Abstract
Here we report a unique cellular reprogramming phenomenon, called stimulus-triggered acquisition of pluripotency (STAP), which requires neither nuclear transfer nor the introduction of transcription factors. In STAP, strong external stimuli such as a transient low-pH stressor reprogrammed mammalian somatic cells, resulting in the generation of pluripotent cells. Through real-time imaging of STAP cells derived from purified lymphocytes, as well as gene rearrangement analysis, we found that committed somatic cells give rise to STAP cells by reprogramming rather than selection. STAP cells showed a substantial decrease in DNA methylation in the regulatory regions of pluripotency marker genes. Blastocyst injection showed that STAP cells efficiently contribute to chimaeric embryos and to offspring via germline transmission. We also demonstrate the derivation of robustly expandable pluripotent cell lines from STAP cells. Thus, our findings indicate that epigenetic fate determination of mammalian cells can be markedly converted in a context-dependent manner by strong environmental cues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
02 July 2014
Several critical errors have been found in our Article and Letter, which led to an in-depth investigation by the RIKEN Institute. The RIKEN investigation committee has categorized some of the errors as misconduct (see Supplementary Data 1 and Supplementary Data 2).
References
Gurdon, J. B. The developmental capacity of nuclei taken from intestinal epithelium cells of feeding tadpoles. J. Embryol. Exp. Morphol.10, 622–640 (1962)
Wakayama, T., Perry, A. C., Zuccotti, M., Johnson, K. R. & Yanagimachi, R. Full-term development of mice from enucleated oocytes injected with cumulus cell nuclei. Nature394, 369–374 (1998)
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell126, 663–676 (2006)
Thorpe, T. A. History of plant tissue culture. Mol. Biotechnol.37, 169–180 (2007)
Jiang, Y. et al. Pluripotency of mesenchymal stem cells derived from adult marrow. Nature418, 41–49 (2002)
D’Ippolito, G. et al. Marrow-isolated adult multilineage inducible (MIAMI) cells, a unique population of postnatal young and old human cells with extensive expansion and differentiation potential. J. Cell Sci.117, 2971–2981 (2004)
Kucia, M. et al. A population of very small embryonic-like (VSEL) CXCR4+SSEA-1+Oct-4+ stem cells identified in adult bone marrow. Leukemia20, 857–869 (2006)
Kuroda, Y. et al. Unique multipotent cells in adult human mesenchymal cell populations. Proc. Natl Acad. Sci. USA107, 8639–8643 (2010)
Obokata, H. et al. The potential of stem cells in adult tissues representative of the three germ layers. Tissue Eng. Part A17, 607–615 (2011)
Lengner, C. J., Welstead, G. G. & Jaenisch, R. The pluripotency regulator Oct4: a role in somatic stem cells? Cell Cycle7, 725–728 (2008)
Berg, J. S. & Goodell, M. A. An argument against a role for Oct4 in somatic stem cells. Cell Stem Cell1, 359–360 (2007)
Hong, H. et al. Suppression of induced pluripotent stem cell generation by the p53-p21 pathway. Nature460, 1132–1135 (2009)
Hanna, J. et al. Direct reprogramming of terminally differentiated mature B lymphocytes to pluripotency. Cell133, 250–264 (2008)
Ohbo, K. et al. Identification and characterization of stem cells in prepubertal spermatogenesis in mice small star, filled. Dev. Biol.258, 209–225 (2003)
Holtfreter, J. Neural induction in explants which have passed through a sublethal cytolysis. J. Exp. Zool.106, 197–222 (1947)
Gerhart, J. Johannes Holtfreter’s contributions to ongoing studies of the organizer. Dev. Dyn.205, 245–256 (1996)
Byrnes, W. M. Ernest Everett Just, Johannes Holtfreter, and the origin of certain concepts in embryo morphogenesis. Mol. Reprod. Dev.76, 912–921 (2009)
Gratama, J. W., Sutherland, D. R. & Keeney, M. Flow cytometric enumeration and immunophenotyping of hematopoietic stem and progenitor cells. Semin. Hematol.38, 139–147 (2001)
Inoue, K. et al. Inefficient reprogramming of the hematopoietic stem cell genome following nuclear transfer. J. Cell Sci.119, 1985–1991 (2006)
Ying, Q. L. et al. The ground state of embryonic stem cell self-renewal. Nature453, 519–523 (2008)
Tesar, P. J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature448, 196–199 (2007)
