Abstract
The determination of protein crystal structures is hampered by the need for macroscopic crystals. X-ray free-electron lasers (FELs) provide extremely intense pulses of femtosecond duration, which allow data collection from nanometre- to micrometre-sized crystals1,2,3,4 in a ‘diffraction-before-destruction’ approach. So far, all protein structure determinations carried out using FELs have been based on previous knowledge of related, known structures1,2,3,4,5. Here we show that X-ray FEL data can be used for de novo protein structure determination, that is, without previous knowledge about the structure. Using the emerging technique of serial femtosecond crystallography1,2,3,4,6, we performed single-wavelength anomalous scattering measurements on microcrystals of the well-established model system lysozyme, in complex with a lanthanide compound. Using Monte-Carlo integration6,7, we obtained high-quality diffraction intensities from which experimental phases could be determined, resulting in an experimental electron density map good enough for automated building of the protein structure. This demonstrates the feasibility of determining novel protein structures using FELs. We anticipate that serial femtosecond crystallography will become an important tool for the structure determination of proteins that are difficult to crystallize, such as membrane proteins1,2,8.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Accessions
Protein Data Bank
Data deposits
Structure factor amplitudes and anomalous differences have been deposited in the Protein Data Bank along with the refined structure with accession code 4N5R, and diffraction patterns of crystal hits will be deposited at http://cxidb.org/.
References
Chapman, H. N. et al. Femtosecond X-ray protein nanocrystallography. Nature 470, 73–77 (2011)
Johansson, L. C. et al. Lipidic phase membrane protein serial femtosecond crystallography. Nature Methods 9, 263–265 (2012)
Boutet, S. et al. High-resolution protein structure determination by serial femtosecond crystallography. Science 337, 362–364 (2012)
Redecke, L. et al. Natively inhibited Trypanosoma brucei cathepsin B structure determined by using an X-ray laser. Science 339, 227–230 (2013)
Kern, J. et al. Room temperature femtosecond X-ray diffraction of photosystem II microcrystals. Proc. Natl Acad. Sci. USA 109, 9721–9726 (2012)
Kirian, R. A. et al. Femtosecond protein nanocrystallography-data analysis methods. Opt. Express 18, 5713–5723 (2010)
White, T. A. et al. CrystFEL: a software suite for snapshot serial crystallography. J. Appl. Cryst. 45, 335–341 (2012)
Kern, J. et al. Simultaneous femtosecond X-ray spectroscopy and diffraction of photosystem II at room temperature. Science 340, 491–495 (2013)
Owen, R. L., Rudino-Pinera, E. & Garman, E. F. Experimental determination of the radiation dose limit for cryocooled protein crystals. Proc. Natl Acad. Sci. USA 103, 4912–4917 (2006)
Spence, J. C. H. et al. Phasing of coherent femtosecond X-ray diffraction from size-varying nanocrystals. Opt. Express 19, 2866–2873 (2011)
Banumathi, S., Zwart, P. H., Ramagopal, U. A., Dauter, M. & Dauter, Z. Structural effects of radiation damage and its potential for phasing. Acta Crystallogr. D 60, 1085–1093 (2004)
Ravelli, R. B. G., Leiros, H.-K. S., Pan, B., Caffrey, M. & McSweeney, S. Specific radiation damage can be used to solve macromolecular crystal structures. Structure 11, 217–224 (2003)
Son, S.-K., Chapman, H. N. & Santra, R. Multiwavelength anomalous diffraction at high X-ray intensity. Phys. Rev. Lett. 107, 218102 (2011)
Weierstall, U., Spence, J. C. H. & Doak, R. B. Injector for scattering measurements on fully solvated biospecies. Rev. Sci. Instrum. 83, 035108 (2012)
Sierra, R. G. et al. Nanoflow electrospinning serial femtosecond crystallography. Acta Crystallogr. D 68, 1584–1587 (2012)
Kirian, R. A. et al. Structure-factor analysis of femtosecond micro-diffraction patterns from protein nanocrystals. Acta Crystallogr. A 67, 131–140 (2011)
Barends, T. R. M. et al. Anomalous signal from S atoms in protein crystallographic data from an X-ray free-electron laser. Acta Crystallogr. D 69, 838–842 (2013)
Girard, E., Chantalat, L., Vicat, J. & Kahn, R. Gd-HPDO3A, a complex to obtain high-phasing-power heavy-atom derivatives for SAD and MAD experiments: results with tetragonal hen egg-white lysozyme. Acta Crystallogr. D 58, 1–9 (2002)
Boutet, S. & Williams, G. J. The Coherent X-ray Imaging (CXI) instrument at the Linac Coherent Light Source (LCLS). New J. Phys. 12, 035024 (2010)
Foucar, L. et al. CASS-CFEL-ASG software suite. Comput. Phys. Commun. 183, 2207–2213 (2012)
