Abstract
How specific features in the environment are represented within the brain is an important unanswered question in neuroscience. A subset of retinal neurons, called direction-selective ganglion cells (DSGCs), are specialized for detecting motion along specific axes of the visual field1. Despite extensive study of the retinal circuitry that endows DSGCs with their unique tuning properties2,3, their downstream circuitry in the brain and thus their contribution to visual processing has remained unclear. In mice, several different types of DSGCs connect to the dorsal lateral geniculate nucleus (dLGN)4,5,6, the visual thalamic structure that harbours cortical relay neurons. Whether direction-selective information computed at the level of the retina is routed to cortical circuits and integrated with other visual channels, however, is unknown. Here we show that there is a di-synaptic circuit linking DSGCs with the superficial layers of the primary visual cortex (V1) by using viral trans-synaptic circuit mapping7,8 and functional imaging of visually driven calcium signals in thalamocortical axons. This circuit pools information from several types of DSGCs, converges in a specialized subdivision of the dLGN, and delivers direction-tuned and orientation-tuned signals to superficial V1. Notably, this circuit is anatomically segregated from the retino-geniculo-cortical pathway carrying non-direction-tuned visual information to deeper layers of V1, such as layer 4. Thus, the mouse harbours several functionally specialized, parallel retino-geniculo-cortical pathways, one of which originates with retinal DSGCs and delivers direction- and orientation-tuned information specifically to the superficial layers of the primary visual cortex. These data provide evidence that direction and orientation selectivity of some V1 neurons may be influenced by the activation of DSGCs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
19 March 2014
Minor edits were made to the numbering of the affiliations list, and minor typographical edits were made to the legends of Figs 3 and 4.
References
Barlow, H. B. & Hill, R. Selective sensitivity to direction of movement in ganglion cells of the rabbit retina. Science 139, 412–414 (1963)
Briggman, K. L., Helmstaedter, M. & Denk, W. Wiring specificity in the direction-selectivity circuit of the retina. Nature 471, 183–188 (2011)
Wei, W. & Feller, M. B. Organization and development of direction-selective circuits in the retina. Trends Neurosci. 34, 638–645 (2011)
Kim, I. J., Zhang, Y., Yamagata, M., Meister, M. & Sanes, J. R. Molecular identification of a retinal cell type that responds to upward motion. Nature 452, 478–482 (2008)
Huberman, A. D. et al. Genetic identification of an On-Off direction-selective retinal ganglion cell subtype reveals a layer-specific subcortical map of posterior motion. Neuron 62, 327–334 (2009)
Rivlin-Etzion, M. et al. Transgenic mice reveal unexpected diversity of on-off direction-selective retinal ganglion cell subtypes and brain structures involved in motion processing. J. Neurosci. 31, 8760–8769 (2011)
Wickersham, I. R. et al. Monosynaptic restriction of transsynaptic tracing from single, genetically targeted neurons. Neuron 53, 639–647 (2007b)
Osakada, F. et al. New rabies virus variants for monitoring and manipulating activity and gene expression in defined neural circuits. Neuron 71, 617–631 (2011)
Chalupa, L. M. & Werner, J. S. The Visual Neurosciences (MIT Press, 2003)
Krahe, T. E., El-Danaf, R. N., Dilger, E. K., Henderson, S. C. & Guido, W. Morphologically distinct classes of relay cells exhibit regional preferences in the dorsal lateral geniculate nucleus of the mouse. J. Neurosci. 31, 17437–17448 (2011)
Grubb, M. S. & Thompson, I. D. Biochemical and anatomical subdivision of the dorsal lateral geniculate nucleus in normal mice and in mice lacking the β2 subunit of the nicotinic acetylcholine receptor. Vision Res. 44, 3365–3376 (2004)
Morin, L. P. & Blanchard, J. H. Forebrain connections of the hamster intergeniculate leaflet: comparison with those of ventral lateral geniculate nucleus and retina. Vis. Neurosci. 16, 1037–1054 (1999)
Rafols, J. A. & Valverde, F. The structure of the dorsal lateral geniculate nucleus of the mouse. A Golgi and electron microscopic study. J. Comp. Neurol. 150, 303–331 (1973)
