Abstract
In plants, post-transcriptional gene silencing (PTGS) is mediated by DICER-LIKE 1 (DCL1)-dependent microRNAs (miRNAs), which also trigger 21-nucleotide secondary short interfering RNAs (siRNAs) via RNA-DEPENDENT RNA POLYMERASE 6 (RDR6), DCL4 and ARGONAUTE 1 (AGO1)1,2,3, whereas transcriptional gene silencing (TGS) of transposons is mediated by 24-nucleotide heterochromatic (het)siRNAs, RDR2, DCL3 and AGO4 (ref. 4). Transposons can also give rise to abundant 21-nucleotide ‘epigenetically activated’ small interfering RNAs (easiRNAs) in DECREASED DNA METHYLATION 1 (ddm1) and DNA METHYLTRANSFERASE 1 (met1) mutants, as well as in the vegetative nucleus of pollen grains5 and in dedifferentiated plant cell cultures6. Here we show that easiRNAs in Arabidopsis thaliana resemble secondary siRNAs, in that thousands of transposon transcripts are specifically targeted by more than 50 miRNAs for cleavage and processing by RDR6. Loss of RDR6, DCL4 or DCL1 in a ddm1 background results in loss of 21-nucleotide easiRNAs and severe infertility, but 24-nucleotide hetsiRNAs are partially restored, supporting an antagonistic relationship between PTGS and TGS. Thus miRNA-directed easiRNA biogenesis is a latent mechanism that specifically targets transposon transcripts, but only when they are epigenetically reactivated during reprogramming of the germ line. This ancient recognition mechanism may have been retained both by transposons to evade long-term heterochromatic silencing and by their hosts for genome defence.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Allen, E., Xie, Z., Gustafson, A. M. & Carrington, J. C. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121, 207–221 (2005)
Ronemus, M., Vaughn, M. W. & Martienssen, R. A. MicroRNA-targeted and small interfering RNA-mediated mRNA degradation is regulated by Argonaute, Dicer, and RNA-dependent RNA polymerase in Arabidopsis. Plant Cell 18, 1559–1574 (2006)
Cuperus, J. T. et al. Unique functionality of 22-nt miRNAs in triggering RDR6-dependent siRNA biogenesis from target transcripts in Arabidopsis. Nature Struct. Mol. Biol. 17, 997–1003 (2010)
Law, J. A. & Jacobsen, S. E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nature Rev. Genet. 11, 204–220 (2010)
Slotkin, R. K. et al. Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell 136, 461–472 (2009)
Tanurdzic, M. et al. Epigenomic consequences of immortalized plant cell suspension culture. PLoS Biol. 6, e302 (2008)
McCue, A. D., Nuthikattu, S., Reeder, S. H. & Slotkin, R. K. Gene expression and stress response mediated by the epigenetic regulation of a transposable element small RNA. PLoS Genet. 8, e1002474 (2012)
Nuthikattu, S. et al. The initiation of epigenetic silencing of active transposable elements is triggered by RDR6 and 21–22 nucleotide small interfering RNAs. Plant Physiol. 162, 116–131 (2013)
Mirouze, M. et al. Selective epigenetic control of retrotransposition in Arabidopsis. Nature 461, 427–430 (2009)
Miura, A. et al. Mobilization of transposons by a mutation abolishing full DNA methylation in Arabidopsis. Nature 411, 212–214 (2001)
German, M. A., Luo, S., Schroth, G., Meyers, B. C. & Green, P. J. Construction of Parallel Analysis of RNA Ends (PARE) libraries for the study of cleaved miRNA targets and the RNA degradome. Nature Protocols 4, 356–362 (2009)
Hsieh, T. F. et al. Genome-wide demethylation of Arabidopsis endosperm. Science 324, 1451–1454 (2009)
Gehring, M. et al. DEMETER DNA glycosylase establishes MEDEA polycomb gene self-imprinting by allele-specific demethylation. Cell 124, 495–506 (2006)
Schwab, R., Speth, C., Laubinger, S. & Voinnet, O. Enhanced microRNA accumulation through stemloop-adjacent introns. EMBO Rep. 14, 615–621 (2013)
Jeong, D. H. et al. Massive analysis of rice small RNAs: mechanistic implications of regulated microRNAs and variants for differential target RNA cleavage. Plant Cell 23, 4185–4207 (2011)