Brons, I. G. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature448, 191–195 (2007)
Kuroda, Y. et al. Isolation, culture and evaluation of multilineage-differentiating stress-enduring (Muse) cells. Nature Protocols8, 1391–1415 (2013)
Niwa, H. How is pluripotency determined and maintained? Development134, 635–646 (2007)
Watanabe, K. et al. Directed differentiation of telencephalic precursors from embryonic stem cells. Nature Neurosci.8, 288–296 (2005)
Kawasaki, H. et al. Induction of midbrain dopaminergic neurons from ES cells by stromal cell-derived inducing activity. Neuron28, 31–40 (2000)
Gouon-Evans, V. et al. BMP-4 is required for hepatic specification of mouse embryonic stem cell-derived definitive endoderm. Nature Biotechnol.24, 1402–1411 (2006)
Ohgushi, M. & Sasai, Y. Lonely death dance of human pluripotent stem cells: ROCKing between metastable cell states. Trends Cell Biol.21, 274–282 (2011)
Ohgushi, M. et al. Molecular pathway and cell state responsible for dissociation-induced apoptosis in human embryonic stem cells. Cell Stem Cell7, 225–239 (2010)
Murakami, K., Araki, K., Ohtsuka, S., Wakayama, T. & Niwa, H. Choice of random rather than imprinted X inactivation in female embryonic stem cell-derived extra-embryonic cells. Development138, 197–202 (2011)
Pera, M. F. & Tam, P. P. Extrinsic regulation of pluripotent stem cells. Nature465, 713–720 (2010)
Surani, M. A. & Barton, S. C. Development of gynogenetic eggs in the mouse: implications for parthenogenetic embryos. Science222, 1034–1036 (1983)
Wakayama, S. et al. Successful serial recloning in the mouse over multiple generations. Cell Stem Cell12, 293–297 (2013)
Nagy, A., Rossant, J., Nagy, R., Abramow-Newerly, W. & Roder, J. C. Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc. Natl Acad. Sci. USA90, 8424–8428 (1993)
Eakin, G. S., Hadjantonakis, A. K., Papaioannou, V. E. & Behringer, R. R. Developmental potential and behavior of tetraploid cells in the mouse embryo. Dev. Biol.288, 150–159 (2005)
Ogawa, K., Matsui, H., Ohtsuka, S. & Niwa, H. A novel mechanism for regulating clonal propagation of mouse ES cells. Genes Cells9, 471–477 (2004)
Hurtado, C. & De Robertis, E. M. Neural induction in the absence of organizer in salamander is mediated by MAPK. Dev. Biol.307, 282–289 (2007)
Hou, P. et al. Pluripotent stem cells induced from mouse somatic cells by small-molecule compounds. Science341, 651–654 (2013)
Eiraku, M. et al. Self-organizing optic-cup morphogenesis in three-dimensional culture. Nature472, 51–56 (2011)
Eilken, M. H., Nishikawa, S. & Schroeder, T. Continuous single-cell imaging of blood generation from haemogenic endothelium. Nature457, 896–900 (2009)
Jentsch, I., Geigl, J., Klein, C. A. & Speicher, M. R. Seven-fluorochrome mouse M-FISH for high-resolution analysis of interchromosomal rearrangements. Cytogenet. Genome Res.103, 84–88 (2003)
Acknowledgements
We thank S. Nishikawa for discussion and J. D. Ross, N. Takata, M. Eiraku, M. Ohgushi, S. Itoh, S. Yonemura, S. Ohtsuka and K. Kakiguchi for help with experiments and analyses. We thank A. Penvose and K. Westerman for comments on the manuscript. H.O. is grateful to T. Okano, S. Tsuneda and K. Kuroda for support and encouragement. Financial support for this research was provided by Intramural RIKEN Research Budget (H.O., T.W. and Y.S.), a Scientific Research in Priority Areas (20062015) to T.W., the Network Project for Realization of Regenerative Medicine to Y.S., and Department of Anesthesiology, Perioperative and Pain Medicine at Brigham and Women’s Hospital to C.A.V.
Author information
Authors and Affiliations
Contributions
H.O. and Y.S. wrote the manuscript. H.O., T.W. and Y.S. performed experiments, and K.K. assisted with H.O.’s transplantation experiments. H.O., T.W., Y.S., H.N. and C.A.V. designed the project. M.P.V. and M.Y. helped with the design and evaluation of the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Several critical errors have been found in our Article and Letter, which led to an in-depth investigation by the RIKEN Institute. The RIKEN investigation committee has categorized some of the errors as misconduct (see Supplementary Data 1 and Supplementary Data 2). Additional errors identified by the authors that are not discussed in RIKEN’s report are listed below.
