Abstract
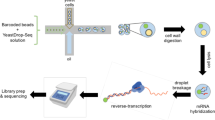
Variation among individuals arises in part from differences in DNA sequences, but the genetic basis for variation in most traits, including common diseases, remains only partly understood. Many DNA variants influence phenotypes by altering the expression level of one or several genes. The effects of such variants can be detected as expression quantitative trait loci (eQTL)1. Traditional eQTL mapping requires large-scale genotype and gene expression data for each individual in the study sample, which limits sample sizes to hundreds of individuals in both humans and model organisms and reduces statistical power2,3,4,5,6. Consequently, many eQTL are probably missed, especially those with smaller effects7. Furthermore, most studies use messenger RNA rather than protein abundance as the measure of gene expression. Studies that have used mass-spectrometry proteomics8,9,10,11,12,13 reported unexpected differences between eQTL and protein QTL (pQTL) for the same genes9,10, but these studies have been even more limited in scope. Here we introduce a powerful method for identifying genetic loci that influence protein expression in the yeast Saccharomyces cerevisiae. We measure single-cell protein abundance through the use of green fluorescent protein tags in very large populations of genetically variable cells, and use pooled sequencing to compare allele frequencies across the genome in thousands of individuals with high versus low protein abundance. We applied this method to 160 genes and detected many more loci per gene than previous studies. We also observed closer correspondence between loci that influence protein abundance and loci that influence mRNA abundance of a given gene. Most loci that we detected were clustered in ‘hotspots’ that influence multiple proteins, and some hotspots were found to influence more than half of the proteins that we examined. The variants that underlie these hotspots have profound effects on the gene regulatory network and provide insights into genetic variation in cell physiology between yeast strains.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
10 January 2014
A minor change was made to the opening paragraph.
References
Rockman, M. V. & Kruglyak, L. Genetics of global gene expression. Nature Rev. Genet. 7, 862–872 (2006)
Smith, E. N. & Kruglyak, L. Gene–environment interaction in yeast gene expression. PLoS Biol. 6, e83 (2008)
Rockman, M. V., Skrovanek, S. S. & Kruglyak, L. Selection at linked sites shapes heritable phenotypic variation in C. elegans. Science 330, 372–376 (2010)
Huang, G. J. et al. High resolution mapping of expression QTLs in heterogeneous stock mice in multiple tissues. Genome Res. 19, 1133–1140 (2009)
West, M. A. L. et al. Global eQTL mapping reveals the complex genetic architecture of transcript-level variation in Arabidopsis. Genetics 175, 1441–1450 (2007)
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511 (2013)
Brem, R. B. & Kruglyak, L. The landscape of genetic complexity across 5,700 gene expression traits in yeast. Proc. Natl Acad. Sci. USA 102, 1572–1577 (2005)
Foss, E. J. et al. Genetic basis of proteome variation in yeast. Nature Genet. 39, 1369–1375 (2007)
Foss, E. J. et al. Genetic variation shapes protein networks mainly through non-transcriptional mechanisms. PLoS Biol. 9, e1001144 (2011)
Ghazalpour, A. et al. Comparative analysis of proteome and transcriptome variation in mouse. PLoS Genet. 7, e1001393 (2011)
Wu, L. et al. Variation and genetic control of protein abundance in humans. Nature 499, 79–82 (2013)
Khan, Z., Bloom, J. S., Garcia, B. A., Singh, M. & Kruglyak, L. Protein quantification across hundreds of experimental conditions. Proc. Natl Acad. Sci. USA 106, 15544–15548 (2009)
Skelly, D. A. et al. Integrative phenomics reveals insight into the structure of phenotypic diversity in budding yeast. Genome Res. 23, 1496–1504 (2013)
Ehrenreich, I. M. et al. Dissection of genetically complex traits with extremely large pools of yeast segregants. Nature 464, 1039–1042 (2010)
Huh, W.-K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003)
Edwards, M. D. & Gifford, D. K. High-resolution genetic mapping with pooled sequencing. BMC Bioinformatics 13, S8 (2012)
Picotti, P. et al. A complete mass-spectrometric map of the yeast proteome applied to quantitative trait analysis. Nature 494, 266–270 (2013)
Brem, R. B., Yvert, G., Clinton, R. & Kruglyak, L. Genetic dissection of transcriptional regulation in budding yeast. Science 296, 752–755 (2002)
Litvin, O., Causton, H. C., Chen, B. J. & Pe’er, D. Modularity and interactions in the genetics of gene expression. Proc. Natl Acad. Sci. USA 106, 6441–6446 (2009)
