Abstract
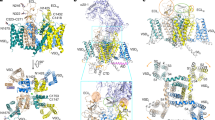
Voltage-gated calcium (CaV) channels catalyse rapid, highly selective influx of Ca2+ into cells despite a 70-fold higher extracellular concentration of Na+. How CaV channels solve this fundamental biophysical problem remains unclear. Here we report physiological and crystallographic analyses of a calcium selectivity filter constructed in the homotetrameric bacterial NaV channel NaVAb. Our results reveal interactions of hydrated Ca2+ with two high-affinity Ca2+-binding sites followed by a third lower-affinity site that would coordinate Ca2+ as it moves inward. At the selectivity filter entry, Site 1 is formed by four carboxyl side chains, which have a critical role in determining Ca2+ selectivity. Four carboxyls plus four backbone carbonyls form Site 2, which is targeted by the blocking cations Cd2+ and Mn2+, with single occupancy. The lower-affinity Site 3 is formed by four backbone carbonyls alone, which mediate exit into the central cavity. This pore architecture suggests a conduction pathway involving transitions between two main states with one or two hydrated Ca2+ ions bound in the selectivity filter and supports a ‘knock-off’ mechanism of ion permeation through a stepwise-binding process. The multi-ion selectivity filter of our CaVAb model establishes a structural framework for understanding the mechanisms of ion selectivity and conductance by vertebrate CaV channels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Accessions
Protein Data Bank
Data deposits
Coordinates and structure factors have been deposited in the Protein Data Bank under accession codes: 4MS2 (TLDDWSN, 15 mM Ca2+), 4MTF (TLDDWSN, 0.5 mM Ca2+), 4MTG (TLDDWSN, 2.5 mM Ca2+), 4MTO (TLDDWSN, 5 mM Ca2+), 4MVM (TLDDWSN, 10 mM Ca2+), 4MVO (TLDDWSN, 15 mM Ca2+), 4MVQ (TLDDWSD, 15 mM Ca2+), 4MVR (TLDDWSD, 100 mM Mn2+), 4MVS (TLDDWSD, 100 mM Cd2+), 4MVZ (TLEDWSD, 15 mM Ca2+), 4MW3 (TLDDWSM, 15 mM Ca2+), 4MVU (TLEDWSM, 15 mM Ca2+), 4MW8 (NavAb, 15 mM Ca2+).
References
Sather, W. A. & McCleskey, E. W. Permeation and selectivity in calcium channels. Annu. Rev. Physiol. 65, 133–159 (2003)
Catterall, W. A. Voltage-gated calcium channels. Cold Spring Harb. Perspect. Biol. 3, a003947 (2011)
Almers, W. & McCleskey, E. W. Non-selective conductance in calcium channels of frog muscle: calcium selectivity in a single-file pore. J. Physiol. (Lond.) 353, 585–608 (1984)
Almers, W., McCleskey, E. W. & Palade, P. T. A non-selective cation conductance in frog muscle membrane blocked by micromolar external calcium ions. J. Physiol. (Lond.) 353, 565–583 (1984)
Hess, P. & Tsien, R. W. Mechanism of ion permeation through calcium channels. Nature 309, 453–456 (1984)
Armstrong, C. M. & Neyton, J. Ion permeation through calcium channels. A one-site model. Ann. NY Acad. Sci. 635, 18–25 (1991)
Dang, T. X. & McCleskey, E. W. Ion channel selectivity through stepwise changes in binding affinity. J. Gen. Physiol. 111, 185–193 (1998)
Lopin, K. V., Obejero-Paz, C. A. & Jones, S. W. Evaluation of a two-site, three-barrier model for permeation in CaV3.1 (α1G) T-type calcium channels: Ca2+, Ba2+, Mg2+, and Na+ . J. Membr. Biol. 235, 131–143 (2010)
Heinemann, S. H., Terlau, H., Stuhmer, W., Imoto, K. & Numa, S. Calcium channel characteristics conferred on the sodium channel by single mutations. Nature 356, 441–443 (1992)