Extended data figures and tables
Extended Data Figure 1 Conversion of haematopoietic cells into Oct4-GFP+ cells by a low-pH exposure.
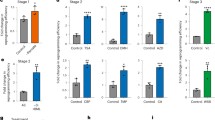
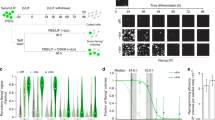
a, Optimization of pH conditions for Oct4-GFP induction. Five days after CD45-positive cells were exposed to acidic solution treatment at different pH, Oct4-GFP expression was analysed by FACS (n = 3, average?±?s.d.). b, Gating strategy for Oct4-GFP+ cell sorting. Top: representative results 7?days after the stress treatment. Bottom: non-treated control. P3 populations were sorted and counted as Oct4-GFP+ cells for all experiments. c, Controls for FACS analysis. In Oct4-GFP+ cell analysis, the grey and white histograms indicate the negative control (non-stress-treated Oct4-gfp haematopoietic cells) and the positive control (Oct4-gfp ES cells), respectively. Also, the green histograms indicate non-treated cells (left) and stress-treated cells at day?7 (right). In CD45+ cell analysis, the grey and white histograms indicate the negative (isotype) and positive controls, respectively. The red histograms indicate non-stress-treated cells (left) and stress-treated cells at day?7 (right). d, Oct4-GFP+ cell generation from various subpopulations of CD45+ cells. Seven days after the stress treatment, Oct4-GFP expression was analysed by FACS (n = 3, average?±?s.d.). Among total CD45+ fraction and its subfractions of CD19+, CD90+, CD34+ and CD34− cells, the efficacy of CD34+ cells was significantly lower than the others. P?<?0.05 by the Newman–Keuls test and P?<?0.01 by one-way ANOVA. e, Comparison of culture conditions for low-pH-induced conversion. Stress-treated cells were cultured in various media. The number of Oct4-GFP-expressing clusters was counted at day 14 (n = 3, average?±?s.d.). ***P?<?0.001 (B27+LIF versus all other groups); Tukey’s test. In the case of 3i medium, although the clusters appeared at a moderate efficiency, they appeared late and grew slowly. ACTH, ACTH-containing ES medium; ES+LIF+FBS, 20% FBS+LIF-containing ES culture medium; B27, DMEM/F12 medium containing 2% B27; B27+LIF, DMEM/F12 medium containing 2% B27+LIF; EpiSC, EpiSC culture medium containing Fgf2+activin. f, Signalling factor dependency of STAP cell generation. Growth factors that are conventionally used for pluripotent cell culture such as LIF, activin, Bmp4 and Fgf2 were added to basal culture medium (B27-supplemented DMEM/F12) in different culture phases (days 0–7, 2–7 and 4–7), and Oct4-GFP expression was analysed by FACS at day 7 (n = 3, average?±?s.d.). g, h, Time course of apoptosis after the low-pH exposure. Stress-treated cells and non-stress-treated control cells were stained with CD45, annexin-V and propidium iodide at day 0 (immediately after stress treatment), day 3 and day 7. g, Blue bars, GFP+CD45−; orange bars, GFP−CD45+. Percentages in total cells included propidium-iodide-positive cells. h, Annexin-V-positive cells in these cell populations were analysed by FACS.
Extended Data Figure 2 Phenotypic change during STAP cell conversion.
a, Oct4 protein expression in STAP cells was detected by immunostaining at day 2 (left) and day 7 (right). b, Live cell imaging of STAP conversion (grey, CD45 antibody; green, Oct4-GFP). See Methods for experimental details to monitor live CD45 immunostaining. c, Immunostaining of a parental CD45+ cell (left) and an Oct4-GFP+ cell (right). Scale bar, 10?μm. d, EdU uptake assay (n = 3, average?±?s.d.). e, Schematic of Tcrb gene rearrangement. f, T-cell-derived STAP cells. Scale bar, 100?μm. g, Genomic PCR analysis of (D)J recombination at the Tcrb gene of T-cell-derived STAP cells. G.L. is the size of the non-rearranged germline type, whereas the smaller ladders correspond to the alternative rearrangements of J exons (confirmed by sequencing). Negative controls (ES cells), positive controls (lymphocytes) and T-cell-derived STAP (two independent preparations on d7) are indicated.