Zitomer, R. S. & Lowry, C. V. Regulation of gene expression by oxygen in Saccharomyces cerevisiae. Microbiol. Rev. 56, 1–11 (1992)
Gaisne, M., Bécam, A. M., Verdiere, J. & Herbert, C. J. A. A ‘natural’ mutation in Saccharomyces cerevisiae strains derived from S288c affects the complex regulatory gene HAP1 (CYP1). Curr. Genet. 36, 195–200 (1999)
Harbison, C. T. et al. Transcriptional regulatory code of a eukaryotic genome. Nature 431, 99–104 (2004)
Butler, G. Hypoxia and gene expression in eukaryotic microbes. Annu. Rev. Microbiol. 67, 291–312 (2013)
Zaman, S., Lippman, S. I., Zhao, X. & Broach, J. R. How Saccharomyces responds to nutrients. Annu. Rev. Genet. 42, 27–81 (2008)
Zaman, S., Lippman, S. I., Schneper, L., Slonim, N. & Broach, J. R. Glucose regulates transcription in yeast through a network of signaling pathways. Mol. Syst. Biol. 5, 245 (2009)
Spor, A. et al. Niche-driven evolution of metabolic and life-history strategies in natural and domesticated populations of Saccharomyces cerevisiae. BMC Evol. Biol. 9, 296 (2009)
Warringer, J. et al. Trait variation in yeast is defined by population history. PLoS Genet. 7, e1002111 (2011)
Fraser, H. B., Moses, A. M. & Schadt, E. E. Evidence for widespread adaptive evolution of gene expression in budding yeast. Proc. Natl Acad. Sci. USA 107, 2977–2982 (2010)
Lewis, J. A. & Gasch, A. P. Natural variation in the yeast glucose-signaling network reveals a new role for the Mig3p transcription factor. G3 Gene Genomes Genetics 2, 1607–1612 (2012)
Henras, A. K. et al. The post-transcriptional steps of eukaryotic ribosome biogenesis. Cell. Mol. Life Sci. 65, 2334–2359 (2008)
Howson, R. et al. Construction, verification and experimental use of two epitope-tagged collections of budding yeast strains. Comp. Funct. Genomics 6, 2–16 (2005)
Tong, A. H. Y. & Boone, C. High-throughput strain construction and systematic synthetic lethal screening in Saccharomyces cerevisiae. Methods in Microbiology 36, 369–707 (2007)
Newman, J. R. S. et al. Single-cell proteomic analysis of S. cerevisiae reveals the architecture of biological noise. Nature 441, 840–846 (2006)
Adey, A. et al. Rapid, low-input, low-bias construction of shotgun fragment libraries by high-density in vitro transposition. Genome Biol. 11, R119 (2010)
Bloom, J. S., Ehrenreich, I. M., Loo, W. T., Lite, T.-L. V. & Kruglyak, L. Finding the sources of missing heritability in a yeast cross. Nature 494, 234–237 (2013)
Meyer, M. & Kircher, M. Illumina Sequencing Library Preparation for Highly Multiplexed Target Capture and Sequencing. Cold Spring Harbor Protocols http://dx.doi.org/10.1101/pdb.prot5448 (2010)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Broman, K. W., Wu, H., Sen, S. & Churchill, G. A. R/qtl: QTL mapping in experimental crosses. Bioinformatics 19, 889–890 (2003)
Yvert, G. et al. Trans-acting regulatory variation in Saccharomyces cerevisiae and the role of transcription factors. Nature Genet. 35, 57–64 (2003)
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003)
Spivak, A. T. & Stormo, G. D. ScerTF: a comprehensive database of benchmarked position weight matrices for Saccharomyces species. Nucleic Acids Res. 40, D162–D168 (2012)
Acknowledgements
We are grateful to C. DeCoste at the Princeton Flow Cytometry Resource Facility for technical assistance and advice on the experiments. This work was supported by National Institutes of Health (NIH) grant R01 GM102308, a James S. McDonnell Centennial Fellowship, and the Howard Hughes Medical Institute (L.K.), German Science Foundation research fellowship AL 1525/1-1 (F.W.A.), a National Science Foundation fellowship (J.S.B.), and NIH postdoctoral fellowship F32 GM101857-02 (S.T.).
Author information
Authors and Affiliations
Contributions
F.W.A. and L.K. conceived the project, designed research and wrote the paper. F.W.A. and A.H.S. performed experiments. F.W.A. analysed the data. S.T. provided advice on yeast strain construction, the initial experimental design and other experimental procedures. J.S.B. provided advice on experimental procedures and data analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 2 Illustration of FACS design.
Shown is GFP intensity and forward scatter (FSC, a measure of cell size) recorded during FACS. The correlation between cell size and GFP intensity is clearly visible. The superimposed collection gates are an illustration, and do not show the actual gates used for this gene. a, The low GFP (blue) and high GFP (red) gates sample extreme levels of GFP within a defined range of cell sizes. b, For the ‘null’ experiments, the same cell size range is collected, but without selecting on GFP.
