Abstract
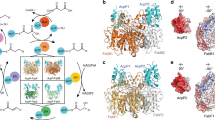
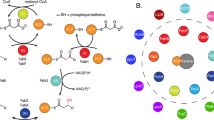
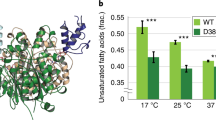
Acyl carrier protein (ACP) transports the growing fatty acid chain between enzymatic domains of fatty acid synthase (FAS) during biosynthesis1. Because FAS enzymes operate on ACP-bound acyl groups, ACP must stabilize and transport the growing lipid chain2. ACPs have a central role in transporting starting materials and intermediates throughout the fatty acid biosynthetic pathway3,4,5. The transient nature of ACP–enzyme interactions impose major obstacles to obtaining high-resolution structural information about fatty acid biosynthesis, and a new strategy is required to study protein–protein interactions effectively. Here we describe the application of a mechanism-based probe that allows active site-selective covalent crosslinking of AcpP to FabA, the Escherichia coli ACP and fatty acid 3-hydroxyacyl-ACP dehydratase, respectively. We report the 1.9 Å crystal structure of the crosslinked AcpP–FabA complex as a homodimer in which AcpP exhibits two different conformations, representing probable snapshots of ACP in action: the 4′-phosphopantetheine group of AcpP first binds an arginine-rich groove of FabA, then an AcpP helical conformational change locks AcpP and FabA in place. Residues at the interface of AcpP and FabA are identified and validated by solution nuclear magnetic resonance techniques, including chemical shift perturbations and residual dipolar coupling measurements. These not only support our interpretation of the crystal structures but also provide an animated view of ACP in action during fatty acid dehydration. These techniques, in combination with molecular dynamics simulations, show for the first time that FabA extrudes the sequestered acyl chain from the ACP binding pocket before dehydration by repositioning helix III. Extensive sequence conservation among carrier proteins suggests that the mechanistic insights gleaned from our studies may be broadly applicable to fatty acid, polyketide and non-ribosomal biosynthesis. Here the foundation is laid for defining the dynamic action of carrier-protein activity in primary and secondary metabolism, providing insight into pathways that can have major roles in the treatment of cancer, obesity and infectious disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chan, D. I. & Vogel, H. J. Current understanding of fatty acid biosynthesis and the acyl carrier protein. Biochem. J. 430, 1–19 (2010)
Rock, C. O. & Cronan, J. E., Jr Acyl carrier protein from Escherichia coli. Methods Enzymol. 71, 341–351 (1981)
Crosby, J. & Crump, M. P. The structural role of the carrier protein—active controller or passive carrier. Nat. Prod. Rep. 29, 1111–1137 (2012)
Magnuson, K., Jackowski, S., Rock, C. O. & Cronan, J. E., Jr Regulation of fatty acid biosynthesis in Escherichia coli. Microbiol. Rev. 57, 522–542 (1993)
Joshi, A. K., Witkowski, A., Berman, H. A., Zhang, L. & Smith, S. Effect of modification of the length and flexibility of the acyl carrier protein–thioesterase interdomain linker on functionality of the animal fatty acid synthase. Biochemistry 44, 4100–4107 (2005)
Issartel, J. P., Koronakis, V. & Hughes, C. Activation of Escherichia coli prohaemolysin to the mature toxin by acyl carrier protein-dependent fatty acylation. Nature 351, 759–761 (1991)
Agarwal, V., Lin, S., Lukk, T., Nair, S. K. & Cronan, J. E., Jr Structure of the enzyme-acyl carrier protein (ACP) substrate gatekeeper complex required for biotin synthesis. Proc. Natl Acad. Sci. USA 109, 17406–17411 (2012)
Anderson, M. S., Bulawa, C. E. & Raetz, C. R. The biosynthesis of gram-negative endotoxin. Formation of lipid A precursors from UDP-GlcNAc in extracts of Escherichia coli. J. Biol. Chem. 260, 15536–15541 (1985)
Jordan, S. W. & Cronan, J. E., Jr A new metabolic link. The acyl carrier protein of lipid synthesis donates lipoic acid to the pyruvate dehydrogenase complex in Escherichia coli and mitochondria. J. Biol. Chem. 272, 17903–17906 (1997)
Lu, Y. J. et al. Acyl-phosphates initiate membrane phospholipid synthesis in Gram-positive pathogens. Mol. Cell 23, 765–772 (2006)
Roujeinikova, A. et al. Crystallization and preliminary X-ray crystallographic studies on acyl-(acyl carrier protein) from Escherichia coli. Acta Crystallogr. D 58, 330–332 (2002)
Leibundgut, M., Jenni, S., Frick, C. & Ban, N. Structural basis for substrate delivery by acyl carrier protein in the yeast fatty acid synthase. Science 316, 288–290 (2007)
Roujeinikova, A. et al. Structural studies of fatty acyl-(acyl carrier protein) thioesters reveal a hydrophobic binding cavity that can expand to fit longer substrates. J. Mol. Biol. 365, 135–145 (2007)
Meier, J. L. & Burkart, M. D. The chemical biology of modular biosynthetic enzymes. Chem. Soc. Rev. 38, 2012–2045 (2009)
Ishikawa, F., Haushalter, R. W. & Burkart, M. D. Dehydratase-specific probes for fatty acid and polyketide synthases. J. Am. Chem. Soc. 134, 769–772 (2012)
Endo, K., Helmkamp, G. M., Jr & Bloch, K. Mode of inhibition of β-hydroxydecanoyl thioester dehydrase by 3-decynoyl-N-acetylcysteamine. J. Biol. Chem. 245, 4293–4296 (1970)
