet. 25, 111–119 (2009)
Mustonen, V. & Lässig, M. Fitness flux and ubiquity of adaptive evolution. Proc. Natl Acad. Sci. USA 107, 4248–4253 (2010)
Desai, M. M. & Fisher, D. S. Beneficial mutation selection balance and the effect of linkage on positive selection. Genetics 176, 1759–1798 (2007)
Rouzine, I. M., Brunet, E. & Wilke, C. O. The traveling-wave approach to asexual evolution: Mullers ratchet and speed of adaptation. Theor. Popul. Biol. 73, 24–46 (2008)
Schiffels, S., Szollosi, G. J., Mustonen, V. & Lässig, M. Emergent neutrality in adaptive asexual evolution. Genetics 189, 1361–1375 (2011)
Good, B. H., Rouzine, I. M., Balick, D. J., Hallatschek, O. & Desai, M. M. Distribution of fixed beneficial mutations and the rate of adaptation in asexual populations. Proc. Natl Acad. Sci. USA 109, 4950–4955 (2012)
Neher, R. A. & Hallatschek, O. Genealogies of rapidly adapting populations. Proc. Natl Acad. Sci. USA 110, 437–442 (2013)
Gerrish, P. J. & Lenski, R. E. The fate of competing beneficial mutations in an asexual population. Genetica 102–103, 127–144 (1998)
Miralles, R., Gerrish, P. J., Moya, A. & Elena, S. F. Clonal interference and the evolution of RNA viruses. Science 285, 1745–1747 (1999)
Ghedin, E. et al. Large-scale sequencing of human influenza reveals the dynamic nature of viral genome evolution. Nature 437, 1162–1166 (2005)
Bush, R. M., Smith, C. B., Cox, N. J. & Fitch, W. M. Effects of passage history and sampling bias on phylogenetic reconstruction of human influenza A evolution. Proc. Natl Acad. Sci. USA 97, 6974–6980 (2000)
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2–approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010)
Swofford, D. L. PAUP*: Phylogenetic Analysis Using likelihood (and Other Methods) 4.0 Beta (Sinauer Associates, 2002)
Weis, W. et al. Structure of the influenza virus haemagglutinin complexed with its receptor, sialic acid. Nature 333, 426–431 (1988)
Wilson, I. A. & Cox, N. J. Structural basis of immune recognition of influenza virus hemagglutinin. Annu. Rev. Immunol. 8, 737–787 (1990)
Skehel, J. J. & Wiley, D. C. Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Annu. Rev. Biochem. 69, 531–569 (2000)
Lin, Y. P. et al. Evolution of the receptor binding properties of the influenza A(H3N2) hemagglutinin. Proc. Natl Acad. Sci. USA 109, 21474–21479 (2012)
Andreasen, V., Lin, J. & Levin, S. A. The dynamics of cocirculating strains conferring partial cross-immunity. J. Math. Biol. 35, 825–842 (1997)
Gog, J. R. & Swinton, J. A status-based approach to multiple strain dynamics. J. Math. Biol. 44, 169–184 (2002)
Koelle, K., Khatri, P., Kamradt, M. & Kepler, T. A two-tiered model for simulating the ecological and evolutionary dynamics of rapidly evolving viruses, with an application to influenza. J. R. Soc. Interface 7, 1257–1274 (2010)
Molinari, N. M. et al. The annual impact of seasonal influenza in the US: measuring disease burden and costs. Vaccine 25, 5086–5096 (2007)
Myers, J. L. et al. Compensatory hemagglutinin mutations alter antigenic properties of influenza viruses. J. Virol. 87, 11168–11172 (2013)
Schulze, I. T. Effects of glycosylation on the properties and functions of influenza virus hemagglutinin. J. Infect. Dis. Aug. 176, (suppl. 1)S24–S28 (1997)
Blackburne, B. P., Hay, A. J. & Goldstein, R. A. Changing selective pressure during antigenic changes in human influenza H3. PLoS Pathog. 4, e1000058 (2008)
Cui, J., Smith, T., Robbins, P. W. & Samuelson, J. Darwinian selection for sites of Asn-linked glycosylation in phylogenetically disparate eukaryotes and viruses. Proc. Natl Acad. Sci. USA 106, 13421–13426 (2009)
Zhang, M. et al. Tracking global patterns of N-linked glycosylation site variation in highly variable viral glycoproteins: HIV, SIV, and HCV envelopes and influenza hemagglutinin. Glycobiology 14, 1229–1246 (2004)
Arinaminpathy, N. & Grenfell, B. Dynamics of glycoprotein charge in the evolutionary history of human influenza. PLoS ONE 5, e15674 (2010)
Ampofo, W. K. et al. Improving influenza vaccine virus selection. Report of a WHO informal consultation held at WHO headquarters, Geneva, Switzerland, 14–16 June 2010. Influenza Other Respir. Viruses 6, 147–152 (2010)