Axtell, M. J. & Bartel, D. P. Antiquity of microRNAs and their targets in land plants. Plant Cell 17, 1658–1673 (2005)
Chen, H. M. et al. 22-Nucleotide RNAs trigger secondary siRNA biogenesis in plants. Proc. Natl Acad. Sci. USA 107, 15269–15274 (2010)
Marí-Ordóñez, A. et al. Reconstructing de novo silencing of an active plant retrotransposon. Nature Genet. 45, 1029–1039 (2013)
Wu, L. et al. DNA methylation mediated by a microRNA pathway. Mol. Cell 38, 465–475 (2010)
Jauvion, V., Rivard, M., Bouteiller, N., Elmayan, T. & Vaucheret, H. RDR2 partially antagonizes the production of RDR6-dependent siRNA in sense transgene-mediated PTGS. PLoS ONE 7, e29785 (2012)
Slotkin, R. K., Freeling, M. & Lisch, D. Heritable transposon silencing initiated by a naturally occurring transposon inverted duplication. Nature Genet. 37, 641–644 (2005)
Stroud, H., Greenberg, M. V. C., Feng, S., Bernatavichute, Y. V. & Jacobsen, S. E. Comprehensive analysis of silencing mutants reveals complex regulation of the Arabidopsis methylome. Cell 152, 352–364 (2013)
Jeddeloh, J. A., Stokes, T. L. & Richards, E. J. Maintenance of genomic methylation requires a SWI2/SNF2-like protein. Nature Genet. 22, 94–97 (1999)
Teixeira, F. K. et al. A role for RNAi in the selective correction of DNA methylation defects. Science 323, 1600–1604 (2009)
Poethig, R. S. Small RNAs and developmental timing in plants. Curr. Opin. Genet. Dev. 19, 374–378 (2009)
Poethig, R. S. Phase change and the regulation of shoot morphogenesis in plants. Science 250, 923–930 (1990)
Calarco, J. P. et al. Reprogramming of DNA methylation in pollen guides epigenetic inheritance via small RNA. Cell 151, 194–205 (2012)
Mosher, R. A. et al. Uniparental expression of PolIV-dependent siRNAs in developing endosperm of Arabidopsis. Nature 460, 283–286 (2009)
Bagijn, M. P. et al. Function, targets, and evolution of Caenorhabditis elegans piRNAs. Science 337, 574–578 (2012)
Shirayama, M. et al. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 150, 65–77 (2012)
Regulski, M. et al. The maize methylome influences mRNA splice sites and reveals widespread paramutation-like switches guided by small RNA. Genome Res. 23, 1651–1662 (2013)
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011)
Zhai, J. et al. MicroRNAs as master regulators of the plant NB-LRR defense gene family via the production of phased, trans-acting siRNAs. Genes Dev. 25, 2540–2553 (2011)
Acknowledgements
We thank V. Colot, O. Voinnet and A. Sarazin for sharing unpublished data and for discussions, J. Simorowski for plant genetics and J. Kendall for computational advice. We thank R.K. Slotkin for sharing data before publication. K.M.C. and M.R. were supported, in part, by a research collaboration with DuPont Pioneer. F.V.E. was supported by a fellowship from the Belgian American Educational Foundation. J.Z. was supported by a University of Delaware Graduate fellowship. Research in the Martienssen laboratory is supported by a grant from the National Institutes of Health (RO1GM067014 to R.A.M.) and by the Howard Hughes Medical Institute and Gordon and Betty Moore Foundation (GBMF3033). The authors acknowledge that this work was performed with assistance from the Cold Spring Harbor Laboratory Shared Resources, which are funded, in part, by the Cancer Center Support Grant (5PP30CA045508).
Author information
Authors and Affiliations
Contributions
R.A.M., B.C.M. and K.M.C. designed the study. F.B., F.V.E., M.R. and K.M.C. performed experiments. J.Z. and K.M.C. analysed data. R.A.M. and K.M.C. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
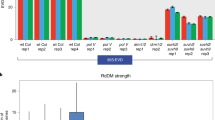
Extended Data Figure 1 Twenty-one-nucleotide easiRNAs originate from transposons in ddm1 and are miRNA and RDR6 dependent.
a, Eighteen-nucleotide to 26-nucleotide small RNA abundance in Col-0, ddm1-2 and ddm1-2 rdr6-15. Normalized reads per million (RPM). nt, nucleotides. b, Eighteen-nucleotide to 26-nucleotide small RNA abundance in Col-0, ddm1-2 and ddm1-2 dcl1-11. Normalized reads per million (RPM).