Land, P. W., Kyonka, E. & Shamalla-Hannah, L. Vesicular glutamate transporters in the lateral geniculate nucleus: expression of VGLUT2 by retinal terminals. Brain Res. 996, 251–254 (2004)
Kay, J. N. et al. Retinal ganglion cells with distinct directional preferences differ in molecular identity, structure, and central projections. J. Neurosci. 31, 7753–7762 (2011)
Dhande, O. S. et al. Genetic dissection of retinal inputs to brainstem nuclei controlling image stabilization. J. Neurosci. 33, 17797–17813 (2013)
Chen, C. & Regehr, W. G. Developmental remodeling of the retinogeniculate synapse. Neuron 28, 955–966 (2000)
Huberman, A. D. et al. Architecture and activity-mediated refinement of axonal projections from a mosaic of genetically identified retinal ganglion cells. Neuron 59, 425–438 (2008)
Völgyi, B., Abrams, J., Paul, D. L. & Bloomfield, S. A. Morphology and tracer coupling patterns of alpha ganglion cells in the mouse retina. J. Comp. Neurol. 492, 66–77 (2005)
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013)
Rivlin-Etzion, M., Wei, W. & Feller, M. B. Visual stimulation reverses the directional preference of direction-selective retinal ganglion cells. Neuron 76, 518–525 (2012)
Li, Y. T., Ibrahim, L. A., Liu, B. H., Zhang, L. I. & Tao, H. W. Linear transformation of thalamocortical input by intracortical excitation. Nature Neurosci. 16, 1324–1330 (2013)
Lien, A. D. & Scanziani, M. Tuned thalamic excitation is amplified by visual cortical circuits. Nature Neurosci. 16, 1315–1323 (2013)
Marshel, J. H., Kaye, A. P., Nauhaus, I. & Callaway, E. M. Anterior-posterior direction opponency in the superficial mouse lateral geniculate nucleus. Neuron 76, 713–720 (2012)
Piscopo, D. M., El-Danaf, R. N., Huberman, A. D. & Niell, C. M. Diverse visual features encoded in mouse lateral geniculate nucleus. J. Neurosci. 33, 4642–4656 (2013)
Livingstone, M. S. Mechanisms of direction selectivity in macaque V1. Neuron 20, 509–526 (1998)
Atallah, B. V., Bruns, W., Carandini, M. & Scanziani, M. Parvalbumin-expressing interneurons linearly transform cortical responses to visual stimuli. Neuron 73, 159–170 (2012)
Lee, S. H. et al. Activation of specific interneurons improves feature selectivity and visual perception. Nature 488, 379–383 (2012)
Katz, L. C., Burkhalter, A. & Dreyer, W. J. Fluorescent latex microspheres as a retrograde neuronal marker for in vivo and in vitro studies of visual cortex. Nature 310, 498–500 (1984)
Hioki, H. et al. Efficient gene transduction of neurons by lentivirus with enhanced neuron-specific promoters. Gene Ther. 14, 872–882 (2007)
Osakada, F. & Callaway, E. M. Design and generation of recombinant rabies virus vectors. Nature Protocols 8, 1583–1601 (2013)
Murphy, G. J. & Rieke, F. Network variability limits stimulus-evoked spike timing precision in retinal ganglion cells. Neuron 52, 511–524 (2006)
Beier, K. T. et al. Transsynaptic tracing with vesicular stomatitis virus reveals novel retinal circuitry. J. Neurosci. 33, 35–51 (2013)
Zack, G. W., Rogers, W. E. & Latt, S. A. Automatic measurement of sister chromatid exchange frequency. J. Histochem. Cytochem. 25, 741–753 (1977)
Pologruto, T. A., Sabatini, B. L. & Svoboda, K. ScanImage: flexible software for operating laser scanning microscopes. Biomed. Eng. Online 2, 13 (2003)
Tian, L. et al. Imaging neural activity in worms, flies and mice with improved GCaMP indicators. Nature Methods 6, 875–881 (2009)
Brainard, D. H. The psychophysics toolbox. Spat. Vis. 10, 433–436 (1997)
Pelli, D. G. The VideoToolbox software for visual psychophysics: transforming numbers into movies. Spat. Vis. 10, 437–442 (1997)
Bokil, H., Andrews, P., Kulkarni, J. E., Mehta, S. & Mitra, P. P. Chronux: a platform for analyzing neural signals. J. Neurosci. Methods 192, 146–151 (2010)
Acknowledgements
We thank the Kleinfeld laboratory for helpful advice, F. Rieke and M. Turner for example DSGC recording, and the Salk Viral Vector and Biophotonics staff. This work was supported by Vision Core P30 EY019005, the Knights Templar Eye Foundation (O.S.D.), Japan Society for the Promotion of Science (F.O.), Kanae Foundation (F.O.), Uehara Memorial Foundation (F.O.), Naito Foundation (F.O.), NINDS Circuits Training Grant (R.N.E.), Gatsby Charitable Trusts (E.M.C. and A.G.), NIH EY022577 and MH063912 (E.M.C.), Whitehall Foundation (A.D.H.), Ziegler Foundation for the Blind (A.D.H.), Pew Charitable Trusts (A.D.H.), The McKnight Foundation (A.D.H.), and NIH R01EY022157 (A.D.H.).
