Abstract
After fertilization, maternal factors direct development and trigger zygotic genome activation (ZGA) at the maternal-to-zygotic transition (MZT). In zebrafish, ZGA is required for gastrulation and clearance of maternal messenger RNAs, which is in part regulated by the conserved microRNA miR-430. However, the factors that activate the zygotic program in vertebrates are unknown. Here we show that Nanog, Pou5f1 (also called Oct4) and SoxB1 regulate zygotic gene activation in zebrafish. We identified several hundred genes directly activated by maternal factors, constituting the first wave of zygotic transcription. Ribosome profiling revealed that nanog, sox19b and pou5f1 are the most highly translated transcription factors pre-MZT. Combined loss of these factors resulted in developmental arrest before gastrulation and a failure to activate >75% of zygotic genes, including miR-430. Our results demonstrate that maternal Nanog, Pou5f1 and SoxB1 are required to initiate the zygotic developmental program and induce clearance of the maternal program by activating miR-430 expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
ten Bosch, J. R., Benavides, J. A. & Cline, T. W. The TAGteam DNA motif controls the timing of Drosophila pre-blastoderm transcription. Development 133, 1967–1977 (2006)
Liang, H. L. et al. The zinc-finger protein Zelda is a key activator of the early zygotic genome in Drosophila. Nature 456, 400–403 (2008)
Tadros, W. & Lipshitz, H. D. The maternal-to-zygotic transition: a play in two acts. Development 136, 3033–3042 (2009)
Andersen, I. S. et al. Epigenetic marking of the zebrafish developmental program. Curr. Top. Dev. Biol. 104, 85–112 (2013)
Vastenhouw, N. L. et al. Chromatin signature of embryonic pluripotency is established during genome activation. Nature 464, 922–926 (2010)
Lindeman, L. C. et al. Prepatterning of developmental gene expression by modified histones before zygotic genome activation. Dev. Cell 21, 993–1004 (2011)
Potok, M. E., Nix, D. A., Parnell, T. J. & Cairns, B. R. Reprogramming the maternal zebrafish genome after fertilization to match the paternal methylation pattern. Cell 153, 759–772 (2013)
Jiang, L. et al. Sperm, but not oocyte, DNA methylome is inherited by zebrafish early embryos. Cell 153, 773–784 (2013)
Kane, D. A. & Kimmel, C. B. The zebrafish midblastula transition. Development 119, 447–456 (1993)
Giraldez, A. J. et al. Zebrafish MiR-430 promotes deadenylation and clearance of maternal mRNAs. Science 312, 75–79 (2006)
Lund, E., Liu, M., Hartley, R. S., Sheets, M. D. & Dahlberg, J. E. Deadenylation of maternal mRNAs mediated by miR-427 in Xenopus laevis embryos. RNA 15, 2351–2363 (2009)
Giraldez, A. J. microRNAs, the cell’s Nepenthe: clearing the past during the maternal-to-zygotic transition and cellular reprogramming. Curr. Opin. Genet. Dev. 20, 369–375 (2010)
Harvey, S. A. et al. Identification of the zebrafish maternal and paternal transcriptomes. Development 140, 2703–2710 (2013)
Aanes, H. et al. Zebrafish mRNA sequencing deciphers novelties in transcriptome dynamics during maternal to zygotic transition. Genome Res. 21, 1328–1338 (2011)
Kaida, D. et al. U1 snRNP protects pre-mRNAs from premature cleavage and polyadenylation. Nature 468, 664–668 (2010)
Bazzini, A. A., Lee, M. T. & Giraldez, A. J. Ribosome profiling shows that miR-430 reduces translation before causing mRNA decay in zebrafish. Science 336, 233–237 (2012)
Chambers, I. & Tomlinson, S. R. The transcriptional foundation of pluripotency. Development 136, 2311–2322 (2009)
Onichtchouk, D. Pou5f1/oct4 in pluripotency control: insights from zebrafish. Genesis 50, 75–85 (2012)
Onichtchouk, D. et al. Zebrafish Pou5f1-dependent transcriptional networks in temporal control of early development. Mol. Syst. Biol. 6, 354 (2010)
Okuda, Y., Ogura, E., Kondoh, H. & Kamachi, Y. B1 SOX coordinate cell specification with patterning and morphogenesis in the early zebrafish embryo. PLoS Genet. 6, e1000936 (2010)
Lunde, K., Belting, H. G. & Driever, W. Zebrafish pou5f1/pou2, homolog of mammalian Oct4, functions in the endoderm specification cascade. Curr. Biol. 14, 48–55 (2004)
Hauptmann, G. et al. spiel ohne grenzen/pou2 is required for zebrafish hindbrain segmentation. Development 129, 1645–1655 (2002)
Belting, H. G. et al. Pou5f1 contributes to dorsoventral patterning by positive regulation of vox and modulation of fgf8a expression. Dev. Biol. 356, 323–336 (2011)
Xu, C. et al. Nanog-like regulates endoderm formation through the Mxtx2-Nodal pathway. Dev. Cell 22, 625–638 (2012)