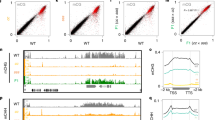
Extended Data Figure 2 miRNA target genes and transposons that do not promote tasiRNA nor easiRNA, respectively, have degradation covering the entire region.
a–f, Read pattern distribution of 21-nucleotide unique reads (represented as a histogram of read density (grey bars)) and PARE signatures at TAS2 (AT2G39681) (a), RCC (AT3G02300) and SEP2 (AT3G02310) (b), MEE58 (AT4G13940) (c), and transposons ATENSPM6 (AT2G06720) (d), ATLINE1_4 (AT2G15540) (e) and ATCOPIA43 (AT3G0410) (f), in track order Col-0, ddm1-2, rdr6-15, ddm1-2 rdr6-15 and ddm1-2 dcl1-11.
Extended Data Figure 3 Relationship between DNA methylation, easiRNA and hetsiRNA at transposons that miRNAs are predicted to target.
a–c, DNA methylation, by CG (red), CHG (blue) and CHH (+ strand, − strand) context (green) and cytosine context (scale: 1 = methylated cytosine; 0 = unmethylated cytosine) at ATCOPIA93 (AT5G17125) (a), the ATMU5 (AT4G08680) DNA transposon and the surrounding region (b) and the ATHILA6A (AT4TE15030) retrotransposon and the surrounding region (c). Twenty-one-nucleotide and 24-nucleotide siRNAs, represented as a histogram of read density (grey bars). sRNA reads: unique (U), mapping to one location of the genome; and multiple (M), mapping to more than one location of the genome. PARE read density. Track order for panels Col-0 (1), ddm1-2 (2), rdr6-15 (3) and ddm1-2 rdr6-15 (4).
Extended Data Figure 4 eamiRNA immature precursor sequence and predicted structure.
eamiRNA immature precursor sequences have methylated cytosines (asterisks) in Col-0 that are unmethylated in ddm1-2. Mature miRNAs are underlined, and the putative stem-loop structures of the precursors are illustrated.
Extended Data Figure 5 Twenty-four-nucleotide hetsiRNAs at transposons in Col-0 are lost in ddm1 and gained in ddm1 rdr6 and ddm1 dcl1.
Twenty-four-nucleotide hetsiRNAs by transposon class in Col-0, ddm1-2, rdr6-15, ddm1-2 rdr6-15 and ddm1-2 dcl1-11. Normalized reads per million (RPM).
Extended Data Figure 6 Overlap of TEs that undergo easiRNA biogenesis, hetsiRNA loss and miRNAs targeting.
a–d, Individual transposons were grouped depending on small RNA abundance in each genotype. a, TEs that lose 24-nucleotide hetsiRNAs in ddm1-2 and those that gain 21-nucleotide easiRNAs in ddm1-2 overlap with those that gain 24-nucleotide hetsiRNAs in ddm1-2 rdr6-15. b, TEs that are targeted and cleaved by miRNAs overlap with those that gain 21-nucleotide easiRNAs in ddm1-2 and 24-nucleotide hetsiRNAs in ddm1-2 rdr6-15. c, TEs that are targeted and cleaved by two or more miRNAs overlap with those that gain 21-nucleotide easiRNAs in ddm1-2 and those that gain 24-nucleotide hetsiRNAs in ddm1-5 rdr6-15. d, TEs that are predicted to be targeted by miRNAs, but without supporting PARE cleavage data, also overlap with those that gain 21-nucleotide easiRNAs in ddm1-2 and 24-nucleotide hetsiRNAs in ddm1-2 rdr6-15.
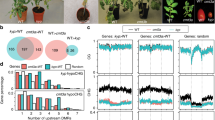
Extended Data Figure 7 Loss of methylation at transposons in ddm1 is partially restored in ddm1 rdr6.