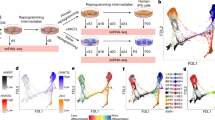
Extended Data Figure 3 Gene expression analyses during STAP conversion and endoderm differentiation assay.
a, Expression of pluripotency marker genes in STAP cells derived from T cells (n = 3, average?±?s.d.). b, Expression of pluripotency marker genes in STAP cells. In this experiment, Oct4-GFP+ cells seen in live cell imaging (Extended Data Fig. 2b) were analysed to confirm their conversion into STAP cells (n = 3, average?±?s.d.). c, Haematopoietic marker expression during STAP conversion from CD45+ cells (n = 3, average?±?s.d.). d, Formation of visceral endoderm-like surface epithelium in differentiating STAP cluster on day 2 (left) and day 8 (right). Scale bars, 50?μm.
Extended Data Figure 4 Teratoma formation assay and characterization of Oct4-GFP-dim cells.
a–c, Teratomas formed from STAP cell clusters included neuroepithelium (a), striated muscle (b) and pancreas (c; right, high-magnification view showing a typical acinar morphology and ductal structures). Scale bars, 100?μm. d, Teratoma-forming ability of Oct4-GFP+ and Oct4-GFP-dim cells (isolated by FACS, top). Oct4-GFP+ cells, but not Oct4-GFP-dim cells, efficiently formed teratomas (table at the bottom). However, because STAP cells were dissociation-intolerant, the teratoma-forming efficiency of dissociated Oct4-GFP+ cells was lower than that of non-dissociated STAP cell clusters. e, Gene expression of Oct4-GFP+ and Oct4-GFP-dim cells (n = 3, the average?±?s.d.). Haematopoietic marker gene expression (left) and early lineage marker gene expression (right) are shown.
Extended Data Figure 5 In vitro characterization of STAP cells.
a, Immunostaining for Ki67 and BrdU. STAP cell clusters (top) and ES cell colonies (bottom) are shown. For BrdU uptake, BrdU was added into each culture medium (10?μM) for 12?h until fixation. Scale bar, 100?μm. b, Transformation assay by soft agar culture. Neither Oct4-GFP+ nor Oct4-GFP-dim cells showed colony formation in soft agar, whereas ES cells and STAP stem cells showed anchorage-independent growth in the same LIF-B27 medium. Scale bar, 100?µm. Proliferated cells were lysed and the amount of DNA in each well was estimated by chemical luminescence (graph). n = 3 , average?±?s.d. c, No substantial change in chromosome number was seen with STAP cells in the CGH array. Genomic DNA derived from CD45+ cells (male) was used as reference DNA. The spikes (for example, those seen in the X chromosome) were nonspecific and also found in the data of these parental CD45+ cells when the manufacturer’s control DNA was used as a reference. d, qPCR analysis for pluripotency markers that highly express in ES cells, but not in EpiSCs. Average?±?s.d. e, Immunostaining of markers for mouse EpiSC and ES cells. Scale bar, 100 μm. f, g, H3K27me3+ foci in female cells, which are indicative of X-chromosomal inactivation. These foci were not observed in male cells. Scale bar, 10?μm. In the case of female STAP cells, ∼40% of cells retained H3K27me3+ foci (g). **P?<?0.001; Tukey’s test. n = 3, average?±?s.d. Although nuclear staining looked to be higher in STAP cells with H3K27me3+ foci (f), this appeared to be caused by some optical artefacts scattering from the strong focal signal. h, qPCR analysis for the tight junction markers Zo-1 and claudin 7, which were highly expressed in EpiSCs, but not in ES cells or STAP cells. **P?<?0.01; ns, not significant; Tukey's test; n = 3, average?±?s.d.
Extended Data Figure 6 Conversion of somatic tissue cells into STAP cells.
a, Alkaline phosphatase expression of STAP cells derived from adipose-derived mesenchymal cells. Scale bar, 100?μm. b, E-cadherin expression of STAP cells derived from adipose-derived mesenchymal cells. Scale bar, 50?μm. c, FACS sorting of dissociated neonatal cardiac muscle cells by removing CD45+ cells. d, Cardiomyocyte marker gene expression during STAP conversion from cardiomyocytes (n = 3, average?±?s.d.).
Extended Data Figure 7 Generation chimaeras with STAP cells.
a, 2N chimaeras generated with STAP cells derived from Oct4-gfp C57BL/6 mice (left) and 129/Sv?×?C57BL/6 F1 mice (right). b, Generation of chimaeric mice from STAP cells by cluster injection. STAP cells used in the experiments above were generated from CD45+ lymphocytes of multiple neonatal spleens (male and female tissues were mixed). *All fetuses were collected at 13.5?d.p.c. to 15.5?d.p.c. and the contribution rate of STAP cells into each organ was examined by FACS. **The contribution of STAP cells into each chimaera was scored as high (>50% of the coat colour of GFP expression). ***B6GFP: C57BL/6 mouse carrying cag-gfp.c, Production of offspring from STAP cells via germline transmission. Chimaeras generated with 129/Sv?×?B6GFP STAP cells (obtained from the experiments shown in b) were used for germline transmission study. d, 4N embryos at E9.5 generated with STAP cells derived from F1 GFP mice (B6GFP and DBA/2 or 129/Sv). B6GFP, C57BL/6 mouse carrying cag-gfp.