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009)
Giraldez, A. J. et al. MicroRNAs regulate brain morphogenesis in zebrafish. Science 308, 833–838 (2005)
Loh, Y. H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nature Genet. 38, 431–440 (2006)
Leichsenring, M., Maes, J., Mossner, R., Driever, W. & Onichtchouk, D. Pou5f1 transcription factor controls zygotic gene activation in vertebrates. Science 341, 1005–1009 (2013)
Foygel, K. et al. A novel and critical role for Oct4 as a regulator of the maternal-embryonic transition. PLoS ONE 3, e4109 (2008)
Tan, M. H. et al. An Oct4-Sall4-Nanog network controls developmental progression in the pre-implantation mouse embryo. Mol. Syst. Biol. 9, 632 (2013)
Keramari, M. et al. Sox2 is essential for formation of trophectoderm in the preimplantation embryo. PLoS ONE 5, e13952 (2010)
Frum, T. et al. Oct4 cell-autonomously promotes primitive endoderm development in the mouse blastocyst. Dev. Cell 25, 610–622 (2013)
Messerschmidt, D. M. & Kemler, R. Nanog is required for primitive endoderm formation through a non-cell autonomous mechanism. Dev. Biol. 344, 129–137 (2010)
Gurdon, J. B., Elsdale, T. R. & Fischberg, M. Sexually mature individuals of Xenopus laevis from the transplantation of single somatic nuclei. Nature 182, 64–65 (1958)
Gurdon, J. B. & Melton, D. A. Nuclear reprogramming in cells. Science 322, 1811–1815 (2008)
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003)
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007)
Wernig, M. et al. In vitro reprogramming of fibroblasts into a pluripotent ES-cell-like state. Nature 448, 318–324 (2007)
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007)
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007)
Masui, S. et al. Pluripotency governed by Sox2 via regulation of Oct3/4 expression in mouse embryonic stem cells. Nature Cell Biol. 9, 625–635 (2007)
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nature Genet. 24, 372–376 (2000)
Soufi, A., Donahue, G. & Zaret, K. S. Facilitators and impediments of the pluripotency reprogramming factors’ initial engagement with the genome. Cell 151, 994–1004 (2012)
Judson, R. L., Babiarz, J. E., Venere, M. & Blelloch, R. Embryonic stem cell-specific microRNAs promote induced pluripotency. Nature Biotechnol. 27, 459–461 (2009)
Subramanyam, D. et al. Multiple targets of miR-302 and miR-372 promote reprogramming of human fibroblasts to induced pluripotent stem cells. Nature Biotechnol. 29, 443–448 (2011)
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010)
Amsterdam, A. et al. Identification of 315 genes essential for early zebrafish development. Proc. Natl Acad. Sci. USA 101, 12792–12797 (2004)
Matter, N. & Konig, H. Targeted ‘knockdown’ of spliceosome function in mammalian cells. Nucleic Acids Res. 33, e41 (2005)
Rösel, T. D. et al. RNA-Seq analysis in mutant zebrafish reveals role of U1C protein in alternative splicing regulation. EMBO J. 30, 1965–1976 (2011)
Iwafuchi-Doi, M. et al. The Pou5f1/Pou3f-dependent but SoxB-independent regulation of conserved enhancer N2 initiates Sox2 expression during epiblast to neural plate stages in vertebrates. Dev. Biol. 352, 354–366 (2011)
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009)
Collins, J. E., White, S., Searle, S. M. & Stemple, D. L. Incorporating RNA-seq data into the zebrafish Ensembl genebuild. Genome Res. 22, 2067–2078 (2012)
Berg, M. G. et al. U1 snRNP determines mRNA length and regulates isoform expression. Cell 150, 53–64 (2012)
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008)
Marson, A. et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134, 521–533 (2008)
Hamatani, T., Carter, M. G., Sharov, A. A. & Ko, M. S. Dynamics of global gene expression changes during mouse preimplantation development. Dev. Cell 6, 117–131 (2004)
Acknowledgements
We thank Y. Kamachi, W. Driever, A. F. Schier, N. Ivanova and I.-H. Park for reagents and fish lines; S. Mane and J. Overton for outstanding sequencing support; E. Zdobnov for hosting genomics data; A. Hubaud, N. Darricarrere and M. Koziol for contribution in the early stages of this project; and all the members of the Giraldez laboratory for discussions. This work was supported by NIH grants F32HD071697-02 (M.T.L.), T32GM007499 (A.R.B), F32HD061194-03 (C.M.T.), Pew Fellows Program in Biomedical Sciences (A.A.B), R01GM081602-06, R01GM103789-01, R01HD074078-02, the Pew Scholars Program in the Biomedical Sciences and the Yale Scholars Program (A.J.G.).