Extended Data Figure 3 Sequence analyses and X-pQTL detection example.
In all panels, physical genomic coordinates are shown on the x-axes. The position of the gene (LEU1) is indicated by the purple vertical line. Top panel: frequency of the BY allele in the high (red) and low (blue) GFP population. SNPs are indicated by dots, and loess-smoothed averages as solid lines. Note the fixation of the BY allele in all segregants at the gene position and at the mating type locus on chromosome III, as well as the fixation of the RM allele at the synthetic genetic array marker integrated at the CAN1 locus on the left arm of chromosome V. Middle panel: subtraction of allele frequencies in the low from those in the high GFP population. SNPs are indicated by grey dots, with the loess-smoothed average indicated in black. Note that, on average, there is no difference between the high and the low populations. Positive difference values correspond to a higher frequency of the BY allele in the high GFP population, which we interpret as higher expression being caused by the BY allele at that locus. The red horizontal lines indicate the 99.99% quantile from the empirical ‘null’ sort experiments. They are shown for illustration only and were not used for peak calling. The blue vertical boxes indicate positions of genome-wide X-pQTL, with the width representing the 2-lod drop interval. Bottom panel: lod scores obtained from MULTIPOOL16. The red horizontal line is the genome-wide significance threshold (lod = 4.5). Stars indicate X-pQTL called by our algorithm; these positions correspond to the blue bars in the middle panel. For this gene, 14 X-pQTL are called.
Extended Data Figure 4 Reproducibility examples.
Shown are allele frequency differences between the high and low GFP populations along the genome of replicates for three genes. The gene positions are indicated by purple vertical lines; note that YMR315W and GCN1 were ‘local’ experiments where peaks at the gene position are visible. The red horizontal lines indicate the 99.99% quantile from the empirical ‘null’ sort experiments. Note the near-perfect agreement for strong X-pQTL, with some differences discernible at weaker loci. See Supplementary Note 1 for details.
Extended Data Figure 5 Example for a local X-pQTL in the gene MAE1.
Shown is the difference in the frequency of the BY allele between the high and the low GFP population along the genome. Red dashed horizontal lines indicate the 99.99% quantile from the empirical ‘null’ sort experiments. They are shown for illustration only and were not used for peak calling.
Extended Data Figure 6 Distributions of X-pQTL effect sizes for X-pQTL with and without a corresponding eQTL.
Effect sizes are shown as the absolute allele frequency differences between the high and low GFP population.
Extended Data Figure 7 The impact of small effect sizes on the π1 estimate.
Each panel shows the P-value distribution obtained from 5,000 tests of a given effect size x, if two groups of 50 individuals each are compared using a t-test. The effect size x is given along with the corresponding variance explained (VE), the π1 estimate, and the fraction of tests that achieved nominal significance (P < 0.05). Note that π1 reaches 0.3 at VE = 0.5% – 1% (middle row, right columns). See Supplementary Note 2 for details.
Extended Data Figure 8 Genes regulated by the hotspots on chromosomes XI, XII and XV.
The table shows genes that have an X-pQTL at three hotspots. For each gene involved in aerobic respiration, we show the X-pQTL lod scores along the genome in the top half of the plot, and the eQTL and pQTL lod scores in the bottom half on an inverted scale. The hotspot locations are shown as grey bars labelled with the names of the causative genes. Purple vertical lines indicate the gene positions. Red dashed horizontal lines are significance thresholds. Stars indicate significant QTL.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-2 and Supplementary Tables 1-3. (PDF 258 kb)
Supplementary Data 1
This file contains full details of the genes studied. (XLSX 67 kb)
Supplementary Data 2
This file contains a list of X-pQTL identified in this study. (XLSX 109 kb)
Supplementary Data 3
This zipped file contains allele count data used in the analyses. (ZIP 32911 kb)
Rights and permissions
About this article
Cite this article
Albert, F., Treusch, S., Shockley, A. et al. Genetics of single-cell protein abundance variation in large yeast populations. Nature 506, 494–497 (2014). https://doi.org/10.1038/nature12904
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12904
This article is cited by
-
Multi-omics analysis identifies drivers of protein phosphorylation
Genome Biology (2023)
-
Divergent patterns of selection on metabolite levels and gene expression
BMC Ecology and Evolution (2021)
-
Fast genetic mapping of complex traits in C. elegans using millions of individuals in bulk
Nature Communications (2019)
-
Integrated analysis of population genomics, transcriptomics and virulence provides novel insights into Streptococcus pyogenes pathogenesis
Nature Genetics (2019)
-
Post-translational buffering leads to convergent protein expression levels between primates
Genome Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.