Author information
Authors and Affiliations
Contributions
A.D.H., A.C.-M., A.G. and R.N.E. designed the experiments. A.D.H., A.C.-M., R.N.E. and P.L.N. carried out and analysed the circuit connectivity experiments. A.C.-M. carried out the in vivo imaging experiments. B.S. and A.C.-M. analysed imaging data. O.S.D. collected data on molecular markers of cell types. E.M.C. and F.O. designed and made the rabies viruses. A.D.H. and A.C.-M. wrote the paper in collaboration with the other authors. A.D.H. and A.C.-M. prepared the figures. A.D.H. oversaw the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 The retino-geniculo-cortical pathway links retinal cells and circuits to the brain.
a, Diagram of retina, dorsal lateral geniculate nucleus (dLGN) and primary visual cortex (V1). The optic tract which carries retinal ganglion cell (RGC) axons and thalamocortical (dLGN to V1) pathway also shown. b, Diagram of retinal layers: PRL, photoreceptor layer; opl, outer plexiform layer; INL, inner nuclear layer; ipl, inner plexiform layer; GCL, ganglion cell layer; nfl, nerve fibre layer. c, Retina diagram with cells shown (labels same as in b).
Extended Data Figure 2 Approach for assessing laminar specificity of mouse geniculocortical projections.
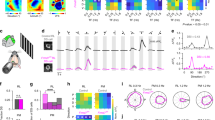
a, Focal retrograde tracer injection to V1. Scale bar, 3 mm. b, Diagram of the three different injection depths used to generate data in Fig. 2. c, Percentage of fluorescence in V1 from superficial (black line) versus deep (grey line) injections. Superficial, peak intensity occurs at 25 μm from pial surface (4 mice). Deep, peak intensity occurs at 350 μm from pial surface. Gray shaded regions, s.e.m. (superficial vs deep = ***P < 0.0001; two-way ANOVA). d, Assessment of retrogradely labelled cells across the width of the dLGN. 0% is at optic tract, 100% is at medial border (see Fig. 2g–i).
Extended Data Figure 3 Retrograde tracers to superficial V1 label cells in the DSGC-RZ.
a–c, Same dLGN as in main Fig. 2f but with GFP+ On-Off DSGC6 axons shown. a, most of the retrogradely labelled cells (magenta/dashed circles) reside in the DSGC-RZ (green terminals). Asterisk, labelled cell outside the DSGC-RZ. Scale bar, 200 μm. b, c, High magnification views of retrogradely labelled dLGN neuron cell bodies with potential contact from GFP+ DSGC axons (arrow in b); c, this cell is in vicinity of DSGC axonal boutons (arrowheads). b, c, Scale, 15 μm. d, Diagram of laminar-specific connections between DSGC-RZ and superficial V1 and dLGN core and deeper V1 layers 4 and 6.
Extended Data Figure 4 Analysis of dLGN neurons retrogradely infected from superficial V1.
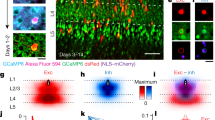
a–f, Example serial sections of anterior, middle and posterior portions of dLGN in a mouse with GFP expressing On-Off DSGC axons that was injected with ΔG-RABV-mCherry in superficial layers of V1. a, DAPI to show cytoarchitectural landmarks and dLGN borders. b, GFP+ DSGC axons and AAV2-Glyco-hGFP-infected cell bodies (see main Fig. 4 and text). c, Mask of GFP+ DSGC axons (Methods). d, ΔG-RABV-mCherry+ dLGN relay neurons. e, GFP+ DSGC axon mask superimposed with mCherry signal; this was used to determine colocalization. f, mCherry and GFP signals merged. Scale bar, 200 μm.