Author information
Authors and Affiliations
Contributions
M.T.L., A.R.B. and A.J.G. designed the project, performed experiments and data analysis. M.T.L., A.R.B., C.M.T. and A.J.G. wrote the manuscript. C.M.T. designed and performed the cycloheximide experiment and contributed to in situ hybridizations. A.A.B. designed and performed ribosome profiling and U1U2 experiments. K.R.D. and E.S.F. assisted with gene validation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
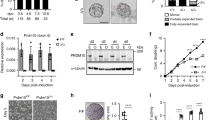
Extended Data Figure 1 Identifying de novo zygotic transcription.
a, Schematic of the sequencing strategy used in this study. Most zebrafish protein-coding genes (>95%) contain introns. De novo transcription produces intronic RNA sequences, which are spliced out of pre-mRNAs by the spliceosome, consisting of several ncRNA species including U1 and U2. b, Typical mRNA-seq applications use poly(A)+ selection to enrich for the mature mRNA population. Sequence reads map predominantly to exonic regions, with very few reads mapping to introns. During embryogenesis, many zygotic transcribed genes are expected to have a maternal contribution in the cytoplasm from the oocyte. The resulting signal will be a mixture of maternal-derived (orange) and zygotic-derived (blue) mRNA molecules, which cannot be deconvoluted without comparing to a reference sample to look for exon expression level change. c, mRNA-seq applications that skip poly(A)+ selection and instead use a rRNA depletion protocol (RiboZero) will not enrich for the mature mRNA population. Thus, transcripts in all stages of biogenesis (pre-mRNA, partially spliced mRNA, spliced introns) will be sequenced, and reads are expected to map both to exons and introns. Because maternally contributed mRNAs are mature, any intron signal detected must derive from de novo zygotic transcription. To determine the background signal for each intron, α-amanitin is used as a negative control for transcription. d, Morpholinos complementary to U1 and U2 injected into one-cell embryos inhibit zygotic splicing. Thus, pre-mRNAs fail to be processed, and the entire population of zygotic mRNAs will be unspliced. There are two benefits: (1) intron signal is amplified, as introns are stabilized in the pre-mRNA compared to spliced out introns; (2) protein production from zygotic mRNAs is effectively halted, as pre-mRNAs are generally not competent for normal translation. Only the first wave of transcription, resulting from activation by maternal factors, is observed. Transcription that requires zygotic proteins (subsequent waves) will be largely absent. e, The proportion of sequencing reads aligning to gene introns. Total RNA sequencing reveals elevated intronic sequence reads, corresponding to de novo zygotic transcription. f, The fate of the 5,318 sphere-stage (4 h.p.f.) zygotic genes that are only detectable through significant changes in intron sequence. At shield stage (6 h.p.f.), 64% of the genes are still detected as zygotically transcribed based only on intron signal. These include genes that have simultaneous zygotic transcription with decay of the maternal contribution. 30% of the genes are detected using both exon and intron signal by shield stage, indicating that transcription levels at sphere stage were too low to detect differences in exons, but were apparent in the introns. g, Number of genes detected in wild-type sphere-stage embryos, sphere embryos injected with U1U2 MO and wild-type shield-stage embryos, at different thresholds of detection. For both groups, a multiple test-corrected P < 0.1 threshold (Benjamin–Hochberg) was used for differential expression of exonic signal. For intronic signal, an uncorrected P < 0.1 was used for the ‘All detected’ group, whereas a multiple test-corrected P < 0.1 was used for the >5 RPKM gain group. h, Quantitative RT–PCR was performed for select genes to confirm zygotic transcription in wild-type sphere-stage embryos (dark blue bars) compared to α-amanitin-treated embryos (light blue bars). Primers were designed to amplify pre-mRNAs across exon–intron boundaries, except for cldne. Expression levels are reported as percentage of CT value compared to a maternally provided housekeeping gene (ef1a) (ΔCT × 100%). Error bars show s.e.m. for three technical replicates. Increased pre-mRNA levels were observed for all zygotic genes tested between wild type and α-amanitin. Maternally provided genes mtATP6 and mtND5 show no increase in wild type. Genes marked with an asterisk represent the bottom 10% of significant differential intron expression based on the RNA-seq data (which quantifies both pre-mRNA and spliced introns). This shows that using intron signal is a reliable indication of zygotic transcription. i, Genes detected in this study were compared to previous annotations of zygotic transcripts13, which used SNPs to identify transcripts derived from paternal alleles, to distinguish zygotic transcription from the maternal contribution. From their genomic sequencing results, we extracted 6,750 genes with informative exonic SNPs, which were consistently called between the two sets of matings. 