Leesong, M., Henderson, B. S., Gillig, J. R., Schwab, J. M. & Smith, J. L. Structure of a dehydratase–isomerase from the bacterial pathway for biosynthesis of unsaturated fatty acids: two catalytic activities in one active site. Structure 4, 253–264 (1996)
Zhuang, Z. et al. Divergence of function in the hot dog fold enzyme superfamily: the bacterial thioesterase YciA. Biochemistry 47, 2789–2796 (2008)
Moynié, L. et al. Structural insights into the mechanism and inhibition of the β-hydroxydecanoyl-acyl carrier protein dehydratase from Pseudomonas aeruginosa. J. Mol. Biol. 425, 365–377 (2013)
Guy, J. E. et al. Remote control of regioselectivity in acyl-acyl carrier protein-desaturases. Proc. Natl Acad. Sci. USA 108, 16594–16599 (2011)
Parris, K. D. et al. Crystal structures of substrate binding to Bacillus subtilis holo-(acyl carrier protein) synthase reveal a novel trimeric arrangement of molecules resulting in three active sites. Structure 8, 883–895 (2000)
Cryle, M. J. & Schlichting, I. Structural insights from a P450 Carrier Protein complex reveal how specificity is achieved in the P450(BioI) ACP complex. Proc. Natl Acad. Sci. USA 105, 15696–15701 (2008)
Babu, M. et al. Structure of a SLC26 anion transporter STAS domain in complex with acyl carrier protein: implications for E. coli YchM in fatty acid metabolism. Structure 18, 1450–1462 (2010)
Qiu, X. & Janson, C. A. Structure of apo acyl carrier protein and a proposal to engineer protein crystallization through metal ions. Acta Crystallogr. D 60, 1545–1554 (2004)
Salzmann, M., Pervushin, K., Wider, G., Senn, H. & Wuthrich, K. TROSY in triple-resonance experiments: new perspectives for sequential NMR assignment of large proteins. Proc. Natl Acad. Sci. USA 95, 13585–13590 (1998)
Hansen, M. R., Mueller, L. & Pardi, A. Tunable alignment of macromolecules by filamentous phage yields dipolar coupling interactions. Nature Struct. Biol. 5, 1065–1074 (1998)
Markwick, P. R. L. & McCammon, J. A. Studying functional dynamics in bio-molecules using accelerated molecular dynamics. Phys. Chem. Chem. Phys. 13, 20053–20065 (2011)
Markwick, P. R. et al. Toward a unified representation of protein structural dynamics in solution. J. Am. Chem. Soc. 131, 16968–16975 (2009)
Frueh, D. P. et al. Dynamic thiolation–thioesterase structure of a non-ribosomal peptide synthetase. Nature 454, 903–906 (2008)
Alekseyev, V. Y., Liu, C. W., Cane, D. E., Puglisi, J. D. & Khosla, C. Solution structure and proposed domain–domain recognition interface of an acyl carrier protein domain from a modular polyketide synthase. Protein Sci. 16, 2093–2107 (2007)
Acknowledgements
M.D.B. and S.-C.T. are supported by GM100305 and GM095970. We thank J. J. LaClair for figure editing. We thank X. Huang for assistance with NMR facilities and experimental setup. Portions of this research were carried out at the Stanford Synchrotron Radiation Lightsource (SSRL), a national user facility operated by Stanford University on behalf of the US Department of Energy, Office of Basic Energy Sciences. The Advanced Light Source is supported by the Office of Basic Energy Sciences of the US Department of Energy under contract no. DE-AC02-05CH11231. J.A.M. is supported by NSF, NIH and HHMI. Portions of the research are supported by the Advanced Light Source, supported by the Director, Office of Science, Office of Basic Energy Sciences, of the US Department of Energy under contract no. DE-AC02-05CH11231.
Author information
Authors and Affiliations
Contributions
C.N., assisted by G.C.-F., D.R.J. and J.B., determined the AcpP–FabA X-ray crystal structures. R.W.H., D.J.L. and B.O. conducted the protein NMR experiments under the supervision of S.J.O. F.I. and K.F. prepared the crosslinking probe. P.R.L.M. conducted molecular dynamics simulations under the supervision of J.A.M. C.N., G.C.-F., R.W.H. and D.J.L. analysed data and contributed to writing of the paper. S.-C.T. and M.D.B. directed the research, provided funding and wrote the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-18, Supplementary Methods, Supplementary Tables 1-9, a Supplementary Discussion and additional references. (PDF 15899 kb)
Rights and permissions
About this article
Cite this article
Nguyen, C., Haushalter, R., Lee, D. et al. Trapping the dynamic acyl carrier protein in fatty acid biosynthesis. Nature 505, 427–431 (2014). https://doi.org/10.1038/nature12810
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12810
This article is cited by
-
Triepoxide formation by a flavin-dependent monooxygenase in monensin biosynthesis
Nature Communications (2023)
-
Acyl carrier protein promotes MukBEF action in Escherichia coli chromosome organization-segregation
Nature Communications (2021)
-
Elucidation of transient protein-protein interactions within carrier protein-dependent biosynthesis
Communications Biology (2021)
-
Helicobacter pylori FabX contains a [4Fe-4S] cluster essential for unsaturated fatty acid synthesis
Nature Communications (2021)
-
Structural basis for selectivity in a highly reducing type II polyketide synthase
Nature Chemical Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.