Osterholm, M. T., Kelley, N. S., Sommer, A. & Belongia, E. A. Efficacy and effectiveness of influenza vaccines: a systematic review and meta-analysis. Lancet Infect. Dis. 12, 36–44 (2012)
Press, W. H., Teukolsky, S. A., Vetterling, W. T. & Flannery, B. P. Numerical Recipes: the Art of Scientific Computing 3rd edn (Cambridge Univ. Press, 2007)
Mustonen, V. & Lässig, M. Adaptations to fluctuating selection in Drosophila. Proc. Natl Acad. Sci. USA 104, 2277–2282 (2007)
Tsimring, L. S., Levine, H. & Kessler, D. A. RNA virus evolution via a fitness-space model. Phys. Rev. Lett. 76, 4440–4443 (1996)
Acknowledgements
We acknowledge discussions with B. D. Greenbaum, B. Grenfell, C. Illingworth, A. Levine, J. W. McCauley, V. Mustonen, S. Pompei and R. Rabadan. This work has been partially supported by Deutsche Forschungsgemeinschaft grant SFB 680 and by German Federal Ministry of Education and Research grant 0315893-Sybacol. Part of this work was performed at the Kavli Institute of Theoretical Physics (Santa Barbara), which has been supported by National Science Foundation grant PHY05-51164.
Author information
Authors and Affiliations
Contributions
Both authors designed research, developed methods, analysed data, interpreted results and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
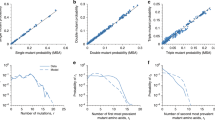
Extended Data Figure 1 Statistical errors of predicted and posterior Wrightian fitness.
The frequency ratio plot ( ,
,  ) of Fig. 2a is shown together with the standard deviation of the predicted ratio
) of Fig. 2a is shown together with the standard deviation of the predicted ratio  in the ensemble of reconstructed trees (vertical bars) and the standard deviation of the posterior ratio
in the ensemble of reconstructed trees (vertical bars) and the standard deviation of the posterior ratio  due to sampling fluctuations of population frequencies (horizontal bars). See sections 1, 3 and 4 of Methods.
due to sampling fluctuations of population frequencies (horizontal bars). See sections 1, 3 and 4 of Methods.
Extended Data Figure 2 Fitness predictions for human influenza A/H1N1.
a, Wrightian fitness: the predicted frequency ratio  is plotted against the posterior ratio
is plotted against the posterior ratio  for 81 HA clades with initial frequency
for 81 HA clades with initial frequency  > 0.1. To be compared with Fig. 2a. b, Dynamics of the influenza strain tree: for each clade, the ancestor node is coloured according to the maximum of the predicted frequency changes,
> 0.1. To be compared with Fig. 2a. b, Dynamics of the influenza strain tree: for each clade, the ancestor node is coloured according to the maximum of the predicted frequency changes,  . To be compared with Fig. 2c. The predictions are based on a sample of 2,136 unique HA1 genotypes observed between 1977 and 2009. We restrict predictions to years when this sample contains at least 12 unique HA1 strains in the winter seasons t and t + 1, which are the years 1990, 1995–1998 and 2005–2008 (see Methods, section 4). These predictions are statistically significant (P < 10−18) but somewhat more noisy than for influenza A/H3N2 (clade growth is correctly predicted in 88% of the cases, decline in 63% of the cases). The reasons include a significantly smaller and more biased strain sample and a less comprehensive knowledge of antigenic epitope sites26. The prediction of influenza A/H1N1 evolution illustrates the broader applicability of our method and highlights the determinants of predictive power.
. To be compared with Fig. 2c. The predictions are based on a sample of 2,136 unique HA1 genotypes observed between 1977 and 2009. We restrict predictions to years when this sample contains at least 12 unique HA1 strains in the winter seasons t and t + 1, which are the years 1990, 1995–1998 and 2005–2008 (see Methods, section 4). These predictions are statistically significant (P < 10−18) but somewhat more noisy than for influenza A/H3N2 (clade growth is correctly predicted in 88% of the cases, decline in 63% of the cases). The reasons include a significantly smaller and more biased strain sample and a less comprehensive knowledge of antigenic epitope sites26. The prediction of influenza A/H1N1 evolution illustrates the broader applicability of our method and highlights the determinants of predictive power.