178 of the genes we call zygotically transcribed at sphere stage at levels >5 RPKM are among the 6,750 informative genes. 87% of these are also found to be transcribed by ref. 13, with agreement between both strictly zygotic genes (Z) and maternal+zygotic genes (M+Z). 24 genes were not detected by ref. 13 (N.D.). At shield stage, 82% of the zygotic genes are also found by ref. 13, with 134 genes not detected. j, These undetected genes nevertheless have highly increased expression pre-64-cell to post-MZT (shield) using the RNA-seq data generated by ref. 13 (left) and in the current study (right). k, Cumulative plots show that SNP density is significantly lower among ref. 13 undetected genes at shield compared to detected genes (P = 1.6 × 10−3, two-sided Wilcoxon rank sum test), suggesting that low SNP density may account for the missed genes. l, Overall, ref. 13 and the current study distinguish a similar number of zygotic versus maternal transcripts at 6 h.p.f., among Ensembl genes with informative SNPs, with 74% agreement. However, 64% of zygotic transcripts identified in the current study do not have informative SNPs, and are thus not called transcribed by ref. 13. m, Genes called transcribed by ref. 13 but not in the current study have significantly higher intron signal than maternal genes (P = 1.4 × 10−95, two-sided Wilcoxon rank sum test), indicating that our significance threshold to detect zygotic transcription is conservative. n, Reference 14 used a time course poly(A)+ RNA-seq strategy to define zygotic transcripts. The comparable r70 Ensembl genes in the ref. 14 maternal+zygotic gene category are largely found in our study; however, we find thousands more transcribed genes based on intron signal—these genes represent transcription that is masked by the maternal contribution. o, Overall, our study captures most of the zygotic genes in the three categories described by ref. 14: maternal–zygotic genes (zygotic genes with maternal contribution, yellow), MBT genes (strictly zygotic genes detected at MBT, 3.5 h.p.f., orange), and post-MBT genes (strictly zygotic genes detected at 5.3 h.p.f., pink). Venn diagrams show the number of comparable r70 Ensembl genes that overlap between the two studies. Left panels include all zygotic genes detected in this study; right panels impose a zygotic expression threshold of >5 RPKM. Percentages within each box are calculated as the number of genes detected in this study (at either time point) that overlap the respective ref. 14 group, divided by the size of the ref. 14 group. The overlap percentages are generally high, indicating that our study recovered genes previously annotated as zygotically transcribed as well as many additional zygotic genes based on the use of intronic reads.
Extended Data Figure 2 Cycloheximide and U1U2 MO transcriptomes show first-wave genes.
a–c, Biplots comparing strictly zygotic genes found by either the current study or ref. 13 at >5 RPKM (N = 202). Zygotic expressed genes of ref. 13 were identified by comparing their raw RNA-seq data at 128-cell (pre-MZT) versus 3.5 h.p.f. In a, zygotic expression in U1U2 MO treated embryos (Total RNA, 4hpf) is compared to ref. 13 embryos treated with cycloheximide (CHX) (poly(A)+, assayed at 3.5 h.p.f.), which shows lagging expression of many first-wave genes (defined as having >5 RPKM in U1U2 MO). Genes verified by RT–PCR as first wave (klf4, nnr, sox11a, isg15, cldne) are highlighted, in addition to cldnb, which misses the threshold for first wave in the U1U2 MO transcriptome, and vox, which was highlighted by ref. 13. In b, c, Embryos treated with CHX and assayed in the current study at 4 h.p.f. and 6 h.p.f. (Total RNA) show gradual increases in expression of zygotic genes. Together these results suggest that expression of first-wave genes is independent of de novo zygotic factors, and that transcription overall is slower in CHX-treated embryos compared to wild type or U1U2 MO. d, Biplot showing gene expression levels (exonic) for all genes in U1U2 MO embryos at 4 h.p.f. compared to CHX-treated embryos assayed at 6 h.p.f. Magenta points, strictly zygotic genes; dark-blue points, maternal+zygotic genes. 97% of the first-wave genes called in U1U2 MO were expressed >1 RPKM in the CHX condition. e, Biplot comparing exonic expression levels between wild-type (4 h.p.f.) and CHX-treated embryos. Magenta points are strictly zygotic genes expressed >5 RPKM in wild type. The dotted line indicates 5 RPKM expression in CHX. f, Box-and-whisker plots comparing exonic expression level differences between wild-type and treated embryos in maternal genes, strictly zygotic multi-exon genes, and strictly zygotic single-exon genes. Both U1U2 MO and CHX-treated embryos show loss of expression in zygotic genes compared to wild type (U1U2 MO: P = 9.4 × 10−207 for multi-exonic, P = 4.2 × 10−4 for single exon, Wilcoxon rank-sum test comparing to maternal; CHX: P = 4.3 × 10−137 multi-exon, P = 1.5 × 10−6 single exon). The box defines the first and third quartiles, with the median indicated with a thick black line. The systemic decreases in expression in