Extended Data Figure 8 Molecular and cellular characterization of STAP stem cells.
a, Compatibility of 2i conditions with STAP stem-cell derivation from STAP cells and STAP stem-cell maintenance. STAP stem cells could not be established directly from STAP cells in 2i + LIF medium (top). However, once established in ACTH medium, STAP stem cells were able to survive and expand in 2i + LIF medium. Scale bar, 100?μm. b, Q-band analysis (n = 4; all cell lines showed the normal karyotype). c, Multicolour FISH analysis (n = 8; all cell lines showed the normal karyotype) of STAP stem cells. d, Methylation status of the Oct4 and Nanog promoters. e, Electron microscope analysis of STAP stem cells. Scale bar, 1?μm. f, g, Beating cardiac muscle (mesoderm; 38%, n = 8). Red line indicates an analysed region for kymograph (g). h, Clonability of STAP stem cells. Clonal expansion from single STAP stem cells was performed. Pluripotency of clonal cell lines was confirmed by teratoma formation assay, showing the formation of neuroectoderm (left), muscle tissue (middle) and bronchial-like epithelium (right). Scale bar, 100?μm. i, Production of chimaeric mice from STAP stem-cell lines using diploid embryos. *These STAP stem-cell lines were generated from independent STAP cell clusters. j, Production of mouse chimaeras from STAP stem-cell lines by the tetraploid complementation method. *These STAP stem-cell lines were generated from independent STAP cell clusters. k, No H3K27me3-dense foci are seen in female STAP stem cells (n = 50; the CD45+ cell is a positive control). Scale bar, 10?μm.
Extended Data Figure 9 Effects of various stressors on STAP conversion.
a, Percentages of Oct4-GFP-expressing cells 7?days after stress treatment. Somatic cells were isolated from various tissues and exposed to different stressors. Oct4-GFP expression was analysed by FACS. b, Oct4 and Oct4-GFP expression induced in the reflux oesophagitis mouse model as an in vivo acid exposure model (top, experimental procedure). Oct4, but not Nanog, expression was observed in the oesophageal epithelium exposed to gastric acid (75% of 12 operated mice).
Supplementary information
Supplementary Information
This file contains Supplementary Table 1. (PDF 102 kb)
Live imaging of low-pH-treated CD45+cells
DIC images during day 0 – day 7, overlaid with oct3/4::GFP (green). A strong contrast of DIC (as compared to video 2) was applied to imaging so that lamellipodia-like processes (frequently seen on and after day 4) could be viewed easily. (MOV 23217 kb)
Live imaging of low-pH-treated CD45+cells (another view)
DIC images during day 0 – day 6, overlaid with oct3/4::GFP (green). The interval of imaging was half (15 min) of that of video 1 (the overall speed of the video is three-times slower than video 1). In this view field where the cell density was relatively low, behaviours of individual cells were more easily seen. In this case, forming clusters were slightly smaller in size. (MOV 11907 kb)
STAP cell-derived embryo (E10.5) from 4N blastocyst injection
STAP cells with constitutive GFP expression were injected into 4N blastocysts and produced normal embryos with heart beating. (MOV 1656 kb)
About this article
Cite this article
Obokata, H., Wakayama, T., Sasai, Y. et al. RETRACTED ARTICLE: Stimulus-triggered fate conversion of somatic cells into pluripotency. Nature 505, 641–647 (2014). https://doi.org/10.1038/nature12968
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12968
This article is cited by
-
Thinking in 3 dimensions: philosophies of the microenvironment in organoids and organs-on-chip
History and Philosophy of the Life Sciences (2023)
-
Postnatal Pluripotent Cells: Quarter of a Century of Research
Bulletin of Experimental Biology and Medicine (2021)
-
Double sperm cloning (DSC) is a promising strategy in mammalian genetic engineering and stem cell research
Stem Cell Research & Therapy (2020)
-
The limited application of stem cells in medicine: a review
Stem Cell Research & Therapy (2018)
-
Analysis of chromosomal aberrations and recombination by allelic bias in RNA-Seq
Nature Communications (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.