Ellinor, P. T., Yang, J., Sather, W. A., Zhang, J. F. & Tsien, R. W. Ca2+ channel selectivity at a single locus for high-affinity Ca2+ interactions. Neuron 15, 1121–1132 (1995)
Yang, J., Ellinor, P. T., Sather, W. A., Zhang, J. F. & Tsien, R. W. Molecular determinants of Ca2+ selectivity and ion permeation in L-type Ca2+ channels. Nature 366, 158–161 (1993)
Kim, M. S., Morii, T., Sun, L. X., Imoto, K. & Mori, Y. Structural determinants of ion selectivity in brain calcium channel. FEBS Lett. 318, 145–148 (1993)
Cibulsky, S. M. & Sather, W. A. The EEEE locus is the sole high-affinity Ca2+ binding structure in the pore of a voltage-gated Ca2+ channel: block by Ca2+ entering from the intracellular pore entrance. J. Gen. Physiol. 116, 349–362 (2000)
Cloues, R. K., Cibulsky, S. M. & Sather, W. A. Ion interactions in the high-affinity binding locus of a voltage-gated Ca2+ channel. J. Gen. Physiol. 116, 569–586 (2000)
Payandeh, J., Scheuer, T., Zheng, N. & Catterall, W. A. The crystal structure of a voltage-gated sodium channel. Nature 475, 353–358 (2011)
Payandeh, J., Gamal El-Din, T. M., Scheuer, T., Zheng, N. & Catterall, W. A. Crystal structure of a voltage-gated sodium channel in two potentially inactivated states. Nature 486, 135–139 (2012)
Zhang, X. et al. Crystal structure of an orthologue of the NaChBac voltage-gated sodium channel. Nature 486, 130–134 (2012)
Ren, D. et al. A prokaryotic voltage-gated sodium channel. Science 294, 2372–2375 (2001)
Yu, F. H. & Catterall, W. A. The VGL-chanome: a protein superfamily specialized for electrical signaling and ionic homeostasis. Sci. STKE 2004, re15 (2004)
Koishi, R. et al. A superfamily of voltage-gated sodium channels in bacteria. J. Biol. Chem. 279, 9532–9538 (2004)
Yue, L., Navarro, B., Ren, D., Ramos, A. & Clapham, D. E. The cation selectivity filter of the bacterial sodium channel, NaChBac. J. Gen. Physiol. 120, 845–853 (2002)
Morais-Cabral, J. H., Zhou, Y. & MacKinnon, R. Energetic optimization of ion conduction rate by the K+ selectivity filter. Nature 414, 37–42 (2001)
Alam, A. & Jiang, Y. Structural analysis of ion selectivity in the NaK channel. Nature Struct. Mol. Biol. 16, 35–41 (2009)
Alam, A., Shi, N. & Jiang, Y. Structural insight into Ca2+ specificity in tetrameric cation channels. Proc. Natl Acad. Sci. USA 104, 15334–15339 (2007)
Derebe, M. G. et al. Tuning the ion selectivity of tetrameric cation channels by changing the number of ion binding sites. Proc. Natl Acad. Sci. USA 108, 598–602 (2011)
McCleskey, E. W. & Almers, W. The Ca channel in skeletal muscle is a large pore. Proc. Natl Acad. Sci. USA 82, 7149–7153 (1985)
Chen, X. H. & Tsien, R. W. Aspartate substitutions establish the concerted action of P-region glutamates in repeats I and III in forming the protonation site of L-type Ca2+ channels. J. Biol. Chem. 272, 30002–30008 (1997)
Cibulsky, S. M. & Sather, W. A. Control of ion conduction in L-type Ca2+ channels by the concerted action of S5–6 regions. Biophys. J. 84, 1709–1719 (2003)
Williamson, A. V. & Sather, W. A. Nonglutamate pore residues in ion selection and conduction in voltage-gated Ca2+ channels. Biophys. J. 77, 2575–2589 (1999)
McCusker, E. C. et al. Structure of a bacterial voltage-gated sodium channel pore reveals mechanisms of opening and closing. Nature Commun. 3, 1102 (2012)
Shaya, D. et al. Structure of a prokaryotic sodium channel pore reveals essential gating elements and an outer ion binding site common to eukaryotic channels. J. Mol. Biol http://dx.doi.org/10.1016/j.jmb.2013.10.010 (published online 10 October 2013)