Extended Data Figure 3 Fitness flux in the evolution of influenza.
a, Fitness flux measures adaptation (schematic, adapted from ref. 29). The cumulative flux Φ(t), as defined in equations (48) and (49), is an aggregate measure of fitness changes due to frequency changes in a population’s history up to a given clade ν at a given time t (shown by uphill arrows)28,29,58. Left: in a static fitness landscape F(x), the flux Φ(t) equals the fitness difference between the initial point and the final point of this history. Right: in a time-dependent fitness seascape F(x, t), the flux Φ(t) is still a typically positive, increasing function of time, even if the population fitness does not increase. b, Mean cumulative fitness flux Φ(t) as given by equation (48) for influenza from 1993 up to season t. The mean flux inferred from our fitness model (black line) shows a continuous increase. The flux for a null model with scrambled clade fitness values (grey lines) fluctuates around 0. Vertical bars indicate the root mean squared fitness (flux) in each year’s strain sample,  , as given by equation (51).
, as given by equation (51).
Extended Data Figure 4 Strain tree with mutations.
a–c, Four classes of HA sequence mutations are mapped onto individual branches of the influenza strain tree: synonymous mutations (a, blue), nonsynonymous epitope mutations (b, green) and nonsynonymous non-epitope mutations (c, red). Each nonsynonymous mutation marks the origination of a clade in the population; each fixed clade (highlighted by bright colours) has an origination on the trunk of the tree (shown as thick line). The fixation probability, that is, the ratio of the number of fixations and the number of originations, is seen to be reduced for informative non-epitope changes and enhanced for nonsynonymous epitope changes compared to the baseline of synonymous changes; cf. Extended Data Fig. 5. The underlying tree (shown here from 1993 to 2012) is a sample from our ensemble of strain trees, which are constructed by maximum likelihood from the HA sequence of 3,944 strains (other equiprobable trees differ only in peripheral branches). The horizontal coordinate D of a node is its mutational distance from the root of the tree. The trunk of the tree (thick line) is the single lineage connecting past and future on timescales beyond the coalescence time.
Extended Data Figure 5 Selection on epitope and non-epitope changes.
The frequency propagator ratio9 g(X), as defined in equation (7), is shown for several classes of nonsynonymous HA mutations: mutations in epitopes A–D (green bullets), mutations in epitope E (green circles), mutations in sialic receptor binding sites (green diamonds), informative non-epitope mutations (red bullets) and non-informative non-epitope mutations (red circles). Error bars indicate sampling uncertainties. Mutations in epitopes A–D, including those in epitopic receptor binding sites, reach values g(X) ≥ 2.5 for large frequencies, signalling substantial positive selection. Mutations in epitope E are under weaker positive selection, with g(X) ≈ 1.5 for large frequencies. Informative non-epitope changes drop to g(X) = 0.6, signalling predominantly negative selection. Non-informative non-epitope changes evolve near the neutral baseline (g = 1, blue line), indicating weak or heterogeneous selection. See section 2 of Methods.
Extended Data Figure 6 Evolution of glycosylation.
a, Number of epitopic glycosylation sites, nep, in the influenza A/H3N2 strain sample between 1968 and 2012 (green lines): population mean value (thin line), root mean squared variation (error bars), and value for trunk lineages (thick line). The same data are shown for non-epitope glycosylation sites (red lines). Epitope sites show substantial changes with a net increase in number and substantial natural variation in some years, whereas non-epitope sites had only one fixation of a glycosylation site. b, Evolution of nep on the influenza strain tree between 1993 and 2012. Trunk strains show a rapid increase to nep = 5 between 1995 and 1997 and maintain this value in later years; the mean nep shows a slower increase between 1995 and 2001. Off-trunk clades drop below nep = 5 also in later years, and there are compensatory mutations back to nep = 5. The data suggest an adaptive increase of nep up to a saturation value nep = 5 after 1996. These observations inform the glycosylation fitness component (equation (22)), which is used to test the predictive value of glycosylation. See section 2 of Methods.
Supplementary information
Supplementary Data
This file contains the GenBank and Gisaid accession numbers of the influenza strains used in the study. (TXT 80 kb)
Rights and permissions
About this article
Cite this article
Łuksza, M., Lässig, M. A predictive fitness model for influenza. Nature 507, 57–61 (2014). https://doi.org/10.1038/nature13087
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13087
This article is cited by
-
Evolutionary implications of SARS-CoV-2 vaccination for the future design of vaccination strategies
Communications Medicine (2023)
-
Steering and controlling evolution — from bioengineering to fighting pathogens
Nature Reviews Genetics (2023)
-
Quantitative systems-based prediction of antimicrobial resistance evolution
npj Systems Biology and Applications (2023)
-
Forecasting influenza hemagglutinin mutations through the lens of anomaly detection
Scientific Reports (2023)
-
Analysis of lineage-specific protein family variability in prokaryotes combined with evolutionary reconstructions
Biology Direct (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.