a, Transposon methylation in Col-0, ddm1-2, rdr6-15 and ddm1-2 rdr6-15 replicates. Scale: 1 = methylated cytosine; 0 = unmethylated cytosine. b–f, Total DNA methylation at transposons, grouped by superfamily in Col-0 (b), ddm1-2 (c), rdr6-15 (d) and ddm1-2 rdr6-15 (e, f) replicates. 1, LTR retrotransposon ATGYPSY; 2, LTR retrotransposon ATCOPIA; 3, non-LTR retrotransposons ATLINE; 4, non-LTR retrotransposon TSCL; 5, TIR DNA transposon MUDR; 6, non-TIR DNA transposon MuDR; 7, DNA transposon ENSPM; 8, DNA transposon HELITRON; 9, other repeats. Total methylation by total converted cytosine to thymine and non-converted cytosine counts (at least ten reads per cytosine). Scale: 0 = unmethylated; 1 = methylated. Boxplots indicate median, range and standard deviations (box). g, h, DNA methylation by CG (red), CHG (blue) and CHH+/− context (green) at ATHILA ORF1 (AT2G10280) (g) and ATCOPIA43 (AT1G36040) (h). Track order: 1, Col-0; 2, ddm1-2; 3, rdr6-15; 4, ddm1-2 rdr6-15.
Extended Data Figure 8 Transposon transcript abundance in ddm1-2 and ddm1 rdr6.
a–c, ATH1 Affymetrix microarray expression (log2 signal intensity) in Col-0 in comparison with ddm1-2, rdr6-15 and ddm1-2 rdr6-15. d, TEs upregulated in ddm1-2 were not further upregulated in ddm1-2 rdr6-15. Red indicates transposons, black indicates genes.
Extended Data Figure 9 miRNA-directed easiRNA biogenesis from activated transposons.
When TEs are epigenetically activated, through the loss of DNA methylation and/or heterochromatin, transposon mRNA transcripts become preferentially targeted by miRNAs (DCL1 dependent) bound by AGO1. Productive cleavage of transposon transcripts engages RDR6 and DCL4, which generate 21-nucleotide easiRNAs from transposon ORFs—in a PTGS mechanism—that are then loaded into AGO1 and, thus, prevents engagement of RDR2 and RdDM. This antagonism accounts for the retention of miRNA binding sites by transposons to evade long-term heritable silencing, elicited by DNA methylation via RDR2. This model also accounts for the retention of the miRNA-directed mechanism by the host organism, in order to generate easiRNAs to silence TEs when they are epigenetically reprogrammed in the germ line.
Extended Data Figure 10 Arabidopsis miRNAs target transposons.
Known Arabidopsis miRNAs and novel eamiRNAs that arise in ddm1-2 are predicted to target transposon transcripts and confirmed to cleave transposon transcripts by PARE (Supplementary Table 3 and Methods). eamiRNAs, some known to be developmentally regulated, and TE-derived eamiRNAs that target specific transposon families. Transposons are identified by EVRY TE identifier (Supplementary Table 2; for further annotation refer to TAIR using ORF identification number). Transposon transcripts giving rise to 21-nucleotide easiRNAs (bold); those that are targeted by multiple miRNA (asterisks); and those miRNAs that target multiple transposons of the same family (italics) are highlighted.
Supplementary information
Supplementary Table 1
21-nt easiRNAs and 24-nt hetsiRNAs sRNA-seq read abundance at transposons. Ths table contains sRNA-sequencing for 21-nt and 24-nt sRNAs at transposons in Col-0, ddm1-2, rdr6-15, ddm1-2 rdr6-15 (HiSeq) and Col-0,ddm1-2 and ddm1-2 dcl1-11 (MiSeq). Normalized reads per million (RPM) for sRNA reads, unique (U) mapping to one location of the genome, and multiple (M) mapping to more than one location of the genome. (XLS 11558 kb)
Supplementary Table 2
Abundance of known Arabidopsis miRNAs and newly discovered eamiRNAs and nt size variants discovered. This table shows known and predicted miRNA abundance, sequence, length (nt) and RNA type indicating if new miRNA length at known immature miRNA precursor for Col-0, ddm1-2, rdr6-15 and ddm1-2 rdr6-15 sRNA-seq libraries. (XLS 65 kb)
Supplementary Table 3
PARE validation of predicted miRNA targeted transposon transcripts. This table shows analysis of PARE libraries by utilizing the predicted miRNA binding site within each transposon-coded open-reading frame (ORF). miRNA sequence and length are shown for each target ORF (ATX) along with the co-ordinate and genomic DNA strand (+/-) of the predicted cleavage site. PARE normalized read abundance ending within small 5-nt (s) and long 31-nt (l) windows at the miRNA binding site are shown in Col-0, ddm1-2, rdr6-15 and ddm1-2 rdr6-15 (Refer to Supplementary Methods for details), followed by the genomic co-ordinates and strand (+/-) of the target ORF. Cleavage products