Faham, S. & Bowie, J. U. Bicelle crystallization: a new method for crystallizing membrane proteins yields a monomeric bacteriorhodopsin structure. J. Mol. Biol. 316, 1–6 (2002)
Faham, S. et al. Crystallization of bacteriorhodopsin from bicelle formulations at room temperature. Protein Sci. 14, 836–840 (2005)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Collaborative Computational Project, 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Laskowski, R. A., Moss, D. S. & Thornton, J. M. Main-chain bond lengths and bond angles in protein structures. J. Mol. Biol. 231, 1049–1067 (1993)
DeLano, W. L. PyMOL molecular viewer (V.1. 2r3pre) (http://www.pymol.org) (2002)
Gamal El-Din, T. M., Martinez, G. Q., Payandeh, J., Scheuer, T. & Catterall, W. A. A gating charge interaction required for late slow inactivation of the bacterial sodium channel NavAb. J. Gen. Physiol. 142, 181–190 (2013)
Acknowledgements
We are grateful to the beamline staff at the Advanced Light Source (BL8.2.1 and BL8.2.2) for their assistance during data collection. Research reported in this publication was supported by the National Institute of Neurological Disorders and Stroke (NINDS) of the National Institutes of Health (NIH) under award number R01NS015751 (W.A.C.), the National Heart, Lung, and Blood Institute (NHLBI) of the NIH under award number R01HL112808 (W.A.C. and N.Z.) and a National Research Service Award from training grant T32GM008268 (T.M.H.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. This work was also supported by the Howard Hughes Medical Institute (N.Z.).
Author information
Authors and Affiliations
Contributions
L.T., T.M.G.E.-D., J.P., T.S., N.Z. and W.A.C. designed the experiments. J.P. initiated the experimental work. L.T. conducted the protein purification, crystallization and diffraction experiments. L.T., J.P. and N.Z. determined and analysed the structures of the apo and cation-bound forms of CaVAb and the intermediate CaVAb constructs. T.M.G.E.-D. and T.S. performed physiological studies of CaVAb and related constructs. G.Q.M. and T.M.H. made the constructs and performed the preliminary data collection. All authors interpreted the structures in light of the physiological data. L.T., N.Z. and W.A.C. wrote the manuscript with input from all co-authors. W.A.C. and N.Z. are co-senior authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-9 and Supplementary Table 1. (PDF 13277 kb)
Rights and permissions
About this article
Cite this article
Tang, L., Gamal El-Din, T., Payandeh, J. et al. Structural basis for Ca2+ selectivity of a voltage-gated calcium channel. Nature 505, 56–61 (2014). https://doi.org/10.1038/nature12775
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12775
This article is cited by
-
Molecular details of ruthenium red pore block in TRPV channels
EMBO Reports (2024)
-
Structure of human NaV1.6 channel reveals Na+ selectivity and pore blockade by 4,9-anhydro-tetrodotoxin
Nature Communications (2023)
-
Calcium-permeable channelrhodopsins for the photocontrol of calcium signalling
Nature Communications (2022)
-
Mechanisms of membrane protein crystallization in ‘bicelles’
Scientific Reports (2022)
-
Transportation of calcium ions through chemically modified nanochannels in a polymeric membrane
Ionics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.