the U1U2 MO or CHX conditions compared to wild type indicate that although maternal factors can activate to a large extent expression of the first-wave genes, additional zygotic contribution of transcription factors (Nanog, SoxB1 and Pou5f1, but possibly others as well) might be required to reach wild-type levels of expression for many genes. This was also observed in ref. 13 for the gene vox. Alternatively, lower expression of first-wave zygotic genes might be caused by reduced level of maternal encoded proteins, as incubation with CHX at 32-cell stage might also decrease translation of the maternally deposited mRNAs. We consistently observe that CHX-treated embryos show lower/delayed expression compared with U1U2-MO-treated embryos, indicating that premature inhibition of maternal mRNA translation has an effect on the rate of activation of the first-wave genes. g, UCSC Genome Browser track showing an example of premature cleavage and polyadenylation (PCPA) for grhl3. Arrows indicate primer sites for RT–PCR. Previously, it was shown that U1 snRNA also serves to protect nascent mRNAs from PCPA, and that U1 inhibition results in 3′-truncation that may affect transcript level quantification56. h, RT–PCR for grhl3 on shield-stage embryos (N = 5). Wild-type (WT), U1U2 MO and CHX-treated embryos all amplify a 381-bp fragment from exon 1 to the beginning of intron 1. U1U2-MO-injected embryos amplify an unspliced 2,164-bp gene product spanning exon 1 to 3, whereas wild-type and CHX-treated embryos have a 294-bp spliced product, with α-amanitin as a negative control. i, Biplots comparing expression levels at the 5′ end of a transcript compared to the 3′ end, to detect PCPA at 4 h.p.f. Read density was assayed in up to 1,000 nucleotides of 5′ and 3′ sequence per transcript. The range of asymmetry values in wild type reflects sequencing biases or transcript annotation irregularities. Several genes in U1U2 MO embryos show elevated asymmetry compared to wild-type (orange dots, >twofold), reflecting a drop-off of read density moving 5′–3′ in the transcript, indicative of PCPA. These genes are included in our annotations of the zygotic first wave of expressed genes. The minor extent of PCPA during embryogenesis may reflect the short length of many of the zygotic genes, as PCPA is associated with longer genes that are likely to harbour cryptic polyadenylation sites. Transcripts in CHX-treated embryos generally do not show this trend.
Extended Data Figure 3 Verification of first-wave gene expression and functional categories.
a, To assay the embryonic specificity of the first-wave genes, we used publicly available microarray data from NCBI GEO across eight normal adult tissue types (brain, GSE11107; liver, GSE11107; heart, GSE17993; skin, GSE24528; kidney, GSE32363; digestive tract, GSE35889; ovary, GSE14979; testis, GSE14979) to classify genes as expressed specifically in the embryo (called ‘present’ by the MAS5 algorithm in 0–2 different adult tissues), genes expressed semi-specifically (present in 3–5 different adult tissues), and genes expressed ubiquitously (present in 6–8 different adult tissues); this latter group would correspond to ‘housekeeping’ genes. Sphere-stage first-wave genes consist of a mixture of specifically expressed and housekeeping genes. Subsequent-wave genes and genes expressed at levels <5 RPKM consist of a larger proportion of genes typically expressed ubiquitously in adult fish, suggesting a widespread activation of genes encoding general cellular processes in addition to developmentally specific ones. b, Gene Ontology enrichment analysis for first-wave, subsequent-wave and the low expressed genes with intronic RPKM >0.5. Top 5 scoring clusters are shown for each gene set. Clusters were defined using DAVID (http://david.abcc.ncifcrf.gov) Gene Functional Annotation Clustering on GO ‘FAT’ annotations and ‘high’ stringency. Clusters are annotated with representative GO terms and corresponding Benjamini–Hochberg FDR corrected P values. c, To validate genes activated in the first wave versus subsequent waves, RT–PCR was performed on shield stage (6 h.p.f.) in wild-type, α-amanitin, U1U2 MO and cycloheximide (CHX)-treated embryos. The unspliced products for nnr, isg15 and klf4 are detected only in U1U2 morphants, confirming that U1U2 is indeed blocking splicing. CHX treatment indicates the single-exon genes cldne and sox11a are activated in the first wave. cldnb is detected at low levels in wild type, as well as both U1U2 MO and CHX-treated embryos; however, based on RNA-seq levels at sphere stage, this gene does not pass the expression threshold to be called first wave. krt4 is significantly reduced in U1U2 MO and CHX-treated embryos, indicating that zygotic factors are required for its activation. Maternal tubb4b is present in all conditions. d–h, UCSC Genome Browser tracks for first-wave genes nnr, isg15, klf4, cldne and sox11a. i, UCSC Genome Browser track for cldnb, which shows low expression levels at sphere stage. j, k, UCSC Genome Browser track for a gene activated in subsequent waves (krt4) and for a maternally provided gene (tubb4b).