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D 66, 479–485 (2010)
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Cowtan, K. ‘dm': An automated procedure for phase improvement by density modification. Joint CCP4 and ESF-EACBM Newsletter on Protein Crystallography 31,. 34–38 (1994)
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nature Protocols 3, 1171–1179 (2008)
Diederichs, K. & Karplus, P. A. Improved R-factors for diffraction data analysis in macromolecular crystallography. Nature Struct. Biol. 4, 269–275 (1997)
Weiss, M. S. Global indicators of X-ray data quality. J. Appl. Crystallogr. 34, 130–135 (2001)
Weiss, M. S., Sicker, T. & Hilgenfeld, R. Soft X-rays, high redundancy, and proper scaling: a new procedure for automated protein structure determination via SAS. Structure 9, 771–777 (2001)
Panjikar, S. & Tucker, P. A. Phasing possibilities using different wavelengths with a xenon derivative. J. Appl. Crystallogr. 35, 261–266 (2002)
Amann, J. et al. Demonstration of self-seeding in a hard-X-ray free-electron laser. Nature Photon. 6, 693–698 (2012)
Acknowledgements
Portions of this research were carried out at the Linac Coherent Light Source, a National User Facility operated by Stanford University on behalf of the US Department of Energy, Office of Basic Energy Sciences. The CXI instrument was funded by the LCLS Ultrafast Science Instruments (LUSI) project funded by the US Department of Energy, Office of Basic Energy Sciences. We acknowledge support from the Max Planck Society and from the EU for an Incoming Scientist Award to R.B.D. We thank the staff at the LCLS for their support and are grateful to S. Pesch and R. van Gessel (Bracco Imaging Konstanz and Singen, Germany) for the gift of the sample of gadoteridol. We thank H. Zimmermann for suggestions, W. Kabsch for discussions and J. Wray for critically reading the manuscript. In addition, we acknowledge L. Hammon and C. Patty for laboratory support, and the MCC staff for the beam they provided. We are indebted to C. Roome and F. Koeck for computing support.
Author information
Authors and Affiliations
Contributions
T.R.M.B. and I.S. conceived the research, I.S. prepared crystals, Sa.B., R.B.D. and R.L.S. performed sample injection. Sé.B., G.J.W., J.E.K. and M.M. performed data collection, T.R.M.B., L.F. and K.N. performed data processing and analysis. T.R.M.B. and I.S. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
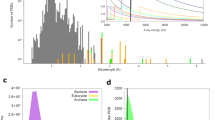
Extended Data Figure 1 Anomalous signal strength of the SFX data (blue lines) as well as the rotating anode data (red lines) as measured by Rano on intensities (solid lines).
The noise in the data is indicated in terms of Rsplit for the SFX data and Rp.i.m. for the rotating anode data (dashed lines).
Extended Data Figure 2 Expected anomalous signal strength for a SAD experiment on lysozyme.
Expected anomalous signal strength for a SAD experiment on lysozyme with 2 gadolinium atoms per protein molecule at 8.5 keV (top panel) and for a sulphur-SAD experiment on lysozyme with 10 sulphur atoms per protein molecule at 6.0 keV (bottom panel). In each case, an optimistic scenario with all anomalous scatterers ordered is shown (green line) as well as a pessimistic scenario in which 60% of the anomalous scatterers are ordered (blue line). This figure was prepared using the anomalous scattering web server at http://skuld.bmsc.washington.edu/scatter/AS_signal.html.
Supplementary information
Supplementary Information
This file contains Supplementary Methods and Supplementary References. (PDF 328 kb)
Rights and permissions
About this article
Cite this article
Barends, T., Foucar, L., Botha, S. et al. De novo protein crystal structure determination from X-ray free-electron laser data. Nature 505, 244–247 (2014). https://doi.org/10.1038/nature12773
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12773
This article is cited by
-
Structural heterogeneity assessment among the isoforms of fungal 1-aminocyclopropane-1-carboxylic acid (ACC) deaminase: a comparative in silico perspective
Journal of Genetic Engineering and Biotechnology (2022)
-
A simple pressure-assisted method for MicroED specimen preparation
Nature Communications (2021)
-
High-brightness self-seeded X-ray free-electron laser covering the 3.5 keV to 14.6 keV range
Nature Photonics (2021)
-
Illumination guidelines for ultrafast pump–probe experiments by serial femtosecond crystallography
Nature Methods (2020)
-
Viscosity-adjustable grease matrices for serial nanocrystallography
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.