Extended Data Figure 5 Putative sites of contact between DSGC axons and a dLGN neuron retrogradely infected from superficial V1.
a–i, GFP+ On-Off DSGC axons (green in all panels except black in b) and mCherry+ dLGN relay neuron (magenta in all panels except white in c) infected by injection to superficial V1. Framed region in a is shown at higher magnification in b–d. Arrowhead (a), thalamocortical axon of mCherry+ dLGN cell. Scale bar in a, 50 μm. Yellow boxed region in c, d, is shown at higher magnification in e–i. Scale bar in d, 15 μm. e–i, Some DSGC axon–dendrite contacts contain VGLUT2 (blue). f–i, Arrowhead, site of GFP/mCherry co-localization that does not contain VGLUT2; arrow, GFP/mCherry/VGLUT2+ contact.
Extended Data Figure 6 The axons of GFP+ On-Off DSGCs and dLGN neurons infected with AAV2-Glyco-hGFP can be distinguished on the basis of their cellular localization.
High magnification view of DSGC-RZ in mouse with GFP+ posterior-tuned On-Off DSGCs that was injected 14 days earlier with AAV2-Glyco-hGFP. Glyco-hGFP+ neurons have nuclear GFP labelling (arrows), whereas DSGCs have GFP in axon terminals (arrowheads). Dashed line, lateral border of dLGN. OT, optic tract. Scale bar, 50 μm.
Extended Data Figure 7 Signature anatomical and physiological characteristics of GFP-tagged On-Off DSGCs.
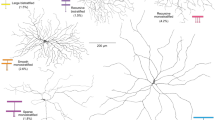
a, b, Flat-mount retina with GFP+ On-Off DSGCs (a) and co-stained with DAPI (b). c, Positions of GFP+ RGCs. Scale bar in c, 150 μm. d–f, High magnification views. Scale bar, 12 μm. g, Targeted fill of a GFP+ DSGC. Scale bar, 50 μm. h, Schematic of On-Off DSGC stratification and starburst amacrine cells (magenta). Labelling as in Extended Data Fig. 1. i, j, Higher magnification of framed region in g stained for VAChT (starburst amacrine processes). Asterisk, ‘looping arborizations’; dashed line, GFP arborization, which matches VAChT plexus. Scale bar, 10 μm. k, l, Side (x–z plane) views of cell in g. GFP+ dendrites co-stratify with both the On and Off sublayers. Scale bar, 5 μm. m, Direction-tuned response of a GFP+ On-Off DSGC targeted for recording and receptive field characterization. The spike count is highest for bars moving towards ∼270° in the cardinal axes.
Extended Data Figure 8 Injections of ΔG-RABV-mCherry into both superficial and deep V1 combined with AAV2-Glyco-hGFP infection of dLGN core.
a, mCherry+ neurons in the DSGC-RZ and the core of the dLGN. b, AAV2-Glyco-hGFP: many neurons throughout the dLGN, but mostly along the medial border and not in the shell/DSGC-RZ express Glyco-hGFP. DSGC-RZ marked by axons of GFP+ On-Off DSGCs. c, Merged of a, b. Scale in a, 100 μm. Boxed regions with arrows: two dLGN neurons; both RABV-mCherry+ and AAV2-Glyco-hGFP+. One or both of these cells infected their presynaptic partner, the RGC shown in Fig. 4 (panels cc-ee) of the main text. Scale bar, 15μm.
Rights and permissions
About this article
Cite this article
Cruz-Martín, A., El-Danaf, R., Osakada, F. et al. A dedicated circuit links direction-selective retinal ganglion cells to the primary visual cortex. Nature 507, 358–361 (2014). https://doi.org/10.1038/nature12989
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12989
This article is cited by
-
Targeted visual cortex stimulation (TVCS): a novel neuro-navigated repetitive transcranial magnetic stimulation mode for improving cognitive function in bipolar disorder
Translational Psychiatry (2023)
-
Developmental loss of ErbB4 in PV interneurons disrupts state-dependent cortical circuit dynamics
Molecular Psychiatry (2023)
-
Thalamic subnetworks as units of function
Nature Neuroscience (2022)
-
Haploinsufficiency of Shank3 increases the orientation selectivity of V1 neurons
Scientific Reports (2022)
-
Untangling the cortico-thalamo-cortical loop: cellular pieces of a knotty circuit puzzle
Nature Reviews Neuroscience (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.