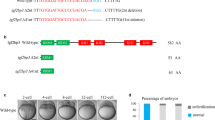
Extended Data Figure 4 Loss-of-function and rescue for Nanog, SoxB1 and Pou5f1.
a, Wild-type embryos were injected with Sox2, Sox3, Sox19a and Sox19b morpholinos individually and in combination (0.125 mM). Consistent with other reports, only quadruple LOF results in severe developmental defects (27 h.p.f.)20. LOF phenotype is rescued by injecting soxb1 mRNA (imaged at 24 h.p.f.). b, Wild-type and MZpou5f1 embryos were injected with SoxB1 MO (0.125mM each) and Nanog MO (0.6mM each) individually and in combination (Nanog + SoxB1). Loss of Nanog results in severe gastrulation defects and failure to progress past 80% epiboly, as previously reported24. Loss of SoxB1 in both wild-type and MZpou5f1 embryos showed developmental delay, whereas combined LOF for Nanog/SoxB1 or Pou5f1/Nanog completely arrested development before epiboly. Triple LOF embryos also arrested and failed to undergo gastrulation. c, Individual LOF for Nanog, SoxB1 and Pou5f1 resulted in developmental abnormalities (top panel). Embryos with Nanog LOF did not progress past 80% epiboly. The LOF phenotypes were rescued by injecting the respective mRNAs (LOF + mRNA) (bottom panel). Embryos imaged at 23 h.p.f. d, e, Wild-type and MZpou5f1 embryos were co-injected with Nanog + SoxB1 MO. LOF embryos arrest at sphere stage and resemble α-amanitin-injected embryos (+MO). Combinatorial LOF is rescued with co-injection of the respective mRNAs (MO + mRNA). Embryos were imaged when wild-type siblings reached 80% epiboly (d) and 24 h.p.f. (e). f, Ribosome profiling was performed at 2 h.p.f. on wild-type embryos and embryos injected with Nanog and SoxB1 morpholino at one-cell stage, to determine the specificity of the morpholinos to repress translation of nanog and soxB1 mRNA. Sequenced ribosome protected fragments (RPFs) were predominantly 28–29 nucleotides long, indicative of the width of the ribosome footprint. UCSC Genome Browser tracks (sense strand) showing ribosome profiling (top 2 tracks per gene) and input mRNA (bottom 2 tracks per gene). nanog and sox19b show significant reduction in RPFs in the Nanog MO + SoxB1 MO injected embryos compared to wild type. Input mRNA is unaffected. Neither h1m, a highly expressed gene, nor oep, a low expressed gene, has any change in either RPFs or input mRNA between wild-type and injected embryos.
Extended Data Figure 5 A transcriptome-wide effect is observed in LOF embryos.
a, b, Biplots comparing log2 RPKM exonic expression levels between time-matched wild-type and Nanog + SoxB1 + Pou5f1 LOF embryos (a); and between wild-type and triple LOF embryos co-injected with mRNA for nanog, soxB1 and pou5f1 (b) at 4 h.p.f., 6 h.p.f. and 8 h.p.f. Dark blue points highlight all strictly zygotic genes, whereas magenta points highlight the first-wave zygotic genes. miR-430 is highlighted at 4 h.p.f. in red, whereas green points indicate expression levels of (left to right) sox2, sox3, sox19a, sox19b and nanog. c, Plots showing proportion of the zygotic transcriptome affected (including first and subsequent waves). For sphere and shield stages and each LOF (Nanog MO, Nanog MO + SoxB1 MO, MZpou5f1 + Nanog MO + SoxB1 MO), dark blue regions represent genes with normal expression compared to wild type; light blue regions represent genes with significant loss of expression. Inner ring comprises zygotic genes with <1 RPKM of maternal contribution; outer ring comprises zygotic genes with maternal contribution. Percentages represent total affected genes in that condition over both gene categories. At sphere stage (4 h.p.f.) the effect for maternal and zygotic (M+Z) genes is weaker than for strictly zygotic genes, which may reflect a reduced power to detect changes due to the maternal contribution (see also Fig. 3b).
Extended Data Figure 6 Zygotic genes fail to be activated with Nanog, SoxB1 and Pou5f1 LOF.
a–f, In situ images showing that loss of Nanog and SoxB1 function results in a significant reduction in zygotic foxa3, blf, vent, foxd3, krt18 and ntla expression. LOF embryos (Nanog + SoxB1 MO) resemble α-amanitin-injected embryos by in situ, as well as in their transcriptome profiles. Loss of Nanog and SoxB1 is rescued by nanog and soxb1 mRNA (MO + mRNA), which is sufficient to restore wild-type expression profiles. g, h, In situ hybridization for zygotically transcribed cldne and cebpb shows that loss of Nanog and SoxB1 (Nanog + SoxB1 MO) has minimal effect on activation of cldne and cebpb. However, triple LOF shows a decrease in expression for both genes, as shown in the UCSC tracks. i–o, RT–PCR analysis (i) and UCSC Genome Browser tracks (j–o) for zygotic genes klf4b, vox, tbx16, mxtx2, her3 and sox32, showing differential expression of zygotic genes in LOF conditions. Expression levels were rescued by injecting nanog and soxb1 mRNA (MO + mRNA). Maternal hist1h2aa was present in the α-amanitin control. RT (−) indicates the absence of reverse transcriptase, to control for genomic DNA contamination. In UCSC tracks, loss of Nanog, SoxB1 and Pou5f1 in each sequenced condition is indicated by (−).