were detected from the same strand or different strand as the target ORF, depending whether cleavage was of the sense or antisense transcript, respectively. For statistical significance of the PARE reads at the miRNA cleavage site, we performed the Chi-square Goodness of Fit Test, calculating the p-value for observed value given the expected value. If we assume PARE tags fall randomly on all the positions in the long window (31-nt), the expected frequency for PARE reads fall into the small window (5-nt) would be 5/ 31, therefore, the expected frequency would be would 26/ 31 for other regions. (For example of the statistical test, refer to Online Methods). (XLS 2948 kb)
Supplementary Table 4
DNA methylation at transposons by cytosine context. This table shows DNA methylation by cytosine context (CG, CHG and CHH) calculated per transposon by overall count of cytosine to thymine at each cytosine position (1 = methylated cytosine, 0 = unmethylated cytosine) for each bisulphite sequencing library: Col-0, ddm1-2, rdr6-15 and two replicates of ddm1-2 rdr6-15. (XLS 5060 kb)
Supplementary Table 5
Transposons upregulated in ddm1 are not further upregulated in ddm1 rdr6. Expression of TEs by fold change normalized to Col-0 WT, in ddm1-2, rdr6-15 and ddm1-2 rdr6-15 by log2 Affymetrix microarray signal intensity. (XLS 57 kb)
Supplementary Table 6
miRNA targets and PARE cleavage data utilized for Extended data Figure 1. For known miRNAs targeting TEs passing stringent criteria based on PARE, confirming targeting, both abundance and strand orientation is given at transposons. (Refer to Methods Summary, to accompany Table 1, adapted from Supplementary Table 3). 86/ 92 TEs with no uniquely mapping easiRNA are cleaved on the antisense strand. (XLS 79 kb)
Supplementary Table 7
eamiRNAs and PARE data for Table 1. Epigenetically-activated miRNAs with relaxed criteria for PARE cleavage data, confirming targeting, with both abundance and strand orientation given at TEs (refer to Methods Summary, to accompany Table 1, adapted from Supplementary Table 3). (XLS 86 kb)
Supplementary Table 8
Sequencing information for libraries described in the study. Read count for sRNA-seq, BIS-seq and PARE libraries used in this study, mapped to the Col-0 reference genome; TAIR10 (refer to Methods Summary for computational details). (XLS 45 kb)
Supplementary Table 9
PARE confirming specific miRNA cleavage at transposon transcripts. Analysis of PARE libraries by utilizing the predicted miRNA binding site within each transposon-coded open-reading frame (ORF). miRNA sequence and length are shown for each target ORF along with the co-ordinate and genomic DNA strand (+/-) of the predicted cleavage site. PARE normalized read abundance ending within the shorter small 1-nt (ss) and shorter long 3-nt (sl) windows at the miRNA binding site are shown in Col-0, ddm1-2, rdr6-15 and ddm1-2 rdr6-15 (Refer to Supplementary Methods for details), followed by the genomic co-ordinates and strand (+/-) of the target ORF. Cleavage products were detected from the same strand or different strand as the target ORF, depending whether cleavage was of the sense or antisense transcript, respectively. (XLS 313 kb)
Supplementary Table 10
Statistical analysis for TE methylation in ddm1 rdr6 replicates compared to ddm1. Exact binomial two-sided test given for transposon methylation in ddm1-2 rdr6-15 replicates as compared to ddm1-2. p-value < 2.2e-16 alternative hypothesis: true probability of success is not equal to 0.5, 95 % confidence interval, sample estimates: probability of success given. (XLS 45 kb)
Source data
Rights and permissions
About this article
Cite this article
Creasey, K., Zhai, J., Borges, F. et al. miRNAs trigger widespread epigenetically activated siRNAs from transposons in Arabidopsis. Nature 508, 411–415 (2014). https://doi.org/10.1038/nature13069
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13069
This article is cited by
-
Accumulation of DNA damage alters microRNA gene transcription in Arabidopsis thaliana
BMC Plant Biology (2022)
-
Plant gene silencing signals move from the phloem to influence gene expression in shoot apical meristems
BMC Plant Biology (2022)
-
Accumulation dynamics of ARGONAUTE proteins during meiosis in Arabidopsis
Plant Reproduction (2022)
-
Cultivar-specific miRNA-mediated RNA silencing in grapes
Planta (2022)
-
How do plants remember drought?
Planta (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.