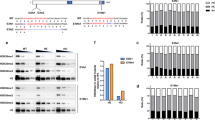
Extended Data Figure 7 Loss of function affects genes across functional categories in a combinatorial manner.
a, Comparisons of the single and double LOF transcriptomes to the triple LOF reveal that regulation is often combinatorial and redundant. Although all three factors seem to exert some influence on most of the transcribed genes, the effects observed in the combined LOF are not usually additive. Nanog seems to have the strongest individual effect of the three factors, but Pou5f1/SoxB1 can often act redundantly, or amplify the effect of Nanog alone. Venn diagrams show overlap between genes significantly downregulated at shield stage in single (pink), double (green) and triple (blue) LOF embryos. n = 2,172, left; n = 2,027, right. b, Pie charts showing the relative influence of each factor in the triple LOF. For each pie chart, genes downregulated in the triple LOF were compared in the single and double LOF transcriptomes. If the downregulation of a gene observed in the single LOF was less than twofold different from that observed in the triple LOF, the gene was considered to be regulated by the single factor alone. Otherwise, if the downregulation in the double was less than twofold different than the triple LOF, the gene was considered regulated by the combination of two factors. All remaining genes display the strongest downregulation in the triple LOF. Note that genes in each category may be affected by other combinations of LOF; however, the effect there is weaker. c, Breakdown of effects showing the redundancy of regulation in genes downregulated in the triple LOF. The largest category of genes seems to be regulated exclusively by Nanog (31%), as loss of Nanog function is equivalent to the triple LOF. 16% of genes seem to be regulated by both Nanog and Pou5f1 together, as loss of either Nanog alone or loss of Pou5f1 alone is sufficient to achieve the loss of function observed in the triple LOF. 16% of genes have equivalent effects with either Nanog LOF or Pou5f1 + SoxB1 double LOF, suggesting that Pou5f1 and SoxB1 act redundantly for these genes to co-regulate with Nanog. 9% of genes show the strongest effect only in the triple LOF. This suggests that there is redundancy between all three factors, as these genes can still be activated when one or two factors are lost. In all, 76% of the affected genes are subject to some form of redundant or combinatorial regulation. Asterisk indicates that for genes where the effect in the triple LOF was equivalent to both the double loss of SoxB1 and Nanog, and the double loss of SoxB1 and Pou5f1, we inferred that the effect was conferred by SoxB1 alone. d, Most genes are affected in the double or triple LOF conditions, across the gene categories defined in Extended Data Fig. 3a, including both embryo-specific genes and housekeeping (ubiquitously expressed) genes. e, Heat map showing specific embryonic functional categories of genes downregulated in LOF embryos. Three GO categories of genes expressed in wild type at shield stage are shown: general transcription factors, gastrulation and cell movement genes, and patterning genes (anterior–posterior axis and dorsal–ventral axis). Expression levels are represented as row-normalized values on a red–green colour scale for wild type (WT), α-amanitin treated (A), Nanog LOF (N), Nanog + SoxB1 LOF (NS), and Nanog + SoxB1 + Pou5f1 triple LOF (NSP). Widespread loss of expression is observed across these functional categories, with the triple LOF exhibiting the greatest similarity to α-amanitin.
Extended Data Figure 8 miR-430 activity requires Nanog function.
a, Schematic representation of miR-430 activity reporter GFP-3×IPT-miR-430 containing three complementary target sites to miR-430 (ref. 26). If maternal factor (M) is present, miR-430 is expressed and represses translation of the target mRNAs (no GFP expressed). Conversely, loss (X) of the maternal factor required for miR-430 activation would lead to a failure to repress miR-430 targets and GFP expression. dsRed is a control mRNA that is not subject to regulation by miR-430 and is co-injected with the target mRNA. b, GFP-reporter and dsRed (injection control) mRNAs were co-injected into embryos at one-cell stage and fluorescence assayed 7–8 h.p.f. GFP-reporter is repressed in wild-type and SoxB1 morphants by endogenous miR-430 (ref. 26), as shown by a decrease in GFP expression. The GFP-reporter fails to be repressed in α-amanitin (that fail to activate zygotic transcription and do not express miR-430) and Nanog-MO-injected embryos, indicating a loss of miR-430 activity. c, In situ hybridization for maternal miR-430 target gene cd82b. At shield stage, cd82b is cleared from wild-type and MZpou5f1 embryos. Combined Nanog, SoxB1 and Pou5f1 LOF causes a failure in clearance (MZpou5f1 + Nanog + SoxB1 MO). Injection of nanog, soxb1 and pou5f1 mRNA rescues the phenotype (MO + mRNA). d, Cumulative plots showing the effect of each LOF condition on miR-430 target repression, as in ref. 16, using Total RNA-seq. Plots show the distribution of log2 fold expression level difference for each condition relative to wild type in three groups of genes defined in ref. 16: miR-430 targets with multiple 7mer or 8mer seed target sites in their 3′ UTR; miR-430 targets with a single 7mer or 8mer seed in the 3′ UTR; and genes lacking miR-430 seed sites in their 3′ UTRs. P values are for two-sided Wilcoxon rank-sum tests comparing each of the two miR-430 target groups to the non-targets. MZdicer expression data are from ref. 16. Displacement of the curve to the left (−) from the grey control line indicates a larger fraction of genes are accumulated (fail to be degraded) in the indicated condition compared to wild type. Nanog has the strongest effect, although there is also an effect from the combined loss of Pou5f1 and SoxB1. e, Cumulative plots showing the effect of triple LOF with and without mRNA rescue on miR-430 target repression, using poly(A)+ selection RNA-seq. At 6 h.p.f., miR-430 targets fail to be degraded in the LOF condition compared to wild type, with expression levels of targets high in the LOF relative to wild type. Co-injection of nanog, soxB1 and pou5f1 mRNAs restores miR-430 activity, and the targets’ expression levels are restored to near wild-type levels. f, At 8 h.p.f., miR-430 targets are still undegraded in the LOF, but are degraded to wild-type levels in the rescue. P values are for two-sided Wilcoxon rank-sum tests comparing each of the two miR-430 target groups to the non-targets.
Extended Data Figure 9 Nanog, Pou5f1 and SoxB1 bind to and regulate embryonic genes.
a, Nanog chromatin immunoprecipitation sequencing binding data in zebrafish at 3.3 h.p.f. (ref. 24) was re-analysed to determine Nanog-bound regions genome wide. Pie charts show percentage of genes in each category that are associated with Nanog bound regions (±5 kb). 74% of first-wave genes detected at sphere were associated with Nanog binding, twofold higher than subsequent-wave genes (P = 3.7 × 10−29, two-sided Fisher’s exact test). Low expressed zygotic genes are also less associated with Nanog-bound regions. For those genes that are nonetheless affected by Nanog LOF, this suggests that they are influenced by Nanog indirectly, rather than through Nanog binding at the gene locus. The enrichment of Nanog binding on the first-wave genes versus subsequent waves supports a model where Nanog has a central role in the regulation of the activation of the first wave of zygotic transcription. b, ChIP-seq data for Nanog, Oct4 and Sox2 in mouse embryonic stem cells57,58 were used to examine the binding profiles of genes transcribed during pre-implantation mouse embryogenesis59, as ChIP data do not exist for early mouse embryos. Three gene groups were analysed: α-amanitin-sensitive genes expressed at early 2-cell stage (minor wave ZGA), α-amanitin sensitive genes expressed at late 2-cell stage (major wave ZGA), and genes expressed during the 4–8-cell stages (mid-preimplantation). Gene promoters (defined to be 5 kb upstream to 50 bp downstream the annotated transcription start site of a gene) are highly enriched in binding sites among the genes comprising ZGA, as compared to the genome as a whole (P = 4.03 × 10−7 for the minor wave, P = 6.05 × 10−18 major wave, two-sided Fisher’s exact test). Genomic coordinates (mm8) for genes were defined by NIA/NIH U-cluster annotations for the microarray probes in ref. 59. Note that not all of the genes expressed during ZGA are necessarily expressed in ES cells; thus, the binding proportions are likely to be underestimates. Although these represent two different states of development, these results are consistent with a role for these factors in activating the earliest waves of zygotic gene expression also in mammals. c, Model showing maternal gene expression in red and zygotic gene expression in blue during the maternal to zygotic transition. Gene expression is depicted on the y axis and time on the x axis. During the MZT, Nanog, SoxB1 and Pou5f1 are required to activate a large fraction of zygotic genes, including miR-430, which in turn is responsible for the clearance of a significant portion of maternal mRNAs. In the loss of function of Nanog, SoxB1 and Pou5f1, there is a reduction in zygotic gene activation, causing a failure in the establishment of the zygotic developmental program, including loss of miR-430 expression and maternal mRNA clearance.
Supplementary information
Supplementary Data 1
This tab-delimited text file contains a list of genes with their expression calls and categories. (TXT 3086 kb)
Rights and permissions
About this article
Cite this article
Lee, M., Bonneau, A., Takacs, C. et al. Nanog, Pou5f1 and SoxB1 activate zygotic gene expression during the maternal-to-zygotic transition. Nature 503, 360–364 (2013). https://doi.org/10.1038/nature12632
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12632
This article is cited by
-
Canalizing cell fate by transcriptional repression
Molecular Systems Biology (2024)
-
Activator-blocker model of transcriptional regulation by pioneer-like factors
Nature Communications (2023)
-
Maternal TDP-43 interacts with RNA Pol II and regulates zygotic genome activation
Nature Communications (2023)
-
Sleeping embryonic genomes are awoken by OBOX proteins
Nature (2023)
-
OBOX regulates mouse zygotic genome activation and early development
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.